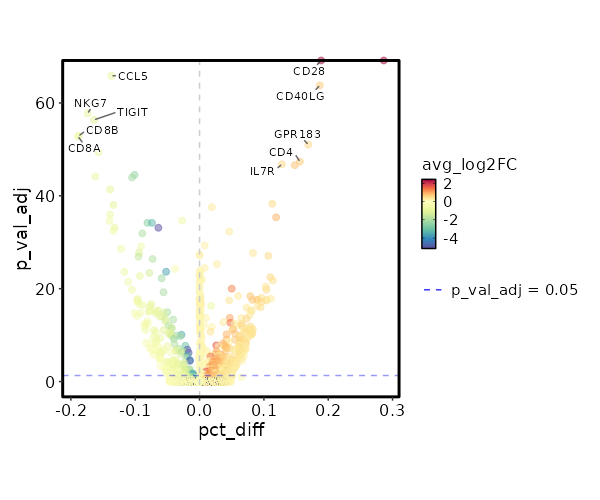
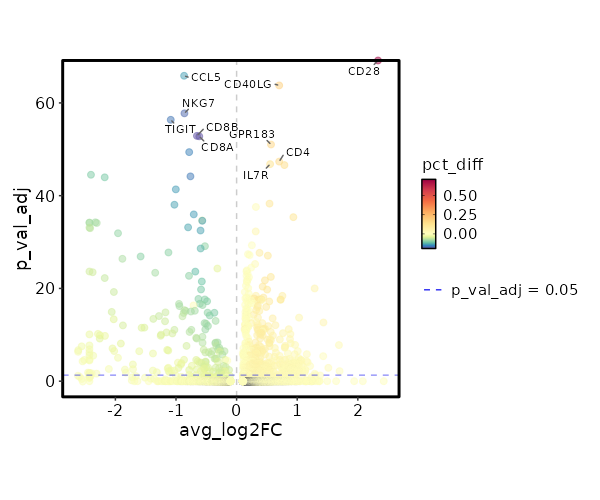
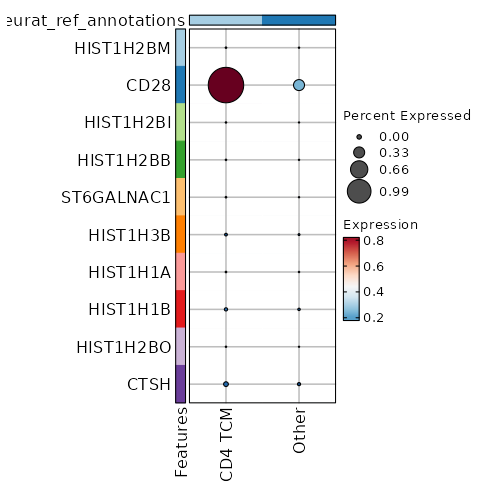
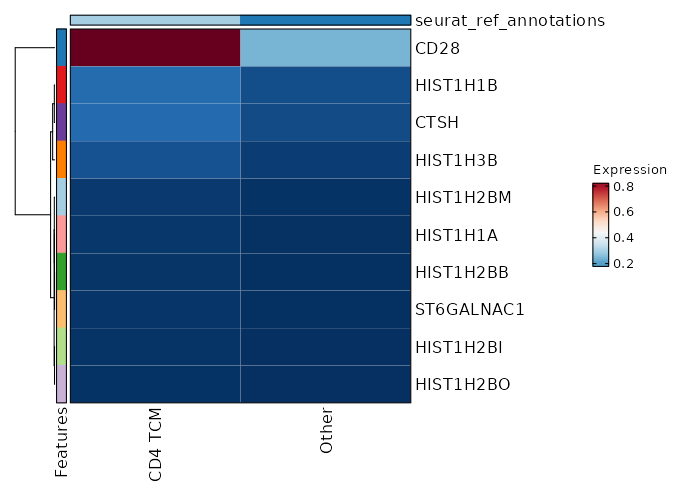
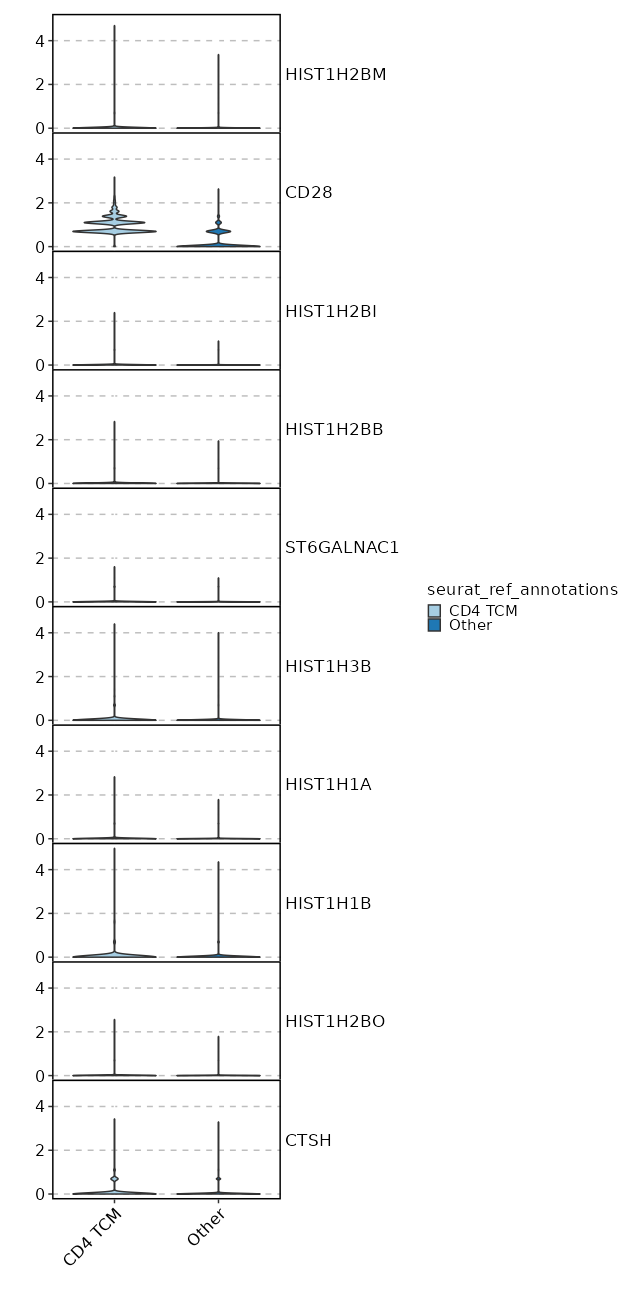
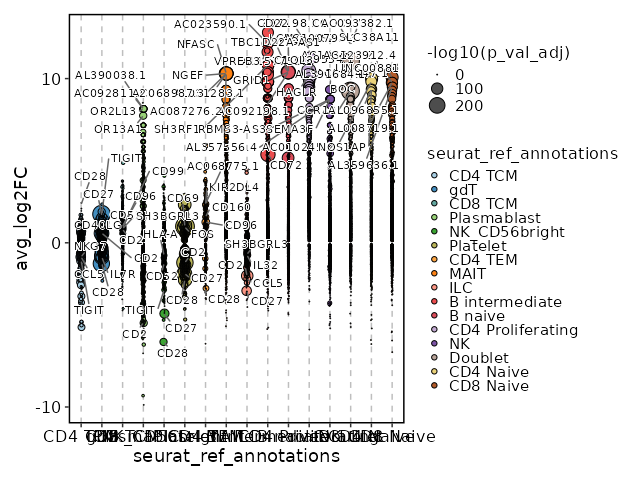
Cluster seurat_ref_annotations: CD4 TCM Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
0 2.3333 0.993 0.282 0 CD28 CD4 TCM 5.051e-69 0.7024 0.498 0.311 1.694e-64 CD40LG CD4 TCM 2.804e-56 0.5677 0.674 0.505 9.403e-52 GPR183 CD4 TCM 1.339e-52 0.7026 0.451 0.295 4.492e-48 CD4 CD4 TCM 4.721e-52 0.5532 0.76 0.632 1.583e-47 IL7R CD4 TCM 7.549e-52 0.7891 0.529 0.381 2.532e-47 AQP3 CD4 TCM 1.555e-43 0.5433 0.819 0.706 5.215e-39 LTB CD4 TCM 8.125e-43 0.3183 0.969 0.95 2.725e-38 FXYD5 CD4 TCM 1.277e-40 0.9366 0.313 0.194 4.284e-36 CCR6 CD4 TCM 1.541e-37 0.3153 0.968 0.922 5.168e-33 DUSP1 CD4 TCM 1.459e-34 0.2517 0.99 0.982 4.893e-30 EEF1B2 CD4 TCM 6.324e-33 0.3779 0.871 0.788 2.121e-28 SPOCK2 CD4 TCM 1.228e-32 0.188 1 1 4.118e-28 EEF1A1 CD4 TCM 1.984e-32 0.2146 1 1 6.655e-28 RPS8 CD4 TCM 2.553e-32 0.5156 0.644 0.537 8.563e-28 ICOS CD4 TCM 1.606e-30 0.3055 0.946 0.919 5.385e-26 LDHB CD4 TCM 9.806e-30 0.2409 0.987 0.979 3.289e-25 EEF1G CD4 TCM 3.846e-29 0.1695 1 0.999 1.29e-24 RPS3A CD4 TCM 6.157e-29 0.162 1 0.999 2.065e-24 RPL32 CD4 TCM 2.053e-28 0.1825 1 1 6.886e-24 RPS12 CD4 TCM
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
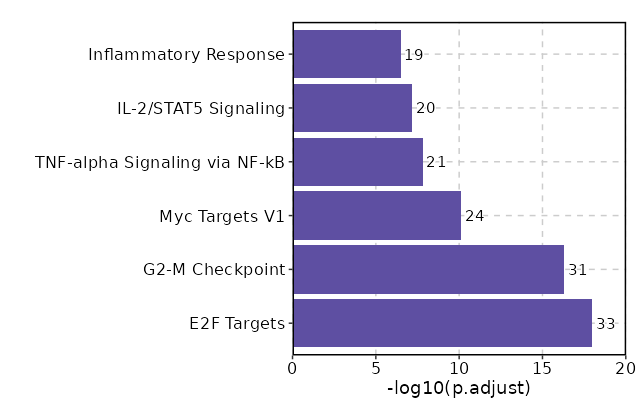
Enrichment Analysis E2F Targets 33/200 2.269e-20 1.067e-18 10.018 453.1361 CDC20;ANP32E;CHEK1;MYC;PLK4;PLK1;CDCA3;CDCA8;PRDX4;MYBL2;CDKN3;BRCA1;RAD51AP1;TK1;TRIP13;TIMELESS;SPC25;CDC25B;MELK;KIF4A;AURKB;DIAPH3;KIF18B;BIRC5;KIF2C;MXD3;CCNB2;CDK1;HMMR;PSIP1;ORC6;RRM2;SPAG5 1 MSigDB_Hallmark_2020 G2-M Checkpoint 31/200 2.171e-18 5.102e-17 9.2502 376.2185 CDC20;CHEK1;RAD54L;MYC;E2F2;PLK4;PLK1;HIF1A;MYBL2;CDKN3;DTYMK;KIF15;PBK;CENPA;NUSAP1;GINS2;UBE2C;CDC25B;KIF4A;AURKB;NDC80;CCNA2;BIRC5;TROAP;KIF2C;CCNB2;NUP50;SFPQ;CDK1;HMMR;ORC6 2 MSigDB_Hallmark_2020 Myc Targets V1 24/200 0 0 6.7514 175.697 RPL22;CNBP;CDC20;MYC;RPL14;RPL18;RPL6;RPL34;PRDX4;RPS10;PABPC4;C1QBP;RPLP0;RPS6;TYMS;FBL;RPS5;RPS2;RPS3;CCNA2;EEF1B2;YWHAQ;RRM1;IMPDH2 3 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 21/200 0 0 5.7635 117.8422 MYC;GPR183;JUNB;DUSP1;CCL20;BTG1;SOCS3;IFNGR2;FOS;CD44;IL6ST;CD69;CD83;PPP1R15A;IL7R;SLC2A3;NFKBIA;TNFAIP8;ATP2B1;KLF2;KLF6 4 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 20/199 0 0 5.4751 102.8147 AHNAK;PNP;FLT3LG;MYC;RNH1;IL18R1;CCR4;ICOS;ADAM19;CD48;CD44;CD83;SLC2A3;RORA;TNFRSF4;ARL4A;PHTF2;TIAM1;KLF6;LTB 5 MSigDB_Hallmark_2020 Inflammatory Response 19/200 0 0 5.1304 87.1841 SLAMF1;MYC;GPR183;HIF1A;CCL20;EMP3;IL18R1;TIMP1;CCR7;IFNGR2;CD48;KCNA3;CD69;CD82;IL7R;NFKBIA;ATP2B1;KLF6;GNA15 6 MSigDB_Hallmark_2020 Allograft Rejection 18/200 0 0 4.8213 74.0211 CD2;CD4;FYB1;HIF1A;RPL9;RPL39;CCR2;CD28;BRCA1;TIMP1;IFNGR2;CD40LG;CD47;RPS9;RPS3A;LTB;ITGB2;TRAT1 7 MSigDB_Hallmark_2020 Mitotic Spindle 15/199 0 0.0001 3.9439 42.8479 PLK1;ANLN;KIF15;NUSAP1;KIF4A;NDC80;BIRC5;KIF2C;PXN;ARHGAP5;CCNB2;CDK1;ECT2;MID1IP1;TIAM1 8 MSigDB_Hallmark_2020 p53 Pathway 14/200 0.0001 0.0004 3.632 34.2278 APP;RPL18;PLK3;RPL36;RPS12;BTG1;HINT1;LDHB;FOS;S100A4;CD82;DGKA;PPP1R15A;SESN1 9 MSigDB_Hallmark_2020 Apoptosis 12/161 0.0001 0.0007 3.8743 34.287 CDC25B;CD2;CD69;APP;LEF1;SATB1;ANXA1;CYLD;BRCA1;TIMP1;CD44;GNA15 10 MSigDB_Hallmark_2020 mTORC1 Signaling 12/200 0.001 0.0044 3.0645 21.1014 PNP;PLK1;PRDX1;SERP1;CXCR4;PPP1R15A;M6PR;SLC2A3;DAPP1;RRM2;ITGB2;ADD3 11 MSigDB_Hallmark_2020 Hypoxia 11/200 0.0032 0.0115 2.7872 15.9921 PLAC8;DUSP1;BTG1;FOS;CXCR4;MAP3K1;S100A4;PPP1R15A;SLC2A3;RORA;KLF6 12 MSigDB_Hallmark_2020 Estrogen Response Late 11/200 0.0032 0.0115 2.7872 15.9921 CDC20;KIF20A;PLK4;FOS;CD44;GINS2;IL6ST;FKBP5;IGFBP4;TIAM1;ADD3 12 MSigDB_Hallmark_2020 Reactive Oxygen Species Pathway 5/49 0.0034 0.0115 5.4026 30.6808 CDKN2D;PRDX4;PRDX2;PRDX1;JUNB 14 MSigDB_Hallmark_2020 UV Response Up 9/158 0.0059 0.0185 2.8843 14.7985 