Cell-Cell Communication Network
Cell-Cell Communication Network
Cell-cell communication plotImage loading error!
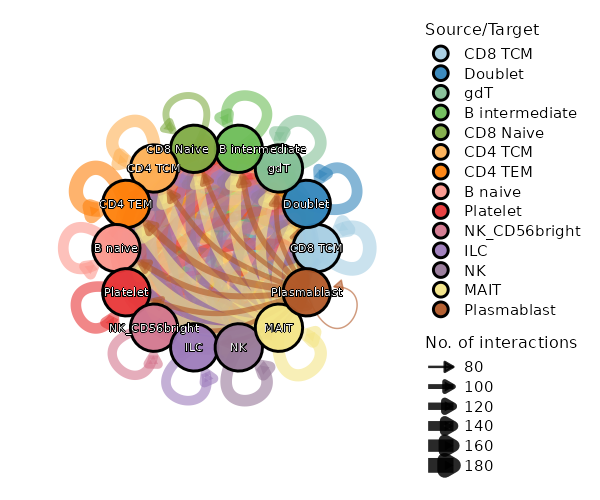
Cell-Cell Communication Circos Plot
Cell-Cell Communication Circos Plot
Cell-cell communication plotImage loading error!
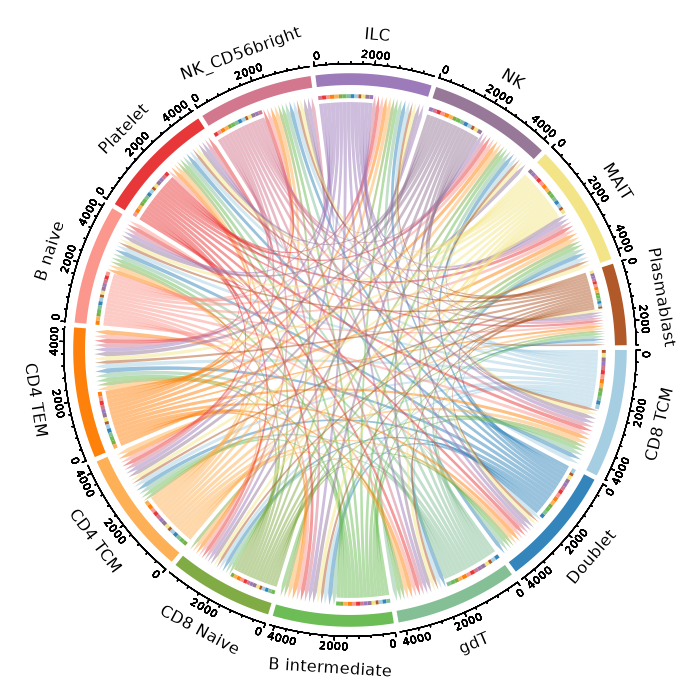
Cell-Cell Communication Heatmap
Cell-Cell Communication Heatmap
Cell-cell communication plotImage loading error!
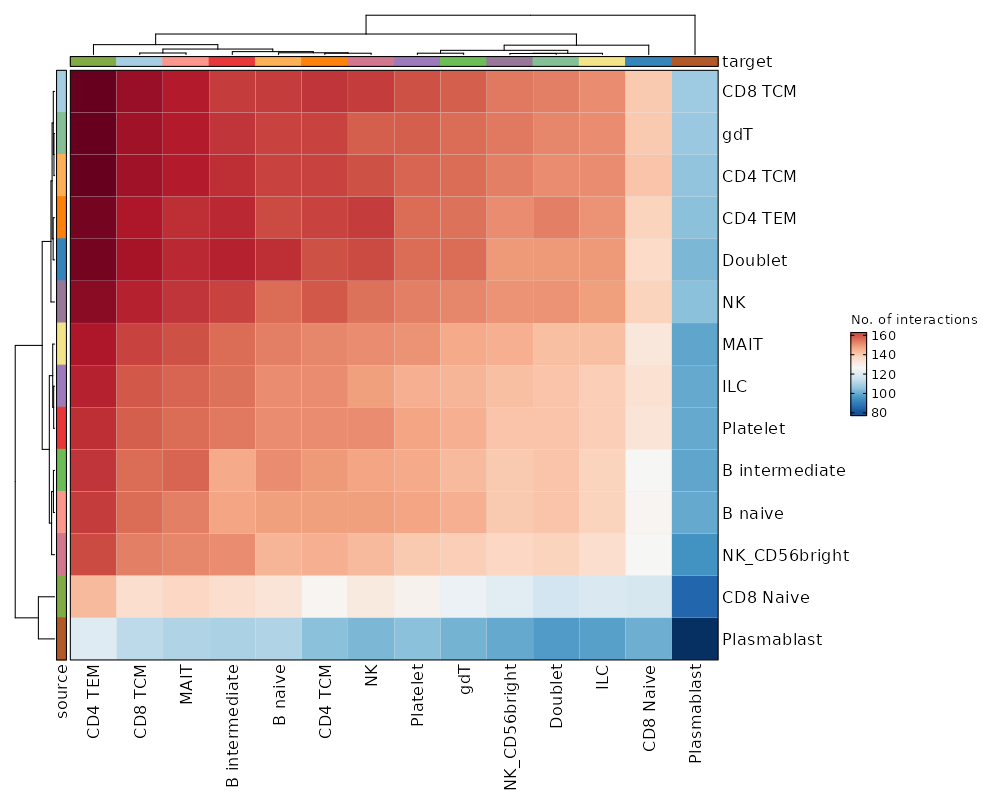
Cell-Cell Communication Interaction (Box Plot)
Cell-Cell Communication Interaction (Box Plot)
Cell-cell communication plotImage loading error!

Running Information
Showing the first job only. Check the workdir for information of other jobs if any.
# Generated by pipen_runinfo v0.9.3
# Lang: R
R version 4.3.3 (2024-02-29)
Platform: x86_64-conda-linux-gnu (64-bit)
Running under: Debian GNU/Linux 12 (bookworm)
Matrix products: default
BLAS/LAPACK: /opt/conda/lib/libopenblasp-r0.3.30.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: Etc/UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] biopipen.utils_0.3.2 scplotter_0.6.2 dplyr_1.1.4
[4] rlang_1.1.6
loaded via a namespace (and not attached):
[1] cubature_2.0.4.6 RcppAnnoy_0.0.22
[3] splines_4.3.3 later_1.4.4
[5] bitops_1.0-9 tibble_3.3.0
[7] polyclip_1.10-7 enrichit_0.0.4
[9] fastDummies_1.7.5 lifecycle_1.0.4
[11] Rdpack_2.6.4 doParallel_1.0.17
[13] globals_0.18.0 lattice_0.22-7
[15] MASS_7.3-60.0.1 magrittr_2.0.3
[17] plotly_4.11.0 httpuv_1.6.16
[19] Seurat_5.3.0 sctransform_0.4.2
[21] spam_2.11-1 sp_2.2-0
[23] spatstat.sparse_3.1-0 reticulate_1.43.0
[25] cowplot_1.2.0 pbapply_1.7-4