IL6ST;AQP3;SIGMAR1;JUNB;GGH;NFKBIA;BTG1;DNAJB1;FOS 15 MSigDB_Hallmark_2020 Estrogen Response Early 10/200 0.0093 0.025 2.5141 11.7588 AQP3;MYC;FOS;CD44;IL6ST;FKBP5;IGFBP4;FCMR;TIAM1;ADD3 16 MSigDB_Hallmark_2020 Interferon Gamma Response 10/200 0.0093 0.025 2.5141 11.7588 IFITM2;PNP;IFI44;HIF1A;BTG1;SOCS3;CD69;ARID5B;NFKBIA;ARL4A 16 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 6/87 0.0096 0.025 3.5236 16.3755 IL6ST;IL18R1;SOCS3;IFNGR2;LTB;CD44 18 MSigDB_Hallmark_2020 Angiogenesis 3/36 0.0384 0.0951 4.3035 14.023 S100A4;APP;TIMP1 19 MSigDB_Hallmark_2020 Wnt-beta Catenin Signaling 3/42 0.0565 0.1246 3.6403 10.461 TCF7;LEF1;MYC 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
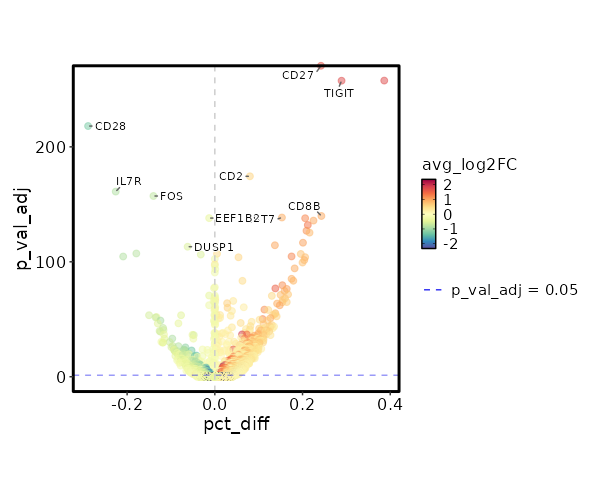
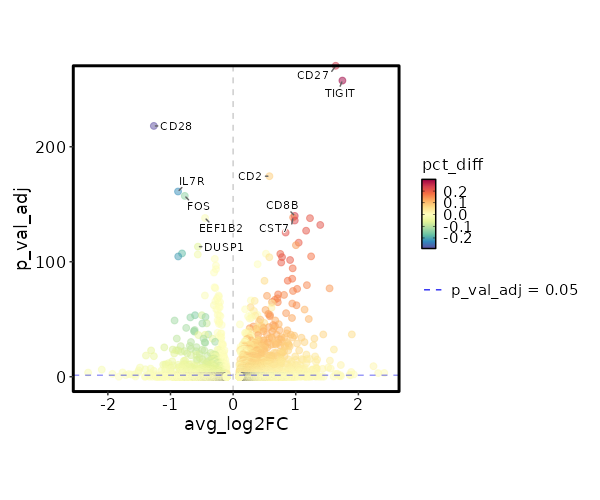
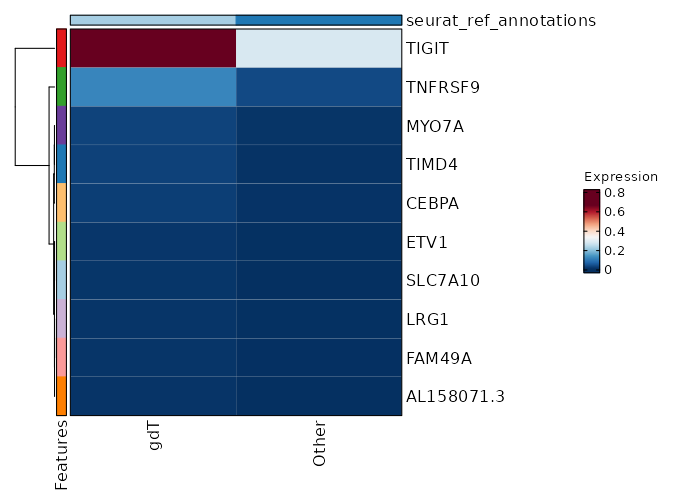
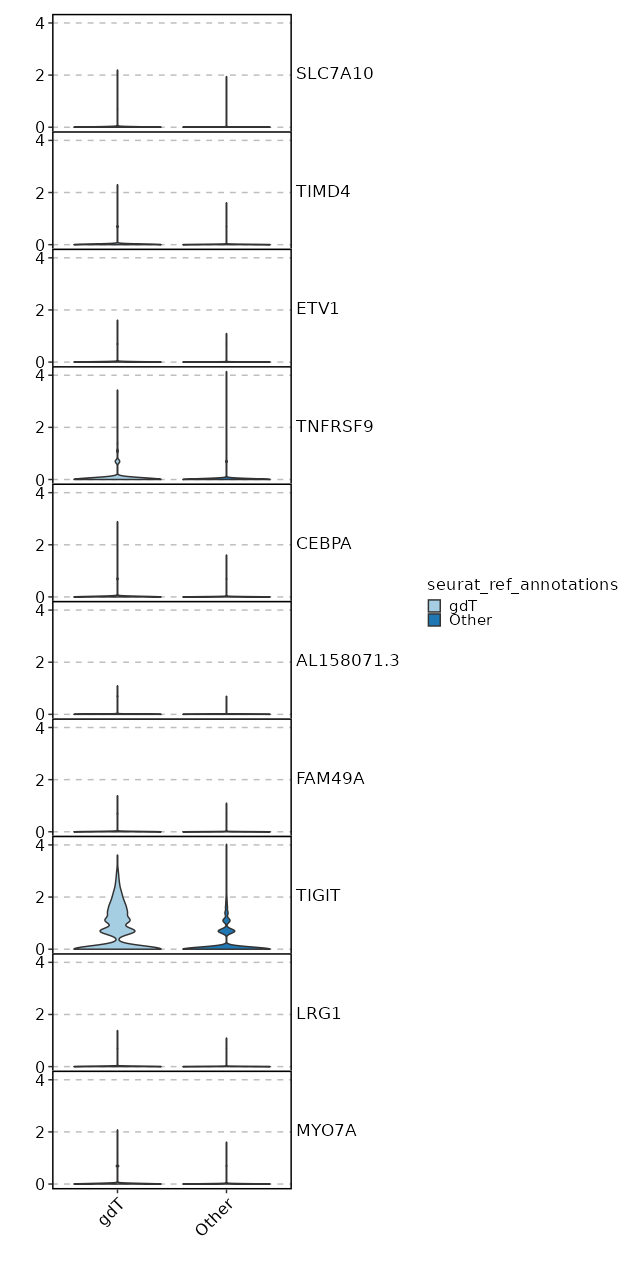
seurat_ref_annotations: gdT Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
0 1.6379 0.806 0.564 0 CD27 gdT 8.751e-263 1.7448 0.565 0.266 2.935e-258 TIGIT gdT 1.202e-179 0.5794 0.968 0.888 4.031e-175 CD2 gdT 4.188e-145 0.9839 0.495 0.252 1.405e-140 CD8B gdT 8.955e-144 0.9578 0.763 0.61 3.003e-139 CST7 gdT 3.744e-143 1.2272 0.586 0.38 1.256e-138 NKG7 gdT 4.191e-141 0.9843 0.521 0.296 1.406e-136 CD8A gdT 3.048e-137 1.3911 0.521 0.309 1.022e-132 LAG3 gdT 3.12e-132 1.1654 0.394 0.185 1.046e-127 GZMB gdT 1.265e-130 0.8386 0.55 0.334 4.244e-126 GZMA gdT 7.532e-122 1.0471 0.573 0.372 2.526e-117 PRF1 gdT 1.198e-119 1.0027 0.707 0.57 4.017e-115 CCL5 gdT 2.995e-112 0.5241 0.987 0.982 1.004e-107 PTPRCAP gdT 4.474e-112 0.7533 0.626 0.43 1.501e-107 CYTOR gdT 6.357e-110 1.2451 0.303 0.128 2.132e-105 GZMH gdT 2.896e-109 0.781 0.483 0.277 9.711e-105 MIR4435-2HG gdT 3.542e-109 0.5779 0.929 0.875 1.188e-104 CD74 gdT 9.175e-107 0.9097 0.467 0.263 3.077e-102 CCL4 gdT 1.372e-104 0.7677 0.436 0.236 4.603e-100 HLA-DRB1 gdT 4.235e-103 0.3911 0.999 0.999 1.42e-98 HLA-A gdT
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
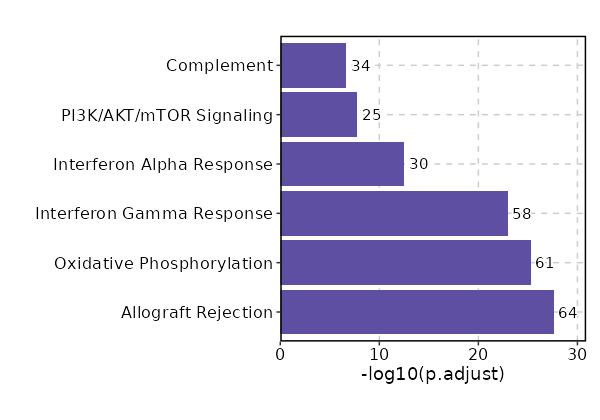
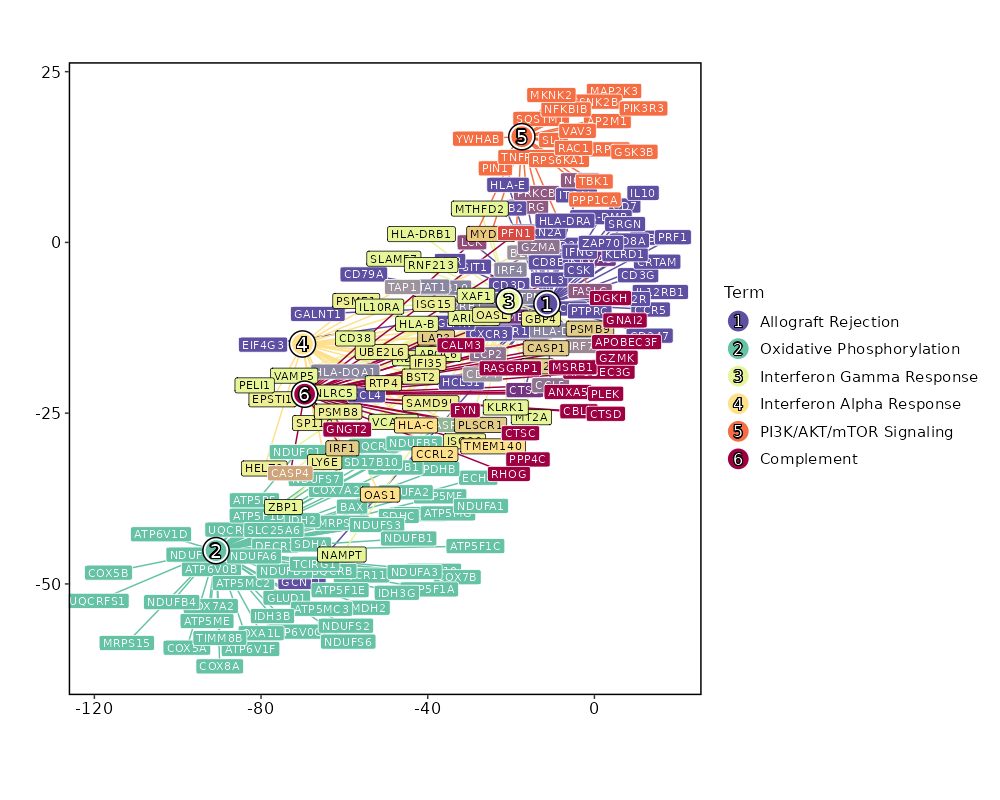
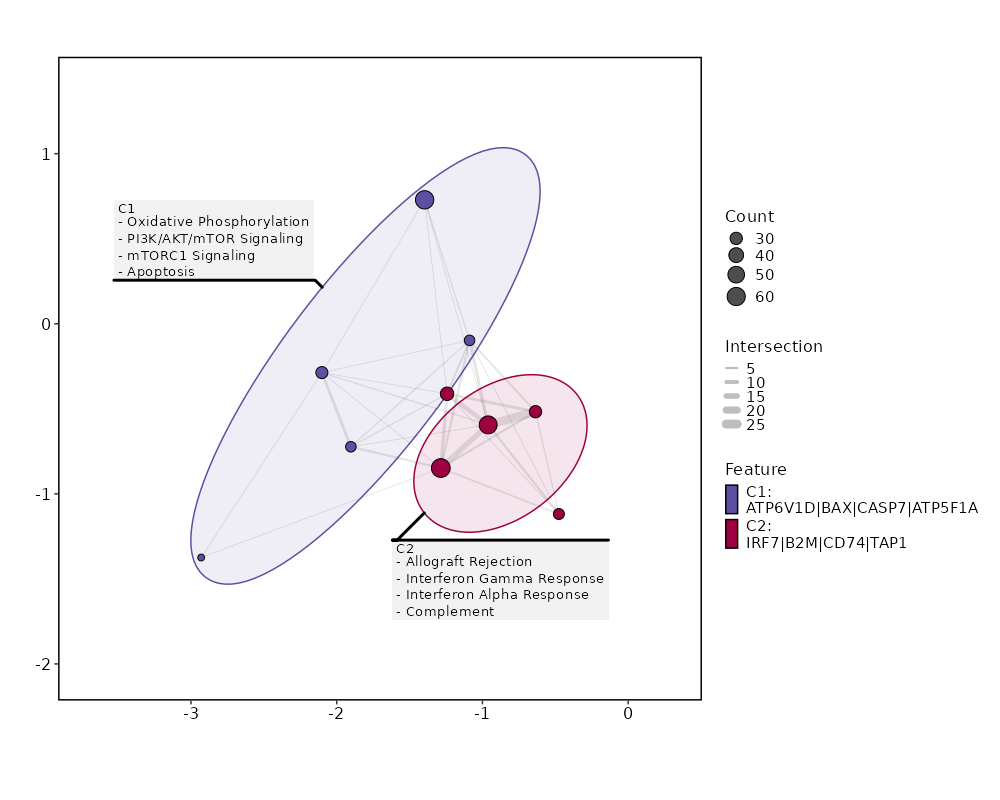
Enrichment Analysis Allograft Rejection 64/200 4.617e-30 2.216e-28 7.8191 528.1646 CD2;CD7;IL2RG;PSMB10;B2M;GBP2;FYB1;HLA-A;PRKCB;HLA-E;PRF1;CD79A;CCR1;IL10;PTPRC;CD3G;CD3E;CD3D;CTSS;CCR5;HLA-DQA1;MAP4K1;CDKN2A;STAT1;HCLS1;GCNT1;WAS;HLA-DMB;CXCR3;HLA-DMA;CD74;CRTAM;TAP1;KLRD1;PTPN6;IL12RB1;F2R;GLMN;CD8B;CD8A;EIF4G3;FASLG;SIT1;SRGN;GALNT1;IRF4;IFNG;BCL3;HLA-DRA;IRF7;LCP2;CD247;GZMB;GZMA;FGR;LCK;ZAP70;IL2RB;ITGB2;ITGAL;CCL5;CCL4;CSK;NCK1 1 MSigDB_Hallmark_2020 Oxidative Phosphorylation 61/200 2.025e-27 4.861e-26 7.2712 446.9163 ECH1;UQCRC2;BAX;PDHB;UQCR10;UQCR11;COX8A;COX7A2L;DECR1;ATP5F1D;ATP5F1E;GLUD1;ATP5ME;COX7B;MRPS15;MRPS12;MRPS11;ATP5MG;ATP5MF;ATP5PF;MDH2;COX6B1;COX5A;COX5B;ATP5F1A;ATP5F1C;SDHC;SDHA;ATP6V0C;ATP6V0B;IDH2;OXA1L;ATP6V1D;UQCRB;ATP5MC3;COX7A2;ATP5MC2;UQCRH;ATP6V1F;NDUFC1;NDUFS8;NDUFS7;NDUFS6;NDUFS3;NDUFS2;NDUFB5;NDUFB4;NDUFB3;NDUFB1;CASP7;UQCRFS1;TIMM8B;IDH3G;NDUFA6;NDUFA3;NDUFA2;NDUFA1;SLC25A6;TCIRG1;HSD17B10;IDH3B 2 MSigDB_Hallmark_2020 Interferon Gamma Response 58/200 7.143e-25 1.143e-23 6.7485 375.2049 SP110;PSMB9;LAP3;MTHFD2;RBCK1;UBE2L6;PSMB10;APOL6;SLAMF7;B2M;EPSTI1;GBP4;HLA-A;HLA-B;PLSCR1;PSME1;VAMP5;HLA-DRB1;IFI35;RNF213;HELZ2;MT2A;KLRK1;IL10RA;BST2;CD38;HLA-DQA1;STAT1;PELI1;PSMB8;NAMPT;HLA-DMA;ZBP1;CD74;TAP1;VCAM1;LY6E;PTPN6;SAMD9L;ARID5B;ISG15;ISG20;MYD88;XAF1;IRF1;IRF4;IRF2;IRF7;LCP2;NLRC5;CASP7;CASP4;CASP1;GZMA;IL2RB;RTP4;OASL;CCL5 3 MSigDB_Hallmark_2020 Interferon Alpha Response 30/97 0 0 7.2481 227.2948 CD74;SP110;TAP1;PSMB9;LAP3;LY6E;SAMD9L;UBE2L6;CCRL2;B2M;GBP2;EPSTI1;GBP4;HLA-C;ISG15;ISG20;PLSCR1;PSME1;IFI35;HELZ2;OAS1;BST2;IRF1;IRF2;IRF7;CASP1;RTP4;TMEM140;OASL;PSMB8 4 MSigDB_Hallmark_2020 PI3K/AKT/mTOR Signaling 25/105 0 0 5.0333 101.6196 PPP1CA;SQSTM1;PFN1;SLA;IL2RG;MAP2K3;PRKCB;MYD88;ARPC3;CSNK2B;PIN1;FASLG;TBK1;AP2M1;TNFRSF1A;VAV3;NFKBIB;GSK3B;YWHAB;PIK3R3;MKNK2;LCK;RPS6KA1;RAC1;NCK1 5 MSigDB_Hallmark_2020 Complement 34/200 0 0 3.3094 57.7294 PSMB9;LAP3;CALM3;PFN1;CBLB;APOBEC3F;APOBEC3G;PLSCR1;PPP4C;ANXA5;RHOG;CTSS;GNGT2;CTSD;CTSC;MSRB1;WAS;GNAI2;DGKH;RASGRP1;IRF1;IRF2;IRF7;LCP2;CASP7;CASP4;CASP1;GZMB;GZMA;GZMK;LCK;PLEK;CCL5;FYN 6 MSigDB_Hallmark_2020 mTORC1 Signaling 29/200 0 0.0001 2.7276 32.1481 MTHFD2;PSMC4;ARPC5L;GLRX;HPRT1;TFRC;TBK1;SLC7A5;CTSC;ELOVL5;TCEA1;NAMPT;G6PD;PSAT1;SQSTM1;SLA;SLC1A4;MAP2K3;SLC9A3R1;ATP6V1D;INSIG1;CORO1A;NFKBIB;GSK3B;PSMD13;PIK3R3;GAPDH;ITGB2;SEC11A 7 MSigDB_Hallmark_2020 Apoptosis 25/161 0 0.0001 2.9519 34.131 TAP1;CD2;BAX;SQSTM1;ROCK1;BCL2L11;F2R;ISG20;BCL2L1;PRF1;FASLG;SAT1;HMOX1;PMAIP1;GSTM1;CFLAR;TGFBR3;IRF1;BTG3;NEDD9;CASP7;CASP4;CD38;CASP1;SOD1 8 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 26/199 0.0001 0.0007 2.4107 21.5389 GBP4;PRKCH;GABARAPL1;PLSCR1;IL10;IL10RA;RHOH;CTLA4;GSTO1;CST7;DCPS;EOMES;BATF;CD81;ODC1;BCL2L1;TNFRSF9;TNFRSF4;TNFRSF1B;IRF4;ITGAE;SYNGR2;MYO1E;IL2RB;SYT11;GALM 9 MSigDB_Hallmark_2020 DNA Repair 21/150 0.0002 0.001 2.6062 21.9813 VPS28;MPG;NT5C;HPRT1;ARL6IP1;DGUOK;MPC2;POLR3GL;EDF1;ADA;SUPT4H1;ADRM1;TAF12;NUDT21;POLR1D;HCLS1;USP11;POLD4;GUK1;POLR2G;POLR2J 10 MSigDB_Hallmark_2020 Reactive Oxygen Species Pathway 10/49 0.0005 0.0023 4.0862 30.9252 G6PD;NDUFS2;NDUFB4;GLRX;NDUFA6;SOD1;LAMTOR5;LSP1;PRDX6;MBP 11 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 24/200 0.0008 0.0028 2.1832 15.5105 TNFSF9;DUSP4;DUSP2;ID2;BTG3;LITAF;SPSB1;NAMPT;TAP1;SQSTM1;CCRL2;MAP2K3;GEM;SAT1;TNFRSF9;CFLAR;TNIP1;IRF1;BCL3;TRIB1;CEBPD;PLEK;CCL5;CCL4 12 MSigDB_Hallmark_2020 Apical Junction 24/200 0.0008 0.0028 2.1832 15.5105 ACTN4;PFN1;MYL12B;FYB1;VASP;RHOF;PTPRC;CAP1;LIMA1;GNAI2;RAC2;VCAM1;ZYX;LAYN;ARPC2;INSIG1;ACTG1;CNN2;CD99;MSN;PECAM1;PIK3R3;EVL;MVD 12 MSigDB_Hallmark_2020 Inflammatory Response 24/200 0.0008 0.0028 2.1832 15.5105 TNFSF9;CHST2;IL10;IL10RA;RHOG;BST2;CXCR6;NAMPT;CD70;LY6E;CD82;CCRL2;RASGRP1;RGS1;TNFRSF9;TNFRSF1B;IRF1;IRF7;LCP2;ADRM1;LCK;IL2RB;RTP4;CCL5 12 MSigDB_Hallmark_2020 UV Response Up 19/158 0.0027 0.0085 2.1834 12.9383 KLHDC3;STARD3;TAP1;SQSTM1;BCL2L11;BSG;AP2S1;RAB27A;HLA-F;TFRC;RASGRP1;HMOX1;ATP6V1F;MARK2;IRF1;TMBIM6;BTG3;CLTB;BAK1 15 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 12/87 0.0052 0.0157 2.5493 13.3946 CCR1;ITGA4;TNFRSF1A;TNFRSF1B;IRF1;IL2RG;CD38;IL12RB1;STAT1;MYD88;HMOX1;BAK1 16 MSigDB_Hallmark_2020 Fatty Acid Metabolism 18/158 0.006 0.017 2.0519 10.4881 ACOT8;ECH1;SDHC;SDHA;HPGD;ODC1;PDHB;UBE2L6;GABARAPL1;PSME1;PRDX6;DECR1;ELOVL5;IDH3G;MDH2;HSD17B10;PCBD1;IDH3B 17 MSigDB_Hallmark_2020 Adipogenesis 21/200 0.0081 0.0194 1.8732 9.0217 ECH1;UQCR10;UQCR11;STOM;COX8A;SCP2;DECR1;CCNG2;COX7B;MDH2;SQOR;ATP1B3;SDHC;UCP2;DNAJC15;REEP5;PIM3;COQ5;NDUFS3;IDH3G;SOD1 18 MSigDB_Hallmark_2020 Estrogen Response Late 21/200 0.0081 0.0194 1.8732 9.0217 FDFT1;HPRT1;TSTA3;DUSP2;SLC7A5;ID2;BTG3;CLIC3;ELOVL5;FABP5;JAK1;BATF;PTPN6;SLC27A2;SLC1A4;IDH2;SLC9A3R1;ISG20;SLC2A8;DNAJC1;ATP2B4 18 MSigDB_Hallmark_2020 p53 Pathway 21/200 0.0081 0.0194 1.8732 9.0217 BAX;TNFSF9;STOM;HMOX1;ADA;CTSD;CDKN2A;TAP1;DEF6;CD82;CD81;F2R;MXD4;SAT1;VDR;MKNK2;CASP1;IP6K2;RALGDS;CEBPA;BAK1 18 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
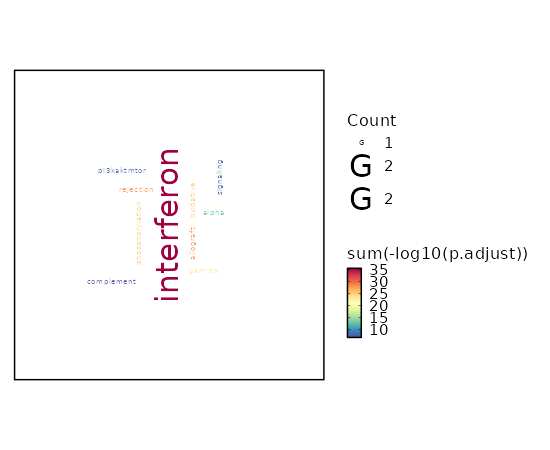
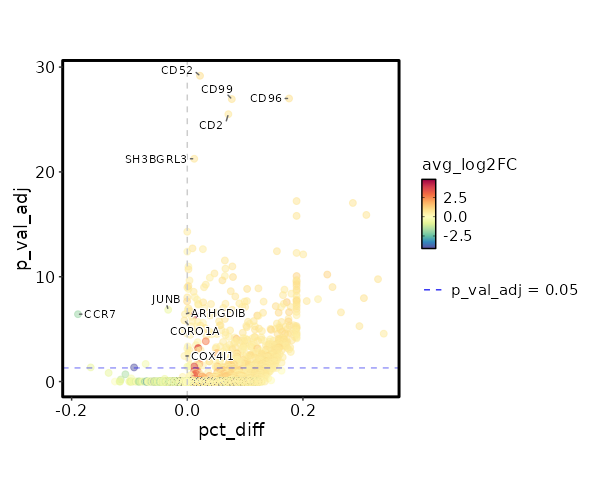
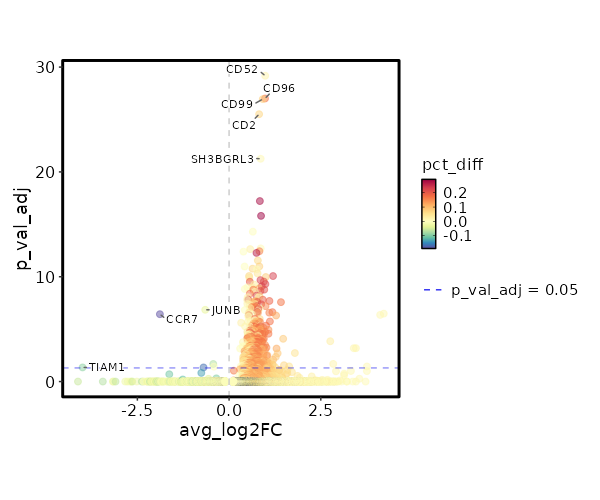
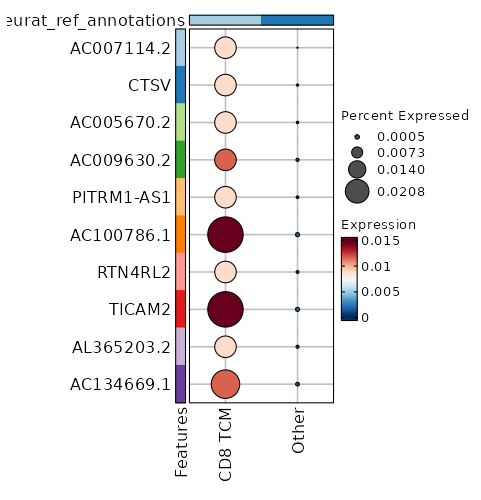
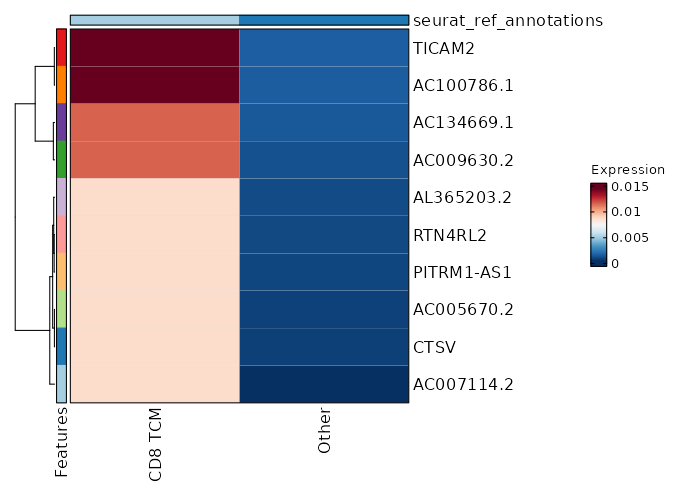
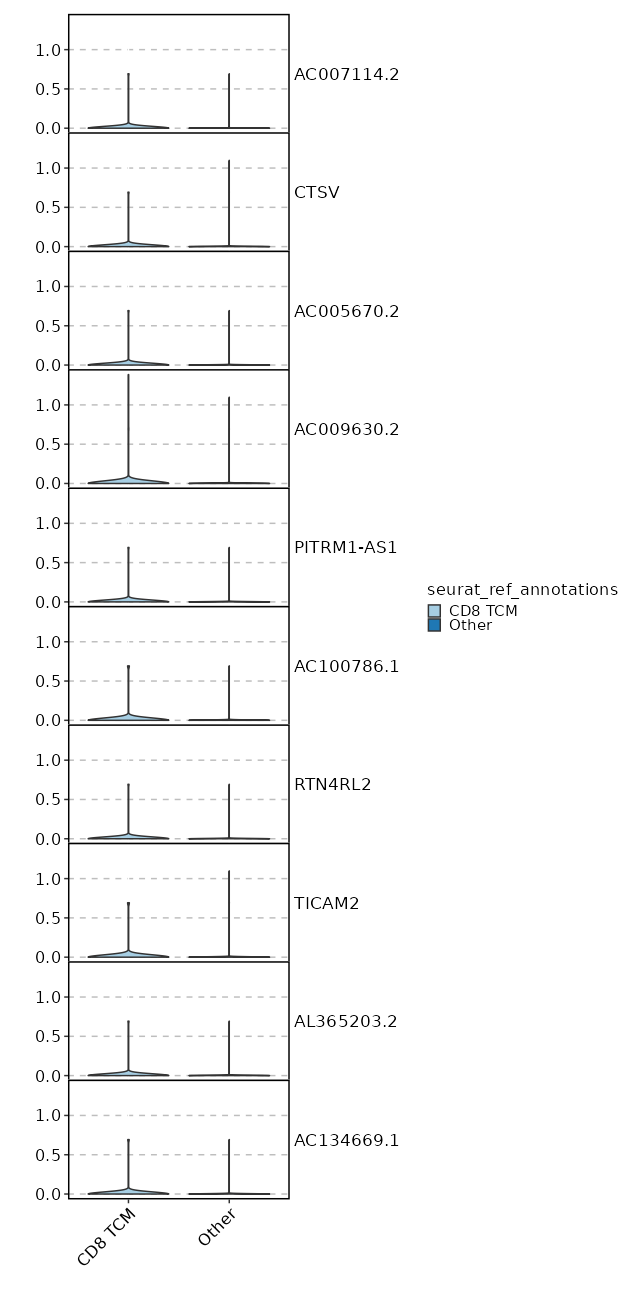
seurat_ref_annotations: CD8 TCM Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
2.006e-34 0.9848 0.996 0.974 6.726e-30 CD52 CD8 TCM 2.941e-32 0.9801 0.904 0.728 9.863e-28 CD96 CD8 TCM 3.362e-32 0.919 0.975 0.898 1.128e-27 CD99 CD8 TCM 9.292e-31 0.8179 0.996 0.925 3.116e-26 CD2 CD8 TCM 1.66e-26 0.8608 0.996 0.984 5.569e-22 SH3BGRL3 CD8 TCM 1.795e-22 0.8391 0.912 0.63 6.021e-18 CCL5 CD8 TCM 4.661e-21 0.871 0.746 0.453 1.563e-16 LINC01871 CD8 TCM 1.507e-19 0.6479 1 1 0 TMSB4X CD8 TCM 5.872e-18 0.8611 1 0.991 0 GAPDH CD8 TCM 6.927e-18 0.5747 0.983 0.956 0 EVL CD8 TCM 1.096e-17 0.8271 0.883 0.728 0 OSTF1 CD8 TCM 1.2e-17 0.3922 1 1 0 B2M CD8 TCM 1.595e-17 0.7468 0.696 0.432 0 GZMA CD8 TCM 8.366e-17 0.7843 0.975 0.91 0 CLIC1 CD8 TCM 3.091e-16 0.8245 0.954 0.876 0 RARRES3 CD8 TCM 3.172e-16 0.4219 1 0.998 0 CD3E CD8 TCM 5.071e-16 0.6394 0.971 0.905 0 LSP1 CD8 TCM 5.859e-16 0.7548 1 0.998 0 PFN1 CD8 TCM 0 0.7773 0.996 0.949 0 ARPC2 CD8 TCM 0 0.5455 0.95 0.885 0 HCST CD8 TCM