[27] RColorBrewer_1.1-3 abind_1.4-5
[29] zlibbioc_1.48.0 Rtsne_0.17
[31] GenomicRanges_1.54.1 purrr_1.1.0
[33] ggraph_2.2.1 BiocGenerics_0.48.1
[35] RCurl_1.98-1.17 tweenr_2.0.3
[37] evmix_2.12 circlize_0.4.16
[39] GenomeInfoDbData_1.2.11 IRanges_2.36.0
[41] S4Vectors_0.40.2 ggrepel_0.9.6
[43] irlba_2.3.5.1 listenv_0.9.1
[45] spatstat.utils_3.1-5 iNEXT_3.0.2
[47] MatrixModels_0.5-4 goftest_1.2-3
[49] RSpectra_0.16-2 scRepertoire_2.2.1
[51] spatstat.random_3.4-1 fitdistrplus_1.2-4
[53] parallelly_1.45.1 codetools_0.2-20
[55] DelayedArray_0.28.0 xml2_1.4.0
[57] ggforce_0.5.0 tidyselect_1.2.1
[59] shape_1.4.6.1 farver_2.1.2
[61] viridis_0.6.5 matrixStats_1.1.0
[63] stats4_4.3.3 spatstat.explore_3.5-2
[65] jsonlite_2.0.0 GetoptLong_1.0.5
[67] ellipsis_0.3.2 tidygraph_1.3.0
[69] progressr_0.15.1 iterators_1.0.14
[71] ggridges_0.5.7 ggalluvial_0.12.5
[73] survival_3.8-3 systemfonts_1.2.3
[75] foreach_1.5.2 tools_4.3.3
[77] ggnewscale_0.5.2 ragg_1.5.0
[79] stringdist_0.9.15 ica_1.0-3
[81] Rcpp_1.1.0 glue_1.8.0
[83] gridExtra_2.3 SparseArray_1.2.2
[85] metap_1.1 qvalue_2.34.0
[87] MatrixGenerics_1.14.0 GenomeInfoDb_1.38.1
[89] withr_3.0.2 fastmap_1.2.0
[91] fansi_1.0.6 ttservice_0.5.3
[93] SparseM_1.84-2 digest_0.6.37
[95] gridGraphics_0.5-1 R6_2.6.1
[97] mime_0.13 textshaping_0.4.0
[99] colorspace_2.1-1 Cairo_1.6-2
[101] scattermore_1.2 tensor_1.5.1
[103] dichromat_2.0-0.1 spatstat.data_3.1-8
[105] tidyr_1.3.1 generics_0.1.4
[107] data.table_1.17.8 graphlayouts_1.2.2
[109] httr_1.4.7 htmlwidgets_1.6.4
[111] S4Arrays_1.2.0 uwot_0.2.3
[113] pkgconfig_2.0.3 gtable_0.3.6
[115] ComplexHeatmap_2.18.0 lmtest_0.9-40