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
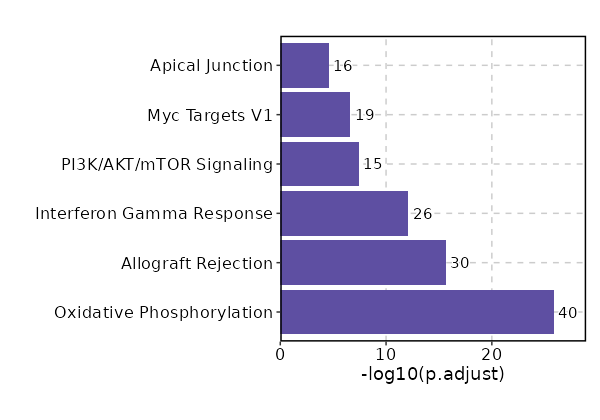
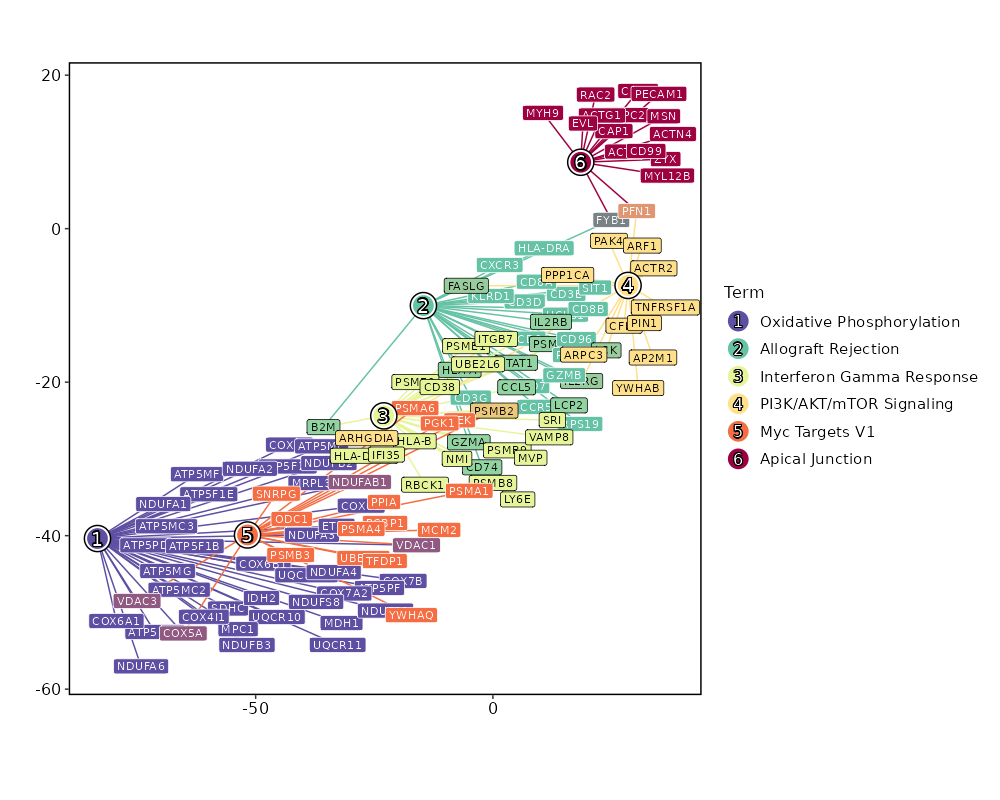
Enrichment Analysis Oxidative Phosphorylation 40/200 2.925e-28 1.375e-26 13.3116 843.9482 UQCR10;UQCR11;COX8A;ATP5F1D;ATP5F1E;COX6C;ATP5ME;COX7B;MRPL34;ATP5MG;ATP5MF;ATP5PF;ATP5PD;COX6B1;NDUFAB1;COX5A;ATP5F1B;ATP5F1C;MDH1;SDHC;COX6A1;IDH2;ATP5MC3;COX7A2;ATP5MC2;MPC1;NDUFS8;NDUFB3;NDUFB2;NDUFB1;UQCRFS1;COX4I1;NDUFA6;NDUFA4;NDUFA3;NDUFA2;NDUFA1;VDAC3;VDAC1;ETFB 1 MSigDB_Hallmark_2020 Allograft Rejection 30/200 9.532e-18 2.24e-16 9.1412 358.2596 CD2;CD7;IL2RG;PSMB10;B2M;FYB1;HLA-A;PRF1;RPS19;CD3G;CD3E;CD3D;CCR5;STAT1;HCLS1;CXCR3;CD74;KLRD1;CD96;CD8B;CD8A;FASLG;SIT1;HLA-DRA;LCP2;GZMB;GZMA;LCK;IL2RB;CCL5 2 MSigDB_Hallmark_2020 Interferon Gamma Response 26/200 0 0 7.657 234.3947 PSMB9;RBCK1;UBE2L6;PSMB10;B2M;HLA-A;HLA-B;VAMP8;PSME2;PSME1;HLA-DRB1;IFI35;NMI;CD38;STAT1;PSMB8;PSMB2;CD74;LY6E;LCP2;GZMA;IL2RB;MVP;SRI;CCL5;ITGB7 3 MSigDB_Hallmark_2020 PI3K/AKT/mTOR Signaling 15/105 0 0 8.3355 162.9586 PPP1CA;PFN1;IL2RG;CFL1;PAK4;ARPC3;PIN1;FASLG;AP2M1;ACTR2;TNFRSF1A;YWHAB;ARF1;LCK;ARHGDIA 4 MSigDB_Hallmark_2020 Myc Targets V1 19/200 0 0 5.2796 91.9622 DEK;UBE2L3;PCBP1;PSMA6;PSMA4;NDUFAB1;PSMA1;COX5A;PSMB3;PSMB2;PPIA;SNRPG;ODC1;TFDP1;MCM2;YWHAQ;PGK1;VDAC3;VDAC1 5 MSigDB_Hallmark_2020 Apical Junction 16/200 0 0 4.3391 54.6577 ACTN4;PFN1;MYL12B;ACTB;FYB1;CAP1;RAC2;ZYX;ARPC2;ACTG1;CNN2;CD99;MSN;MYH9;PECAM1;EVL 6 MSigDB_Hallmark_2020 Interferon Alpha Response 11/97 0 0 6.3333 78.2854 CD74;PSMB9;LY6E;UBE2L6;B2M;HLA-C;PSME2;PSME1;IFI35;NMI;PSMB8 7 MSigDB_Hallmark_2020 Complement 15/200 0 0.0001 4.0353 44.8651 PSMB9;PFN1;LGALS3;PPP4C;ANXA5;CTSV;CTSD;CTSC;GNB2;LCP2;GZMB;GZMA;LCK;CCL5;FYN 8 MSigDB_Hallmark_2020 mTORC1 Signaling 15/200 0 0.0001 4.0353 44.8651 PPA1;TPI1;SLC7A5;ALDOA;SSR1;CTSC;PSMA4;PPIA;SLC9A3R1;CORO1A;ACTR2;MCM2;PGK1;RRM2;GAPDH 8 MSigDB_Hallmark_2020 Adipogenesis 12/200 0.0008 0.0038 3.152 22.4321 UQCR10;UQCR11;STOM;COX8A;ALDOA;COX7B;NDUFAB1;SDHC;PDCD4;COX6A1;REEP5;ETFB 10 MSigDB_Hallmark_2020 Protein Secretion 7/96 0.0034 0.0139 3.8547 21.9424 COPE;ARF1;AP2S1;TMED2;CTSC;GNAS;AP2M1 11 MSigDB_Hallmark_2020 DNA Repair 9/150 0.0035 0.0139 3.1357 17.6906 RBX1;ADA;SUPT4H1;TMED2;DAD1;HCLS1;GUK1;POLR2E;POLR2G 12 MSigDB_Hallmark_2020 Hypoxia 9/200 0.021 0.0706 2.3089 8.915 MT1E;PRDX5;TPI1;ALDOA;TPST2;S100A4;MYH9;PGK1;GAPDH 13 MSigDB_Hallmark_2020 Inflammatory Response 9/200 0.021 0.0706 2.3089 8.915 NMI;CXCR6;LY6E;RGS1;LCP2;LCK;IL2RB;SRI;CCL5 13 MSigDB_Hallmark_2020 TGF-beta Signaling 4/54 0.0237 0.0741 3.8993 14.5979 RHOA;PPP1CA;ID2;FKBP1A 15 MSigDB_Hallmark_2020 Fatty Acid Metabolism 7/158 0.0422 0.1236 2.2648 7.1692 SDHC;ODC1;UBE2L6;PSME1;LGALS1;ALDOA;MDH1 16 MSigDB_Hallmark_2020 G2-M Checkpoint 8/200 0.0514 0.1236 2.0364 6.0441 EZH2;JPT1;SLC7A5;ODC1;HMGN2;TFDP1;TOP1;MCM2 17 MSigDB_Hallmark_2020 Estrogen Response Late 8/200 0.0514 0.1236 2.0364 6.0441 COX6C;SLC7A5;ID2;CLIC3;PDCD4;IDH2;SLC9A3R1;ETFB 17 MSigDB_Hallmark_2020 Glycolysis 8/200 0.0514 0.1236 2.0364 6.0441 RBCK1;CHST12;TPI1;ALDOA;MDH1;SDHC;PPIA;PGK1 17 MSigDB_Hallmark_2020 Androgen Response 5/100 0.0526 0.1236 2.5658 7.5556 B2M;TMEM50A;MYL12A;SRP19;DBI 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
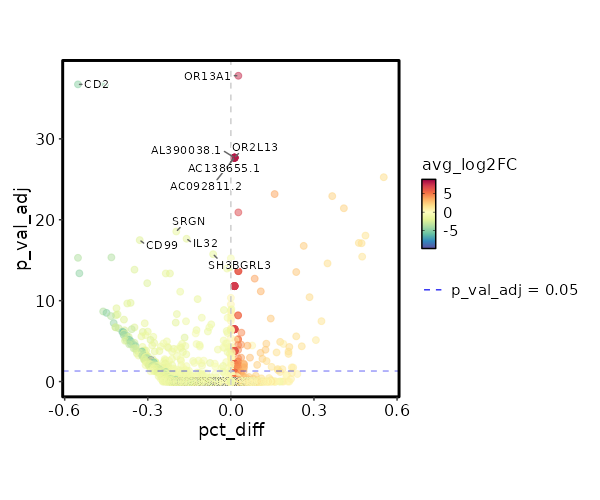
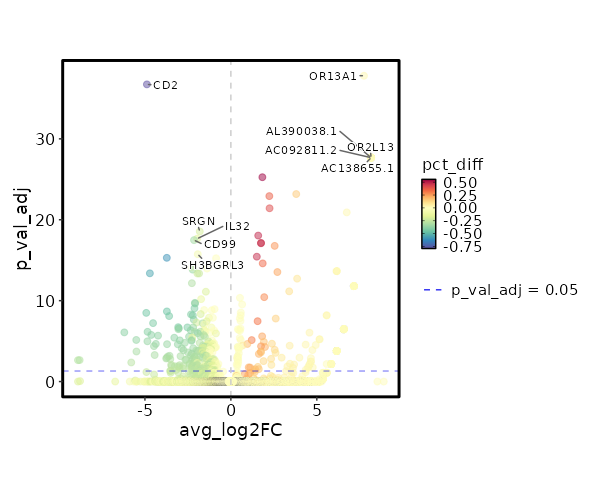
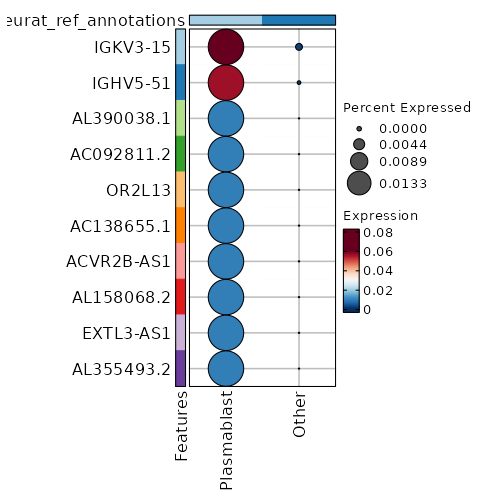
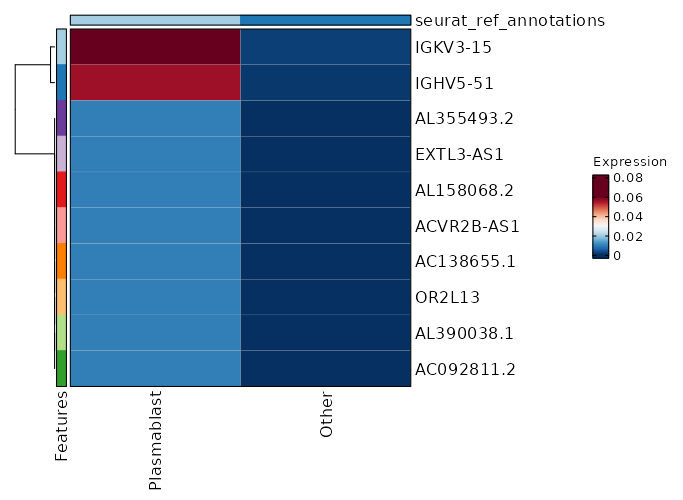
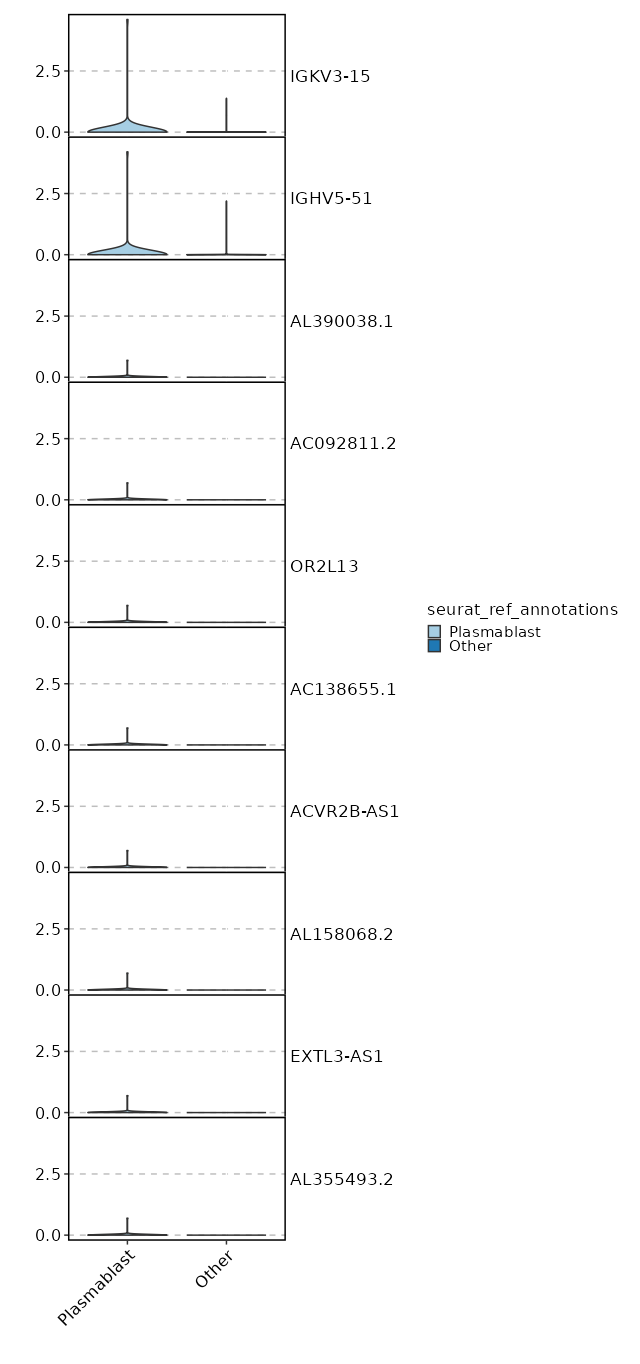
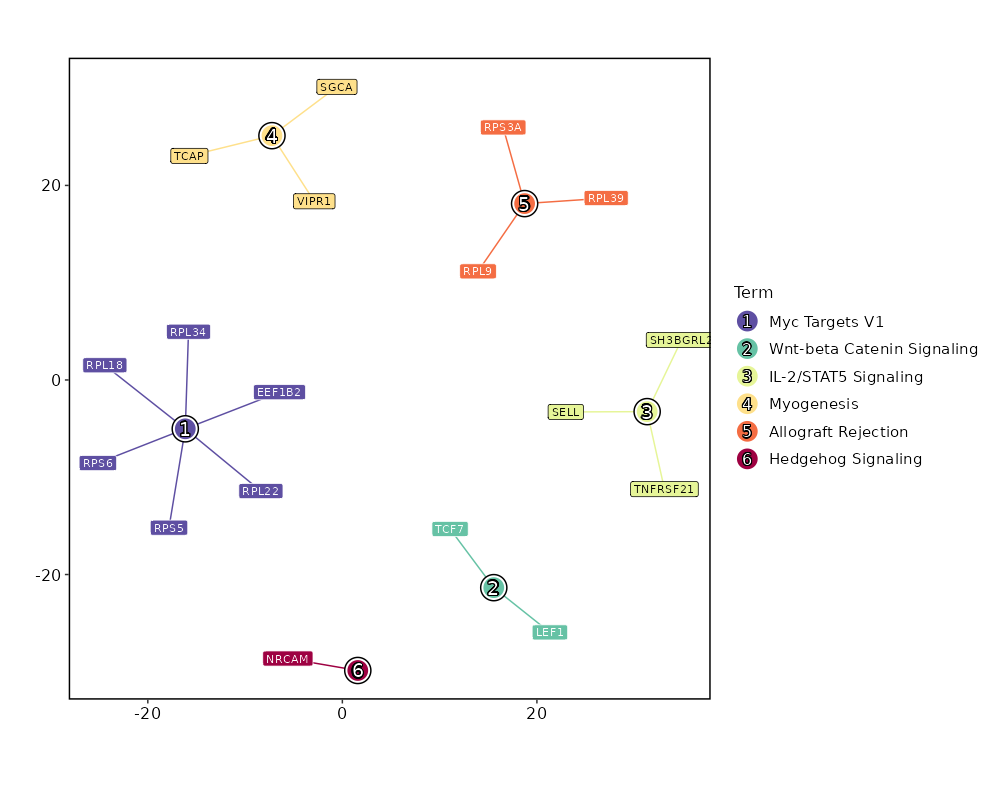
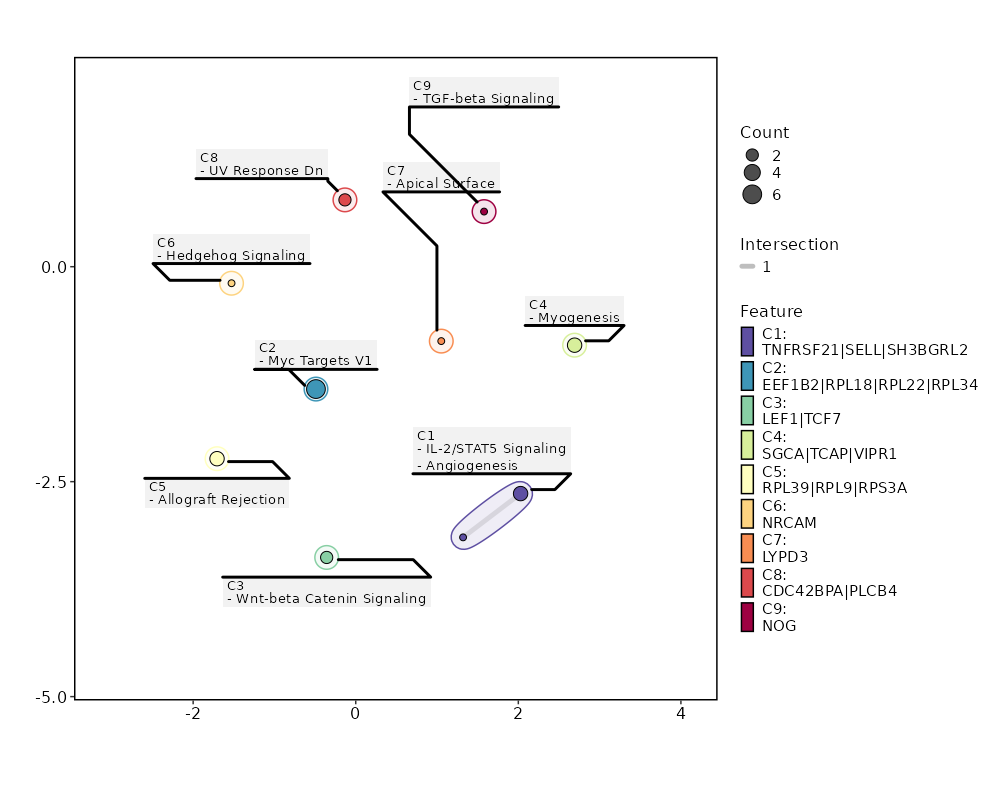
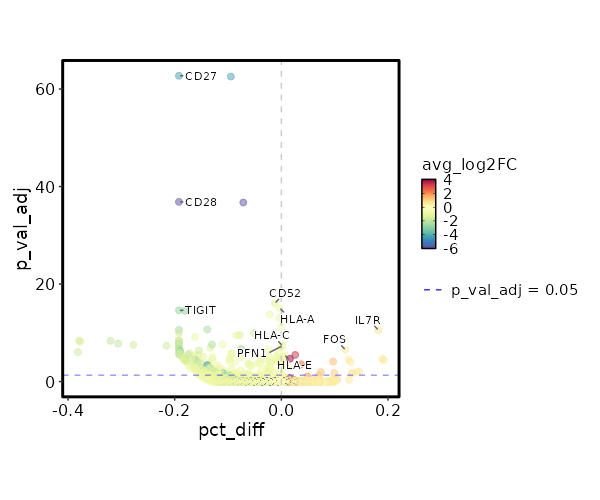
seurat_ref_annotations: Plasmablast Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