[117] SingleCellExperiment_1.24.0 XVector_0.42.0
[119] htmltools_0.5.8.1 dotCall64_1.2
[121] clue_0.3-66 SeuratObject_5.2.0
[123] scales_1.4.0 Biobase_2.62.0
[125] png_0.1-8 spatstat.univar_3.1-4
[127] ggdendro_0.2.0 reshape2_1.4.4
[129] rjson_0.2.23 nlme_3.1-168
[131] cachem_1.1.0 zoo_1.8-14
[133] GlobalOptions_0.1.2 stringr_1.5.2
[135] KernSmooth_2.23-26 parallel_4.3.3
[137] miniUI_0.1.2 logger_0.4.0
[139] pillar_1.11.0 grid_4.3.3
[141] vctrs_0.6.5 RANN_2.6.2
[143] VGAM_1.1-13 promises_1.3.3
[145] xtable_1.8-4 cluster_2.1.8.1
[147] truncdist_1.0-2 cli_3.6.5
[149] compiler_4.3.3 crayon_1.5.3
[151] future.apply_1.20.0 labeling_0.4.3
[153] plyr_1.8.9 forcats_1.0.0
[155] stringi_1.8.7 viridisLite_0.4.2
[157] tidyseurat_0.8.0 deldir_2.0-4
[159] assertthat_0.2.1 gsl_2.1-8
[161] lazyeval_0.2.2 spatstat.geom_3.5-0
[163] quantreg_6.1 Matrix_1.6-5
[165] RcppHNSW_0.6.0 patchwork_1.3.2
[167] future_1.67.0 ggplot2_3.5.2
[169] shiny_1.11.1 plotthis_0.8.2
[171] SummarizedExperiment_1.32.0 rbibutils_2.3
[173] evd_2.3-7.1 ROCR_1.0-11
[175] gridtext_0.1.5 igraph_2.1.4
[177] memoise_2.0.1 bracer_1.2.1
# Generated by pipen-runinfo v0.9.3
Command: stdbuf -oL Rscript /mnt/disks/.cwd/.pipen/Immunopipe/CellCellCommunicationPlots/0/job.script
Voluntary context switches: 1039
Involuntary context switches: 321
Percentage of CPU this job got: 117%
Major page faults: 0
Minor page faults: 262961
Maximum resident set size (kB): 1056120
Elapsed real time (s): 25.74
System (kernel) time (s): 0.46
User time (s): 29.79
Exit status: 0
# Generated by pipen-runinfo v0.9.3
Scheduler
---------
local
Powered by
pipen v0.17.24
and
pipen-report v0.23.14