4.77e-43 7.7436 0.027 0 1.6e-38 OR13A1 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 AL390038.1 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 AC092811.2 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 OR2L13 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 AC138655.1 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 ACVR2B-AS1 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 AL158068.2 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 EXTL3-AS1 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 AL355493.2 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 SGCA Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 CARD14 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 PLCB4 Plasmablast 6.328e-33 8.1587 0.013 0 2.122e-28 WFDC2 Plasmablast 1.647e-30 1.8326 0.787 0.235 5.524e-26 CCR7 Plasmablast 1.989e-28 3.8032 0.173 0.015 6.672e-24 AK5 Plasmablast 3.572e-28 2.2376 0.453 0.087 1.198e-23 ACTN1 Plasmablast 1.134e-26 2.2511 0.533 0.125 3.804e-22 S1PR1 Plasmablast 3.669e-26 6.7436 0.027 0 1.23e-21 SMARCA1 Plasmablast 2.699e-23 1.5905 0.72 0.234 9.051e-19 SELL Plasmablast 2.191e-22 1.7539 0.733 0.271 7.349e-18 LEF1 Plasmablast
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
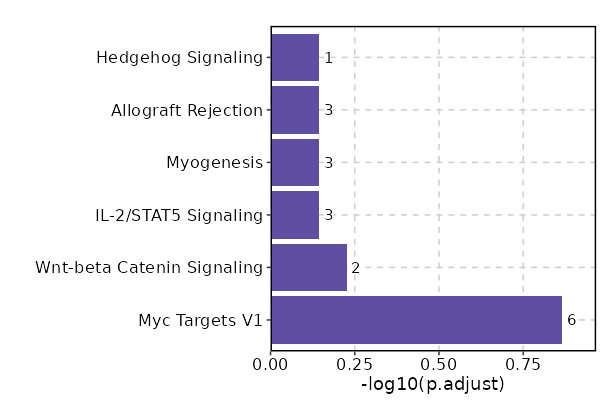
Enrichment Analysis Myc Targets V1 6/200 0.0049 0.1367 4.0515 21.5637 RPL22;RPL18;RPL34;RPS6;RPS5;EEF1B2 1 MSigDB_Hallmark_2020 Wnt-beta Catenin Signaling 2/42 0.0425 0.5946 6.4299 20.3116 TCF7;LEF1 2 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 3/199 0.2033 0.7161 1.9656 3.131 SH3BGRL2;SELL;TNFRSF21 3 MSigDB_Hallmark_2020 Myogenesis 3/200 0.2053 0.7161 1.9555 3.0958 VIPR1;TCAP;SGCA 4 MSigDB_Hallmark_2020 Allograft Rejection 3/200 0.2053 0.7161 1.9555 3.0958 RPL9;RPL39;RPS3A 4 MSigDB_Hallmark_2020 Hedgehog Signaling 1/36 0.2458 0.7161 3.6514 5.1232 NRCAM 6 MSigDB_Hallmark_2020 Angiogenesis 1/36 0.2458 0.7161 3.6514 5.1232 TNFRSF21 6 MSigDB_Hallmark_2020 Apical Surface 1/44 0.2917 0.7161 2.9709 3.66 LYPD3 8 MSigDB_Hallmark_2020 UV Response Dn 2/144 0.3098 0.7161 1.8019 2.1116 CDC42BPA;PLCB4 9 MSigDB_Hallmark_2020 TGF-beta Signaling 1/54 0.3452 0.7161 2.4091 2.5625 NOG 10 MSigDB_Hallmark_2020 Apoptosis 2/161 0.3582 0.7161 1.6079 1.6509 LEF1;BCL2L10 11 MSigDB_Hallmark_2020 Mitotic Spindle 2/199 0.4611 0.7161 1.2952 1.0027 ARHGEF11;CDC42BPA 12 MSigDB_Hallmark_2020 Estrogen Response Early 2/200 0.4637 0.7161 1.2886 0.9904 MPPED2;SYNGR1 13 MSigDB_Hallmark_2020 Apical Junction 2/200 0.4637 0.7161 1.2886 0.9904 ACTN1;LDLRAP1 13 MSigDB_Hallmark_2020 Inflammatory Response 2/200 0.4637 0.7161 1.2886 0.9904 CCR7;SELL 13 MSigDB_Hallmark_2020 p53 Pathway 2/200 0.4637 0.7161 1.2886 0.9904 RPL18;RPS12 13 MSigDB_Hallmark_2020 KRAS Signaling Dn 2/200 0.4637 0.7161 1.2886 0.9904 LYPD3;EDAR 13 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.4948 0.7161 1.4822 1.043 TNFRSF21 18 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.533 0.7161 1.3272 0.8351 SELL 19 MSigDB_Hallmark_2020 Androgen Response 1/100 0.5439 0.7161 1.2867 0.7836 ACTN1 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
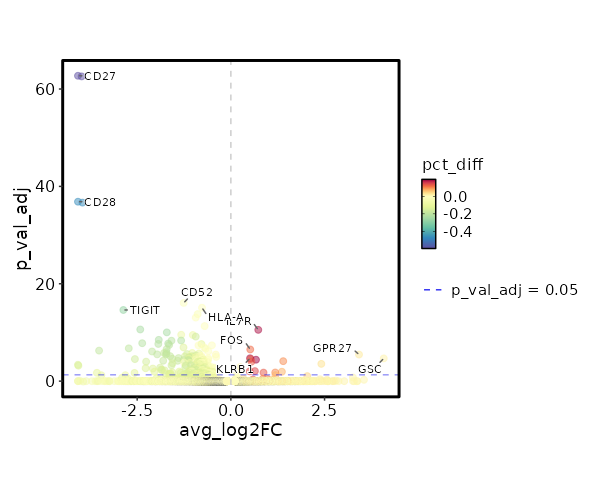
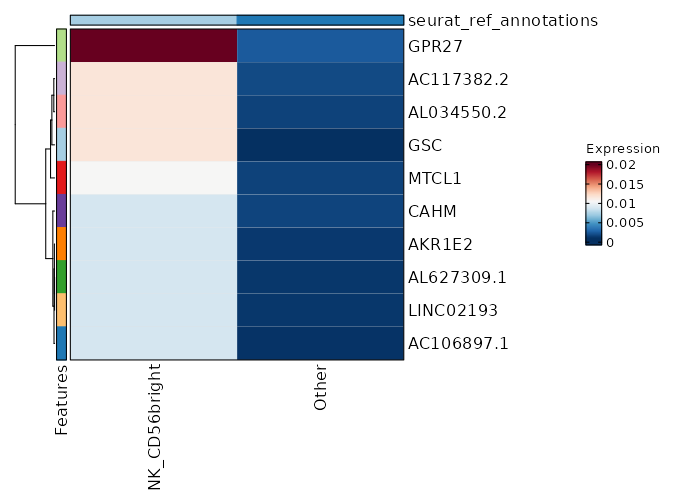
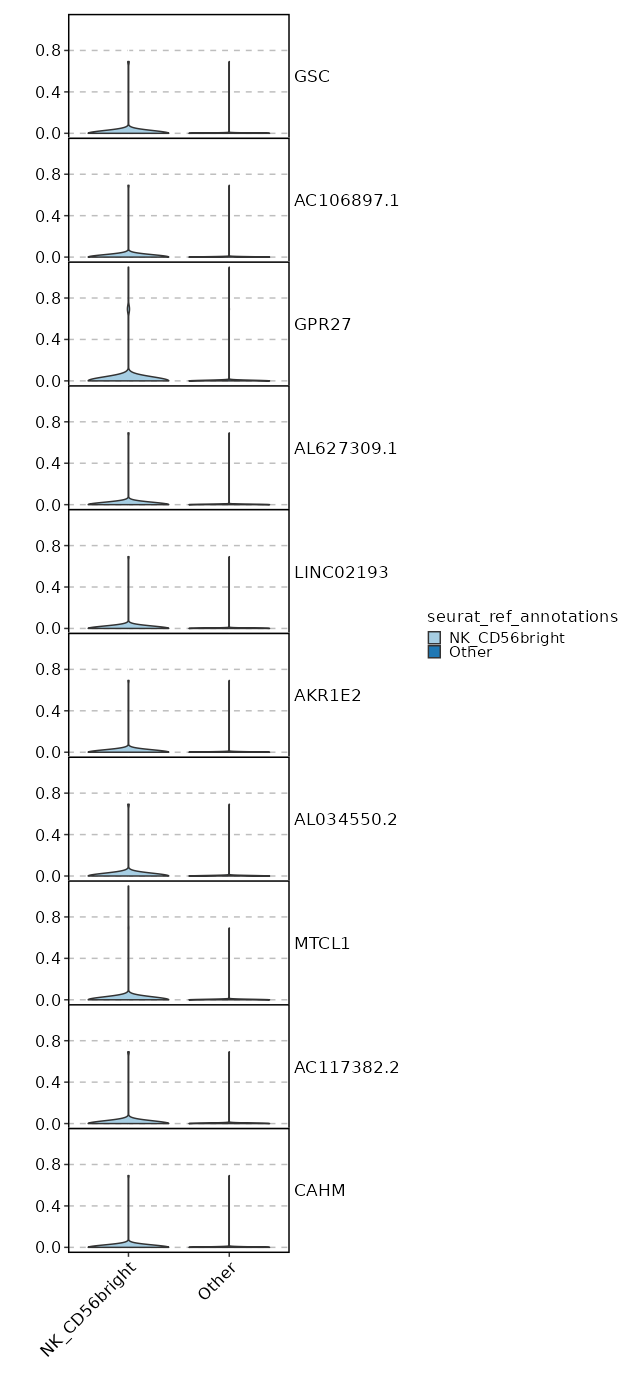
seurat_ref_annotations: NK_CD56bright Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
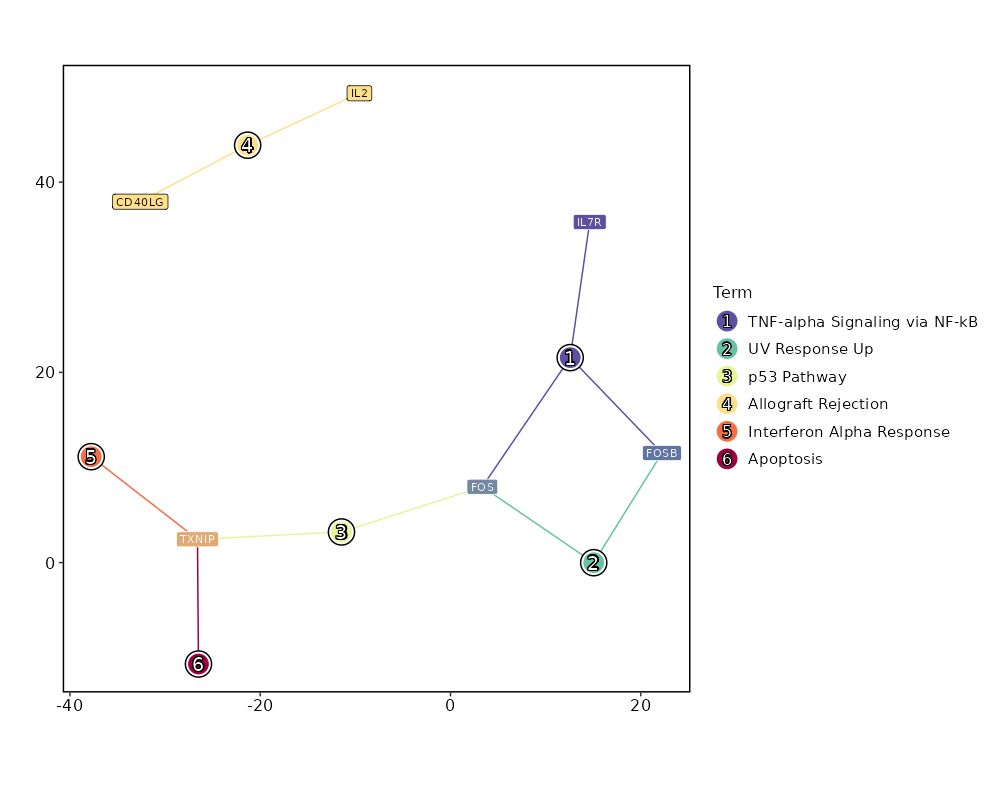
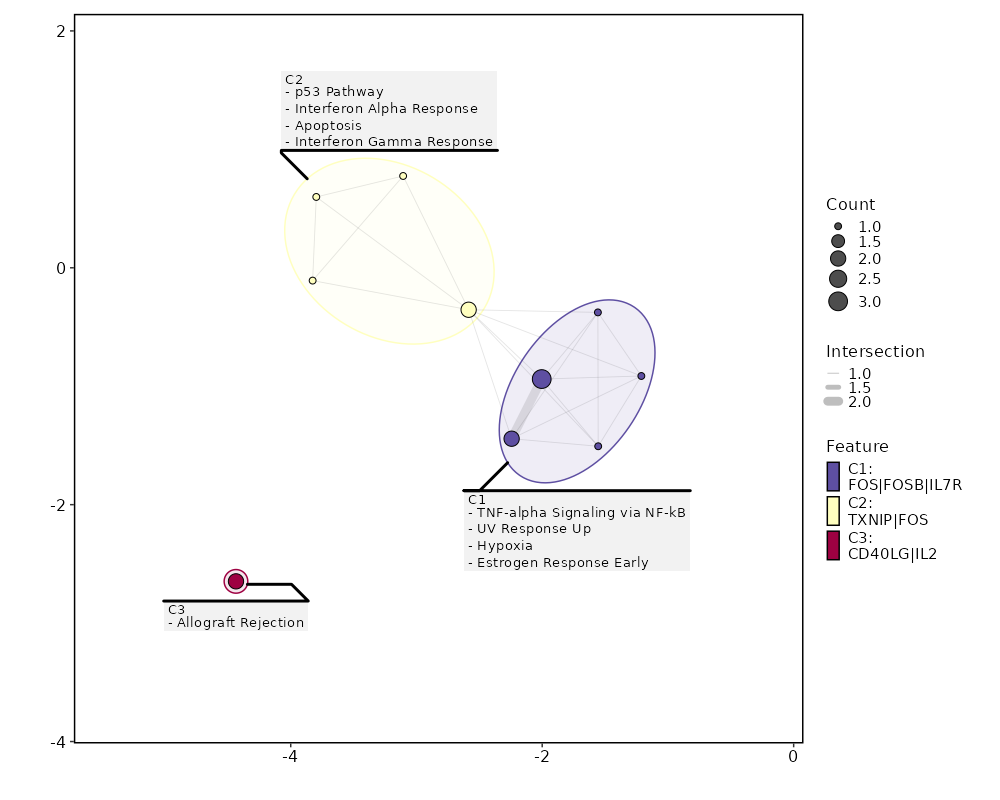
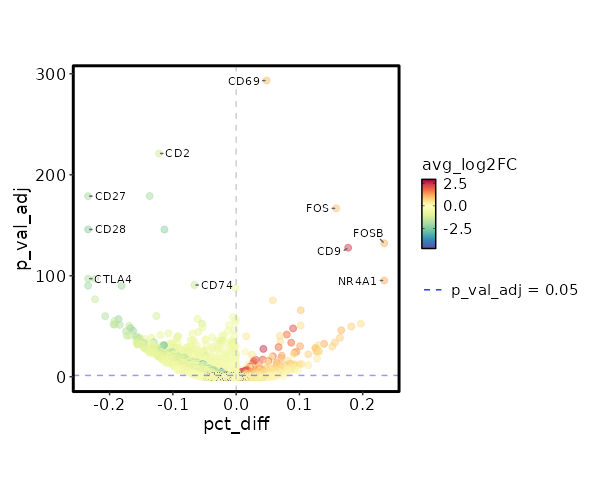
8.719e-16 0.7312 0.842 0.66 0 IL7R NK_CD56bright 0 0.5174 0.958 0.838 0 FOS NK_CD56bright 0 3.4187 0.029 0.003 0 GPR27 NK_CD56bright 0 0.5116 0.758 0.569 0 KLRB1 NK_CD56bright 0 4.0797 0.017 0.001 0 GSC NK_CD56bright 0 0.5283 0.875 0.749 0 TXNIP NK_CD56bright 0 0.6703 0.546 0.354 0 CD40LG NK_CD56bright 0 1.3972 0.162 0.065 0.0001 TTC39C-AS1 NK_CD56bright 0 0.5489 0.254 0.124 0.0001 IL2 NK_CD56bright 0 2.4138 0.046 0.009 0.0003 TRIQK NK_CD56bright 0 0.2507 0.996 0.984 0.0013 EEF1B2 NK_CD56bright 0 0.6414 0.367 0.221 0.0087 CCR6 NK_CD56bright 0 0.5011 0.771 0.63 0.0098 FOSB NK_CD56bright 0 1.3613 0.125 0.051 0.0117 TBXAS1 NK_CD56bright 0 1.1835 0.204 0.105 0.0165 TNFSF13B NK_CD56bright 0 0.8715 0.321 0.189 0.0176 AC020571.1 NK_CD56bright 0 0.1608 1 1 0.0373 EEF1A1 NK_CD56bright
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
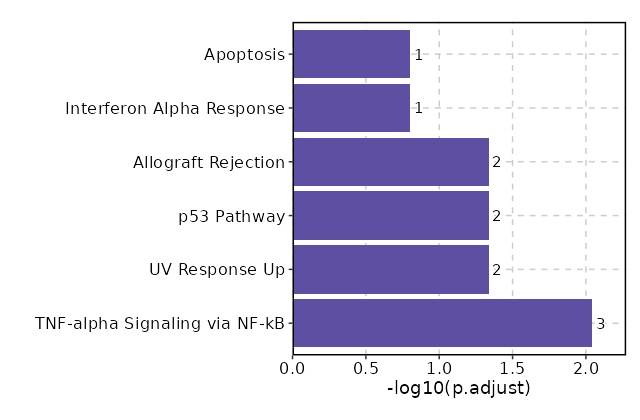
Enrichment Analysis TNF-alpha Signaling via NF-kB 3/200 0.0006 0.0091 21.5221 159.5186 FOS;FOSB;IL7R 1 MSigDB_Hallmark_2020 UV Response Up 2/158 0.0078 0.046 16.9462 82.2448 FOS;FOSB 2 MSigDB_Hallmark_2020 p53 Pathway 2/200 0.0123 0.046 13.3232 58.6413 FOS;TXNIP 3 MSigDB_Hallmark_2020 Allograft Rejection 2/200 0.0123 0.046 13.3232 58.6413 CD40LG;IL2 3 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.0794 0.1571 12.9473 32.8056 TXNIP 5 MSigDB_Hallmark_2020 Apoptosis 1/161 0.1284 0.1571 7.7434 15.8923 TXNIP 6 MSigDB_Hallmark_2020 Hypoxia 1/200 0.1571 0.1571 6.2136 11.4999 FOS 7 MSigDB_Hallmark_2020 Estrogen Response Early 1/200 0.1571 0.1571 6.2136 11.4999 FOS 7 MSigDB_Hallmark_2020 Estrogen Response Late 1/200 0.1571 0.1571 6.2136 11.4999 FOS 7 MSigDB_Hallmark_2020 Interferon Gamma Response 1/200 0.1571 0.1571 6.2136 11.4999 TXNIP 7 MSigDB_Hallmark_2020 Complement 1/200 0.1571 0.1571 6.2136 11.4999 CD40LG 7 MSigDB_Hallmark_2020 Myc Targets V1 1/200 0.1571 0.1571 6.2136 11.4999 EEF1B2 7 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.1571 0.1571 6.2136 11.4999 IL7R 7 MSigDB_Hallmark_2020 KRAS Signaling Up 1/200 0.1571 0.1571 6.2136 11.4999 IL7R 7 MSigDB_Hallmark_2020 KRAS Signaling Dn 1/200 0.1571 0.1571 6.2136 11.4999 CD40LG 7 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
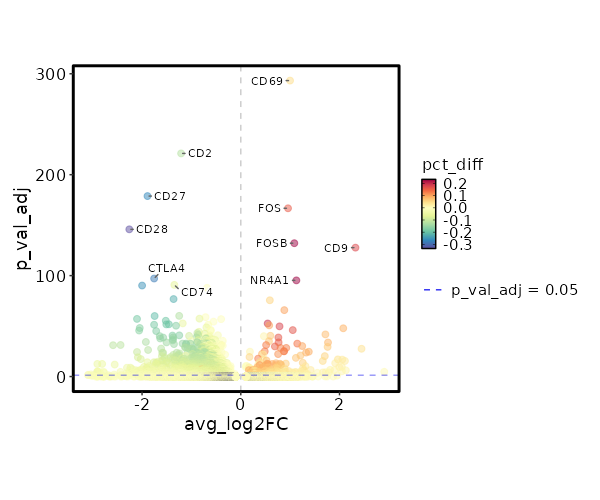
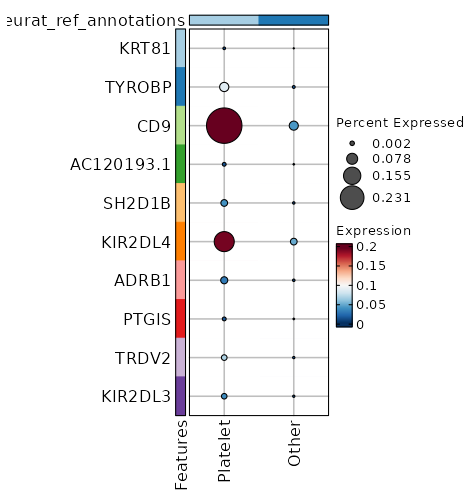
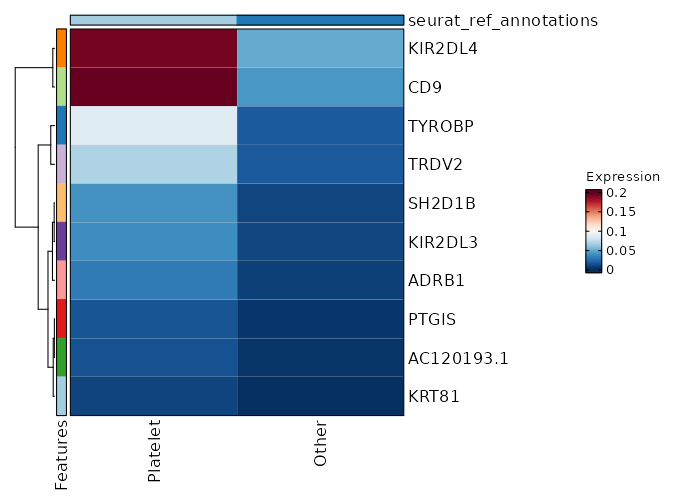
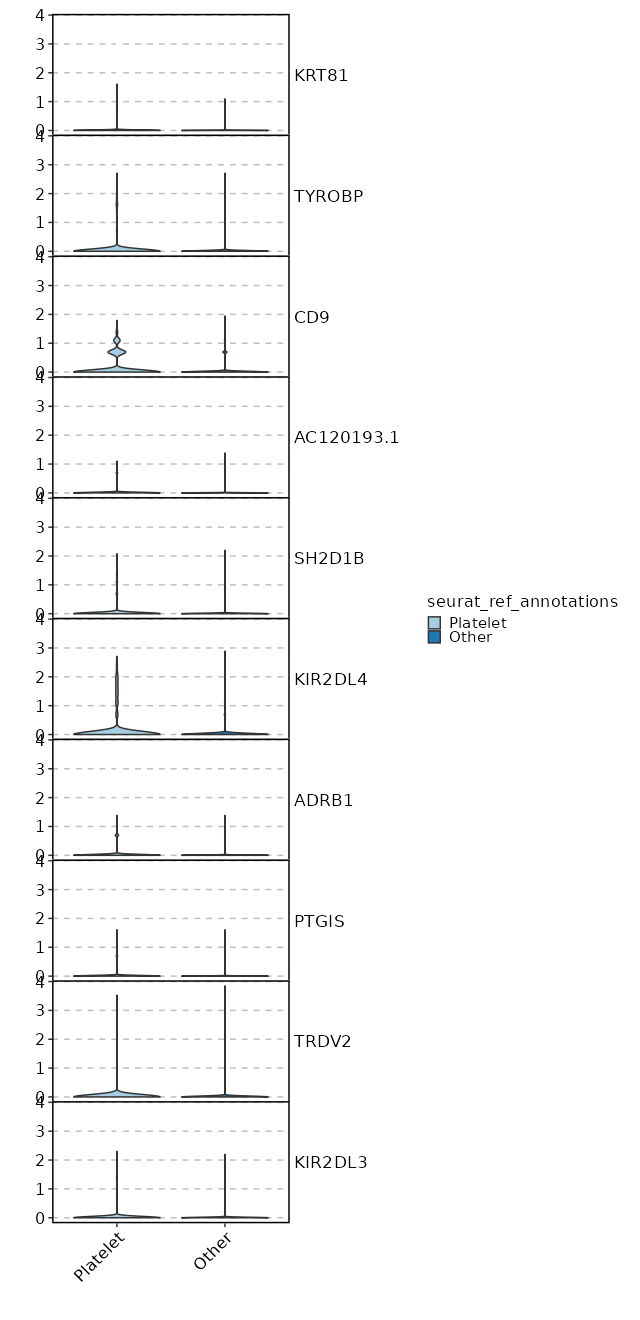
seurat_ref_annotations: Platelet Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
1.709e-298 0.998 0.999 0.951 5.733e-294 CD69 Platelet 5.689e-172 0.9566 0.975 0.817 1.908e-167 FOS Platelet 2.263e-137 1.0844 0.832 0.598 7.59e-133 FOSB Platelet 5.784e-133 2.3228 0.231 0.054 1.94e-128 CD9 Platelet 1.532e-100 1.126 0.706 0.472 5.14e-96 NR4A1 Platelet 7.717e-81 0.5883 0.983 0.925 2.588e-76 DUSP1 Platelet 4.57e-71 0.8816 0.89 0.788 1.533e-66 JUN Platelet 8.212e-58 0.5443 0.831 0.634 2.754e-53 IL7R Platelet 5.918e-56 0.5751 0.857 0.755 1.985e-51 PPP1R15A Platelet 5.636e-55 0.7848 0.661 0.479 1.89e-50 MYADM Platelet 4.438e-53 2.0756 0.127 0.037 1.488e-48 KIR2DL4 Platelet 2.837e-51 1.0527 0.397 0.231 9.513e-47 EGR1 Platelet 5.24e-47 1.7141 0.112 0.032 1.758e-42 TRDC Platelet 1.176e-45 0.5996 0.893 0.824 3.943e-41 BTG2 Platelet 3.035e-45 0.468 0.975 0.959 1.018e-40 JUNB Platelet 6.071e-44 0.7617 0.569 0.405 2.036e-39 AC020916.1 Platelet 2.896e-40 0.5709 0.893 0.823 9.714e-36 NR4A2 Platelet 3.736e-39 0.766 0.422 0.266 1.253e-34 CAPG Platelet 5.555e-39 1.7665 0.143 0.056 1.863e-34 KLRC2 Platelet 7.281e-38 1.1436 0.342 0.203 2.442e-33 AREG Platelet
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
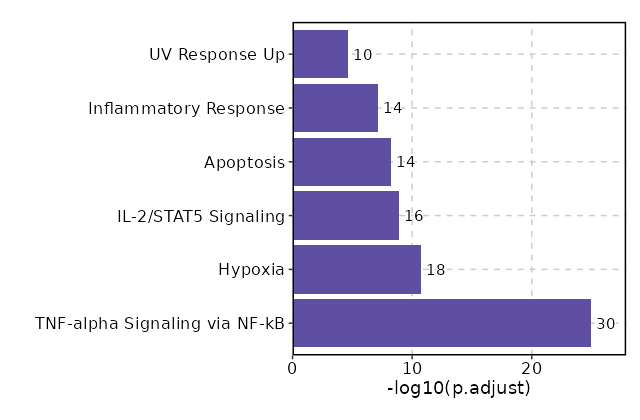
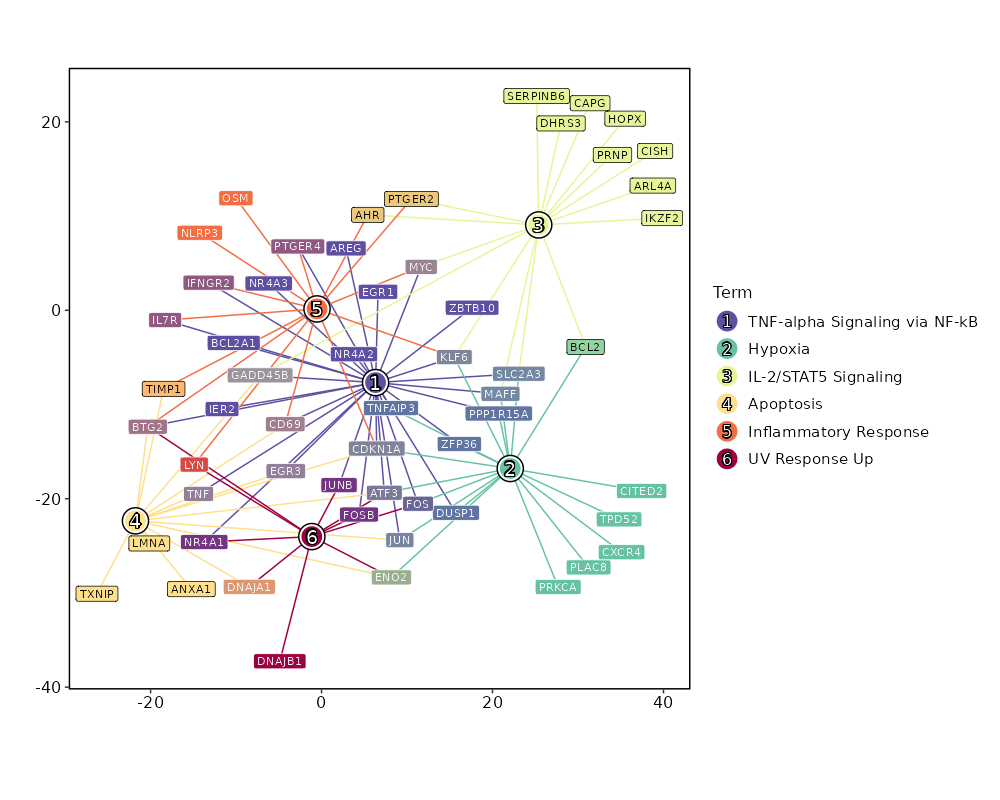
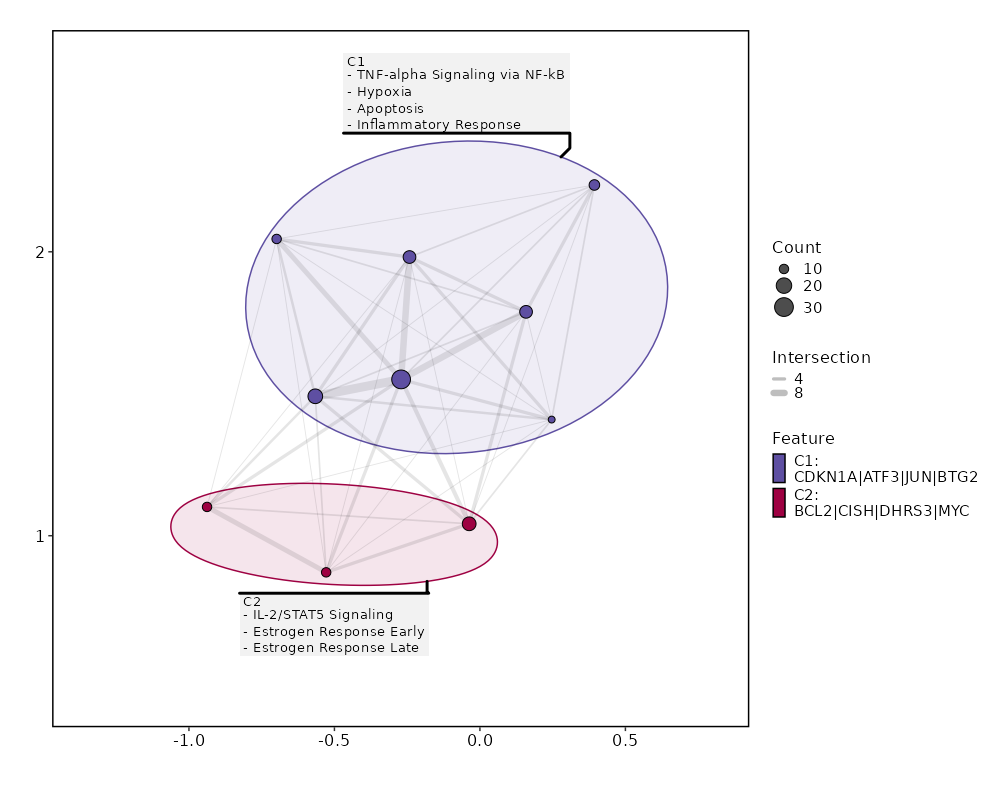
Enrichment Analysis TNF-alpha Signaling via NF-kB 30/200 2.497e-27 1.124e-25 21.5261 1318.5782 ATF3;MYC;EGR1;EGR3;GADD45B;IER2;JUNB;JUN;DUSP1;BTG2;AREG;IFNGR2;FOS;FOSB;CDKN1A;PTGER4;ZBTB10;CD69;PPP1R15A;TNF;ZFP36;MAFF;NR4A3;NR4A2;IL7R;SLC2A3;NR4A1;BCL2A1;TNFAIP3;KLF6 1 MSigDB_Hallmark_2020 Hypoxia 18/200 0 0 11.2204 312.5682 ATF3;PRKCA;ENO2;PLAC8;JUN;DUSP1;CITED2;TPD52;FOS;CDKN1A;CXCR4;PPP1R15A;ZFP36;MAFF;SLC2A3;BCL2;TNFAIP3;KLF6 2 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 16/199 0 0 9.8053 227.2341 MYC;GADD45B;IKZF2;DHRS3;SERPINB6;PTGER2;CAPG;AHR;CISH;PRNP;HOPX;MAFF;SLC2A3;BCL2;ARL4A;KLF6 3 MSigDB_Hallmark_2020 Apoptosis 14/161 0 0 10.5795 226.979 CD69;ATF3;TNF;LMNA;EGR3;GADD45B;ENO2;ANXA1;JUN;BTG2;TIMP1;DNAJA1;TXNIP;CDKN1A 4 MSigDB_Hallmark_2020 Inflammatory Response 14/200 0 0 8.3446 155.425 MYC;LYN;BTG2;TIMP1;IFNGR2;CDKN1A;PTGER2;PTGER4;OSM;CD69;AHR;NLRP3;IL7R;KLF6 5 MSigDB_Hallmark_2020 UV Response Up 10/158 0 0 7.3395 93.5006 ATF3;ENO2;NR4A1;JUNB;LYN;BTG2;DNAJB1;FOS;DNAJA1;FOSB 6 MSigDB_Hallmark_2020 Allograft Rejection 11/200 0 0 6.3439 79.4183 RPL9;LYN;NCR1;TIMP1;TPD52;IFNGR2;CD40LG;CAPG;TNF;NLRP3;IL2 7 MSigDB_Hallmark_2020 Estrogen Response Early 10/200 0 0.0001 5.7049 60.8446 RETREG1;MYC;EGR3;MYBL1;AREG;FOS;DHRS3;CISH;BCL2;DYNLT3 8 MSigDB_Hallmark_2020 Estrogen Response Late 10/200 0 0.0001 5.7049 60.8446 CD9;EGR3;AREG;FOS;ZFP36;PERP;CISH;BCL2;DYNLT3;LSR 8 MSigDB_Hallmark_2020 Epithelial Mesenchymal Transition 9/200 0.0001 0.0006 5.0792 45.2786 VIM;GADD45B;ENO2;JUN;AREG;TIMP1;CAPG;TNFAIP3;THBS1 10 MSigDB_Hallmark_2020 p53 Pathway 8/200 0.0007 0.0026 4.4665 32.4943 ATF3;JUN;BTG2;FOS;CDKN1A;PPP1R15A;PERP;TXNIP 11 MSigDB_Hallmark_2020 KRAS Signaling Up 8/200 0.0007 0.0026 4.4665 32.4943 SPON1;LAT2;FCER1G;CXCR4;PPP1R15A;IL7R;TNFAIP3;SPRY2 11 MSigDB_Hallmark_2020 Apical Surface 4/44 0.0008 0.0028 10.5717 75.2314 LYN;GSTM3;CD160;MAL 13 MSigDB_Hallmark_2020 TGF-beta Signaling 4/54 0.0018 0.0056 8.453 53.6325 PPP1R15A;IFNGR2;THBS1;JUNB 14 MSigDB_Hallmark_2020 Interferon Gamma Response 7/200 0.0032 0.0089 3.8666 22.2618 CDKN1A;CD69;AUTS2;XCL1;ARL4A;TNFAIP3;TXNIP 15 MSigDB_Hallmark_2020 mTORC1 Signaling 7/200 0.0032 0.0089 3.8666 22.2618 CD9;SKAP2;BTG2;CDKN1A;CXCR4;PPP1R15A;SLC2A3 15 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 4/87 0.0097 0.0257 5.0837 23.57 JUN;CD9;TNF;IFNGR2 17 MSigDB_Hallmark_2020 Coagulation 5/138 0.0105 0.0258 3.9769 18.1117 CD9;MAFF;ANXA1;TIMP1;THBS1 18 MSigDB_Hallmark_2020 Reactive Oxygen Species Pathway 3/49 0.0114 0.0258 6.8558 30.646 FES;PRNP;JUNB 19 MSigDB_Hallmark_2020 UV Response Dn 5/144 0.0125 0.0258 3.8041 16.6759 MYC;PRKCA;DUSP1;MGLL;CITED2 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
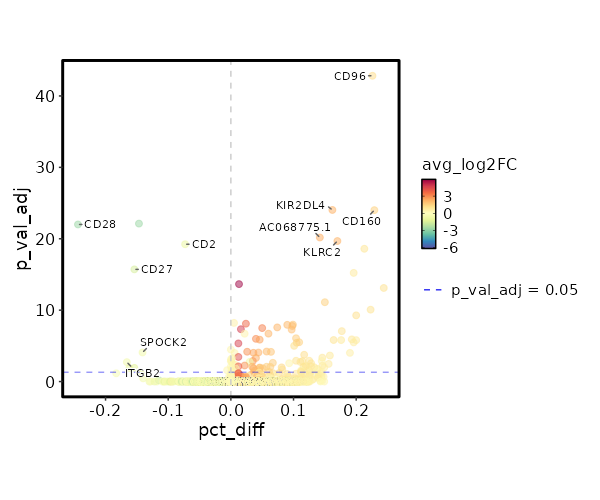
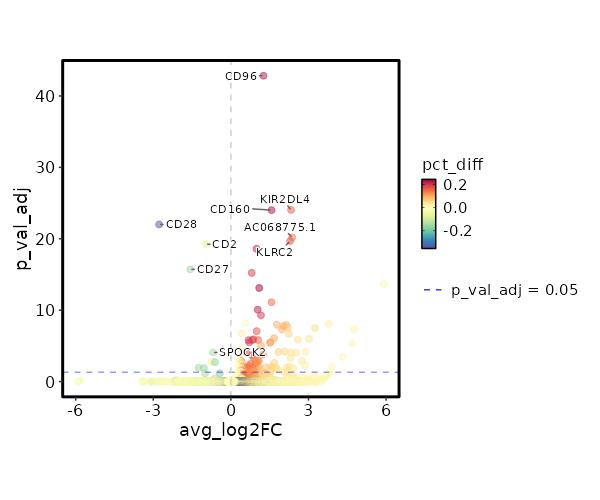
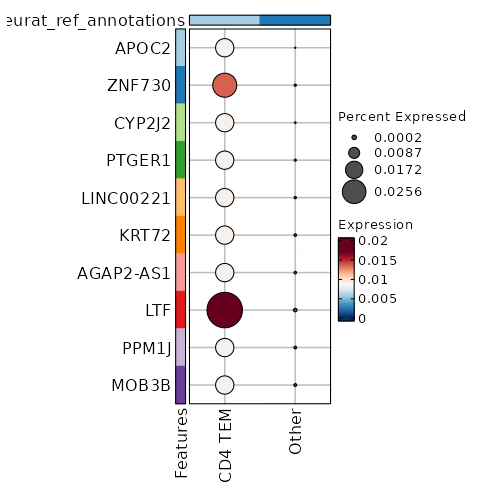
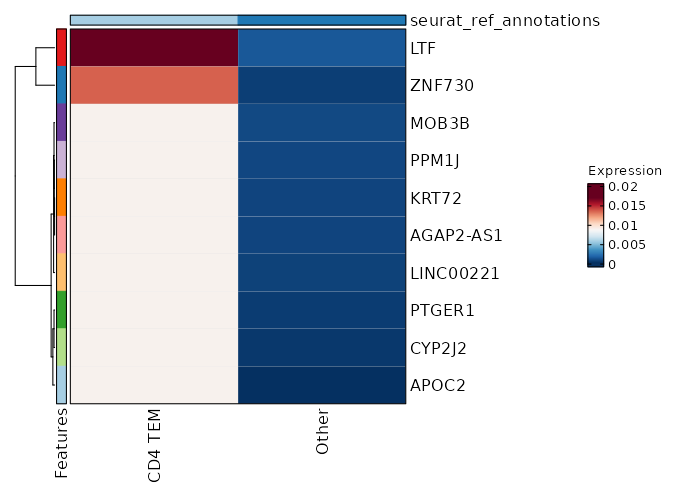
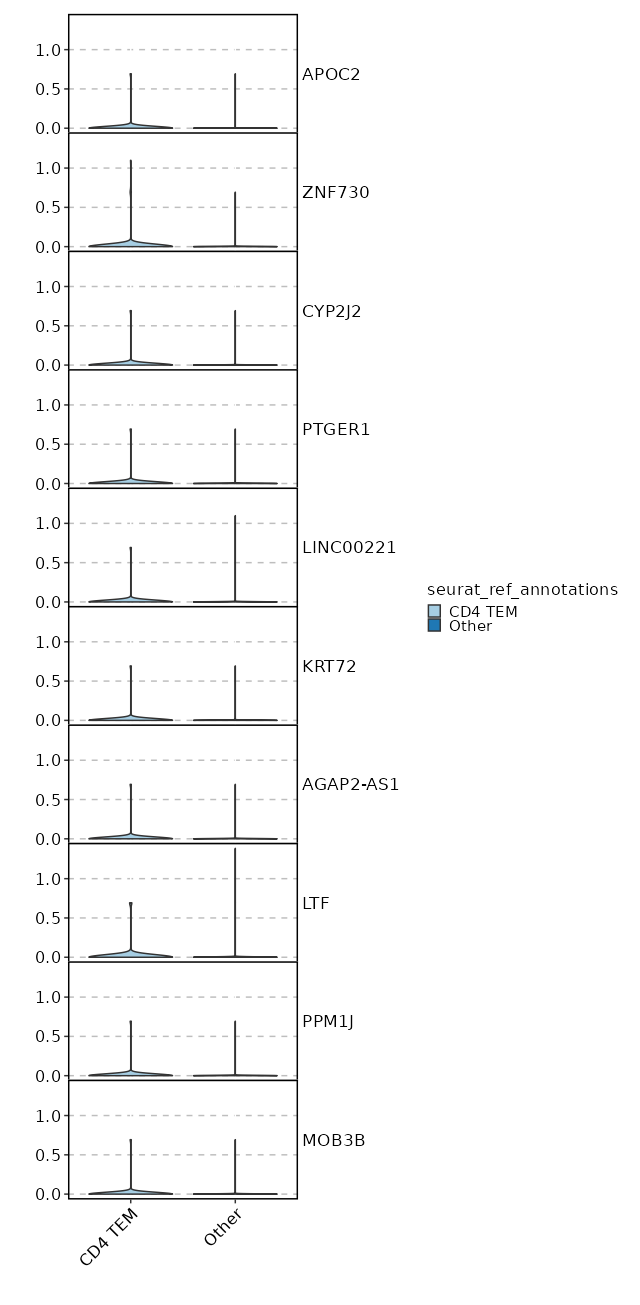
seurat_ref_annotations: CD4 TEM Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
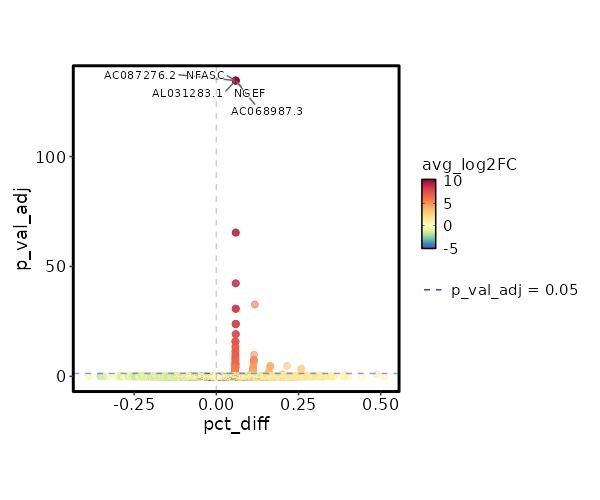
4.429e-48 1.2618 0.953 0.727 1.485e-43 CD96 CD4 TEM 2.76e-29 2.328 0.209 0.047 9.257e-25 KIR2DL4 CD4 TEM 2.924e-29 1.5741 0.329 0.1 9.807e-25 CD160 CD4 TEM 1.982e-25 2.3689 0.184 0.042 6.647e-21 AC068775.1 CD4 TEM 6.195e-25 2.2894 0.235 0.065 2.078e-20 KLRC2 CD4 TEM 7.537e-24 0.9922 0.325 0.112 2.528e-19 XCL2 CD4 TEM 1.799e-20 0.8058 0.303 0.107 6.032e-16 XCL1 CD4 TEM 6.702e-19 5.9106 0.013 0 0 APOC2 CD4 TEM 2.322e-18 1.0954 0.615 0.371 0 HOPX CD4 TEM 2.302e-16 1.5717 0.231 0.081 0 KLRC3 CD4 TEM 0 1.0375 0.56 0.337 0 ITGA1 CD4 TEM 0 1.1626 0.419 0.219 0 TMIGD2 CD4 TEM 0 0.5497 0.996 0.991 0 FTL CD4 TEM 0 3.7793 0.026 0.002 0 LTF CD4 TEM 0 1.7753 0.141 0.042 0 TRDC CD4 TEM 0 2.138 0.124 0.034 0 FES CD4 TEM 0 2.0406 0.141 0.043 0 TRDV1 CD4 TEM 0 2.1996 0.098 0.024 0 SPECC1 CD4 TEM 0 3.2525 0.06 0.01 0 TRDV2 CD4 TEM 0 4.7586 0.017 0.001 0 ZNF730 CD4 TEM
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
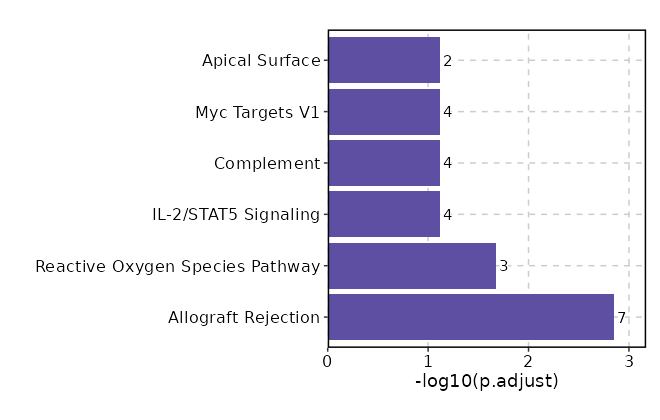
Enrichment Analysis Allograft Rejection 7/200 0.0001 0.0014 7.7695 75.3214 RPS19;LYN;NCR1;CRTAM;KLRD1;CD96;CCL5 1 MSigDB_Hallmark_2020 Reactive Oxygen Species Pathway 3/49 0.0018 0.0211 13.4885 84.9673 FES;FTL;HHEX 2 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 4/199 0.0172 0.0769 4.255 17.2881 MAP3K8;HOPX;ITGAE;ABCB1 3 MSigDB_Hallmark_2020 Complement 4/200 0.0175 0.0769 4.2331 17.1292 LYN;FCER1G;LTF;CCL5 4 MSigDB_Hallmark_2020 Myc Targets V1 4/200 0.0175 0.0769 4.2331 17.1292 PABPC1;HNRNPA2B1;RPS6;BUB3 4 MSigDB_Hallmark_2020 Apical Surface 2/44 0.02 0.0769 9.7491 38.1151 LYN;CD160 6 MSigDB_Hallmark_2020 Apoptosis 3/161 0.0459 0.1507 3.9049 12.0353 PDCD4;NEDD9;DFFA 7 MSigDB_Hallmark_2020 Mitotic Spindle 3/199 0.0762 0.1773 3.1417 8.0885 ARHGEF7;NEDD9;TBCD 8 MSigDB_Hallmark_2020 Inflammatory Response 3/200 0.0771 0.1773 3.1256 8.0106 LYN;NMUR1;CCL5 9 MSigDB_Hallmark_2020 KRAS Signaling Up 3/200 0.0771 0.1773 3.1256 8.0106 LAT2;FCER1G;ABCB1 9 MSigDB_Hallmark_2020 Pperoxisome 2/104 0.094 0.1965 4.0022 9.4649 PABPC1;ABCB1 11 MSigDB_Hallmark_2020 UV Response Up 2/158 0.1843 0.3532 2.6097 4.4139 LYN;ABCB1 12 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 2/200 0.2605 0.3995 2.0518 2.7598 MAP3K8;CCL5 13 MSigDB_Hallmark_2020 Estrogen Response Late 2/200 0.2605 0.3995 2.0518 2.7598 PDCD4;LTF 13 MSigDB_Hallmark_2020 Interferon Gamma Response 2/200 0.2605 0.3995 2.0518 2.7598 XCL1;CCL5 13 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.3512 0.5049 2.3511 2.4601 MAP3K8 16 MSigDB_Hallmark_2020 PI3K/AKT/mTOR Signaling 1/105 0.4069 0.5505 1.9424 1.7466 CAMK4 17 MSigDB_Hallmark_2020 Spermatogenesis 1/135 0.4894 0.6016 1.5053 1.0756 CAMK4 18 MSigDB_Hallmark_2020 Coagulation 1/138 0.497 0.6016 1.4721 1.0292 APOC2 19 MSigDB_Hallmark_2020 G2-M Checkpoint 1/200 0.6312 0.6312 1.0103 0.4649 BUB3 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
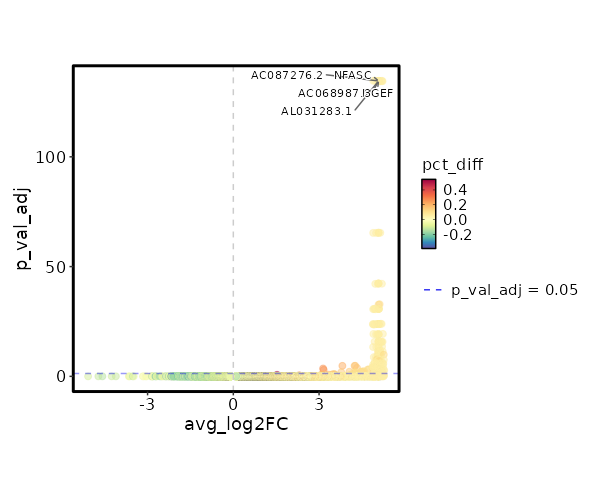
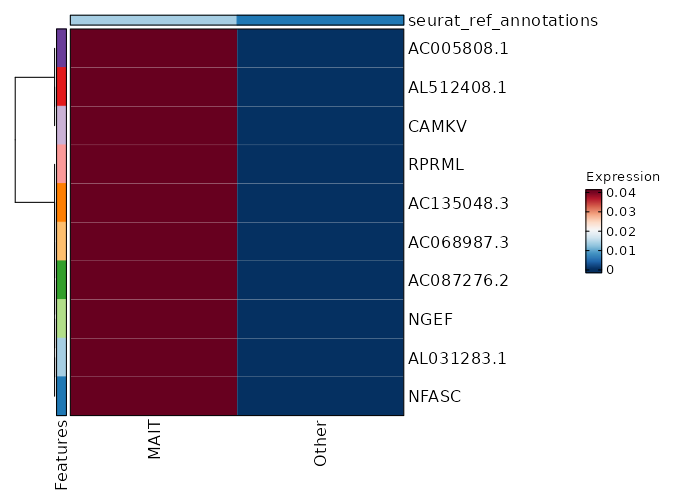
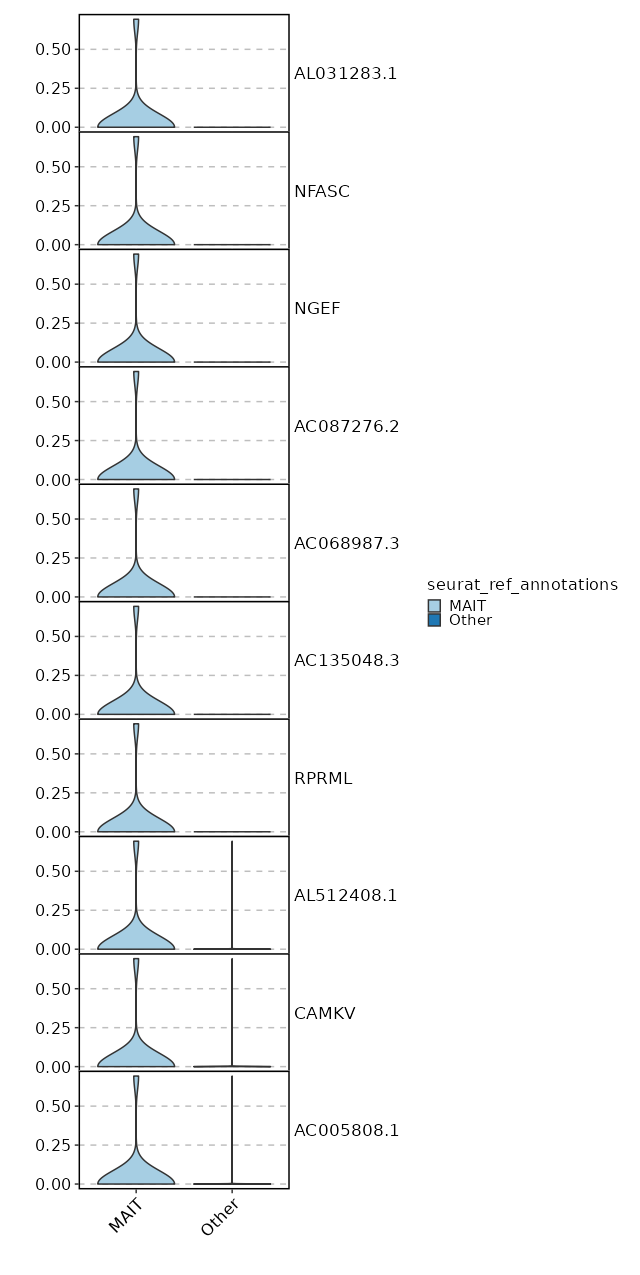
seurat_ref_annotations: MAIT Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
8.047e-140 10.3078 0.059 0 2.699e-135 AL031283.1 MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 NFASC MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 NGEF MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 AC087276.2 MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 AC068987.3 MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 AC135048.3 MAIT 8.047e-140 10.3078 0.059 0 2.699e-135 RPRML MAIT 1.159e-70 9.3078 0.059 0 3.887e-66 AL512408.1 MAIT 1.159e-70 9.3078 0.059 0 3.887e-66 CAMKV MAIT 1.159e-70 9.3078 0.059 0 3.887e-66 AC005808.1 MAIT 1.427e-47 8.7228 0.059 0 4.787e-43 AL358473.1 MAIT 1.427e-47 8.7228 0.059 0 4.787e-43 AL158834.2 MAIT 1.427e-47 8.7228 0.059 0 4.787e-43 AC004771.5 MAIT 6.259e-38 7.0854 0.118 0.001 2.099e-33 ZNRF3 MAIT 5.225e-36 8.3078 0.059 0 1.752e-31 AL031985.4 MAIT 5.225e-36 8.3078 0.059 0 1.752e-31 AL139161.1 MAIT 5.225e-36 8.3078 0.059 0 1.752e-31 ENHO MAIT 5.225e-36 8.3078 0.059 0 1.752e-31 AP003031.2 MAIT 5.225e-36 8.3078 0.059 0 1.752e-31 AL031595.2 MAIT 4.646e-29 7.9859 0.059 0 1.558e-24 PCDHGA7 MAIT
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
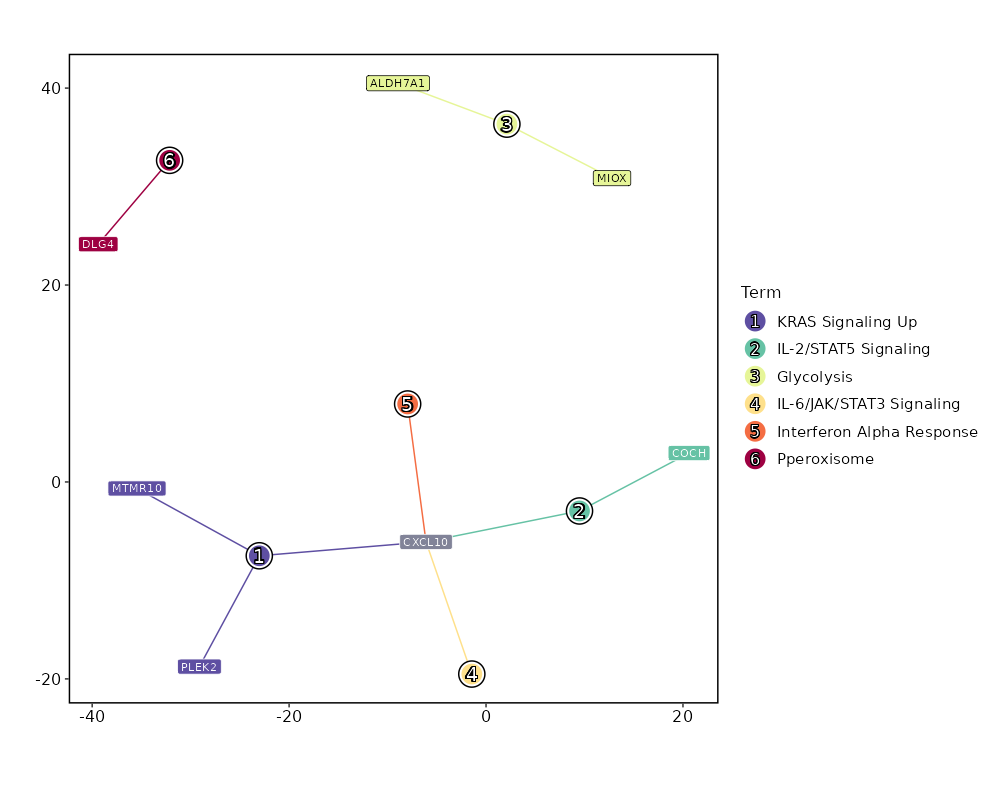
Enrichment Analysis KRAS Signaling Up 3/200 0.0962 0.6666 2.8293 6.6256 PLEK2;MTMR10;CXCL10 1 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 2/199 0.2955 0.6666 1.8686 2.2777 CXCL10;COCH 2 MSigDB_Hallmark_2020 Glycolysis 2/200 0.2976 0.6666 1.8591 2.2535 ALDH7A1;MIOX 3 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.379 0.6666 2.1323 2.0686 CXCL10 4 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.4122 0.6666 1.9092 1.692 CXCL10 5 MSigDB_Hallmark_2020 Pperoxisome 1/104 0.4344 0.6666 1.7789 1.4833 DLG4 6 MSigDB_Hallmark_2020 Spermatogenesis 1/135 0.523 0.6666 1.3652 0.8849 OAZ3 7 MSigDB_Hallmark_2020 UV Response Up 1/158 0.5797 0.6666 1.1638 0.6345 DLG4 8 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 1/200 0.6666 0.6666 0.9162 0.3716 CXCL10 9 MSigDB_Hallmark_2020 Hypoxia 1/200 0.6666 0.6666 0.9162 0.3716 CAVIN1 9 MSigDB_Hallmark_2020 Adipogenesis 1/200 0.6666 0.6666 0.9162 0.3716 CAVIN1 9 MSigDB_Hallmark_2020 Estrogen Response Late 1/200 0.6666 0.6666 0.9162 0.3716 CKB 9 MSigDB_Hallmark_2020 Myogenesis 1/200 0.6666 0.6666 0.9162 0.3716 CKB 9 MSigDB_Hallmark_2020 Interferon Gamma Response 1/200 0.6666 0.6666 0.9162 0.3716 CXCL10 9 MSigDB_Hallmark_2020 Apical Junction 1/200 0.6666 0.6666 0.9162 0.3716 NFASC 9 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.6666 0.6666 0.9162 0.3716 CXCL10 9 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
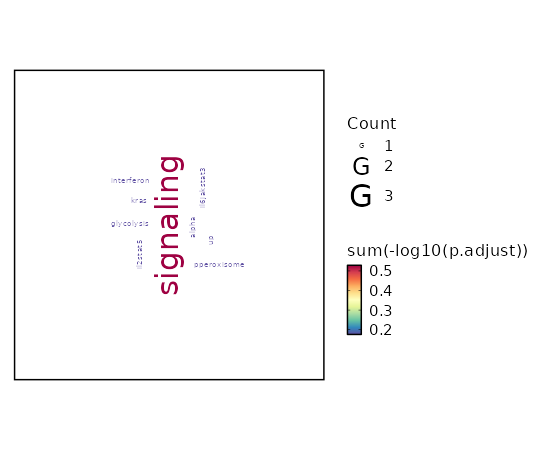
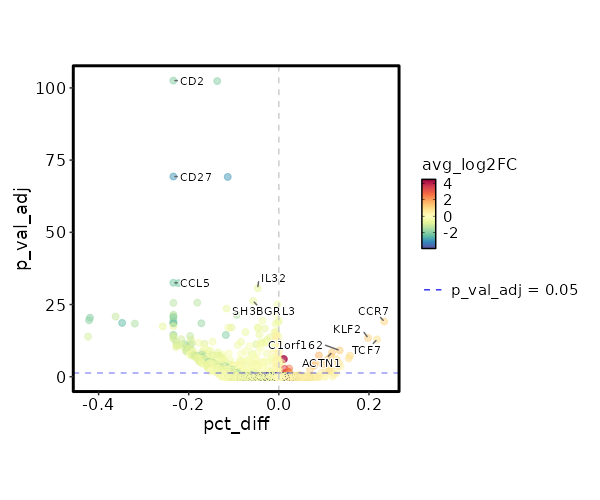
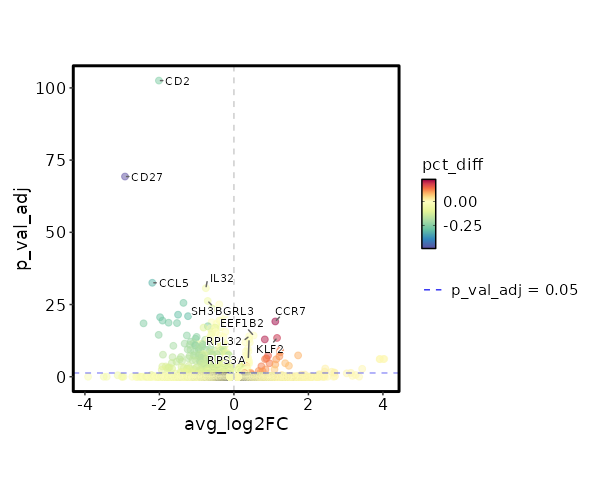
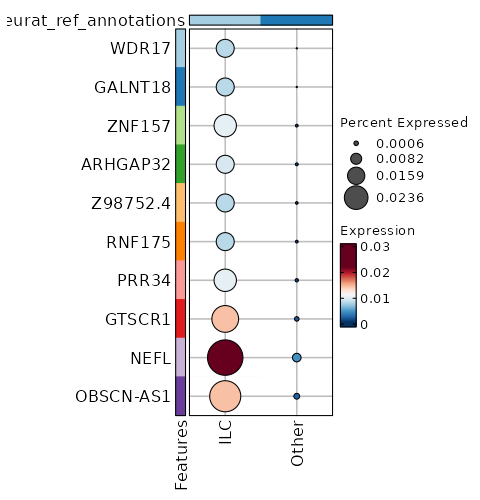
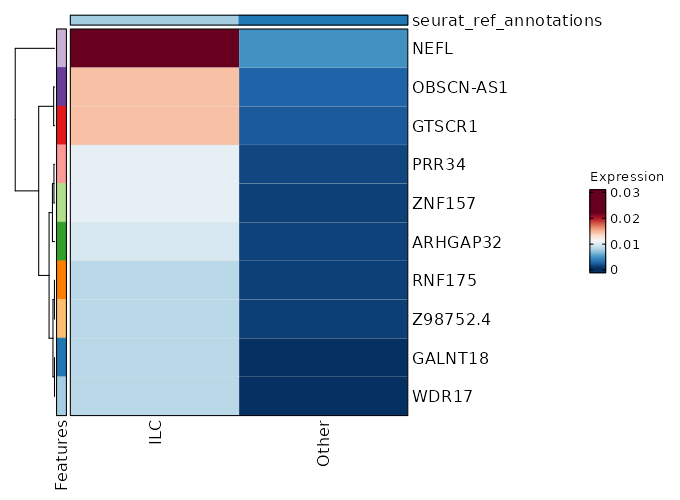
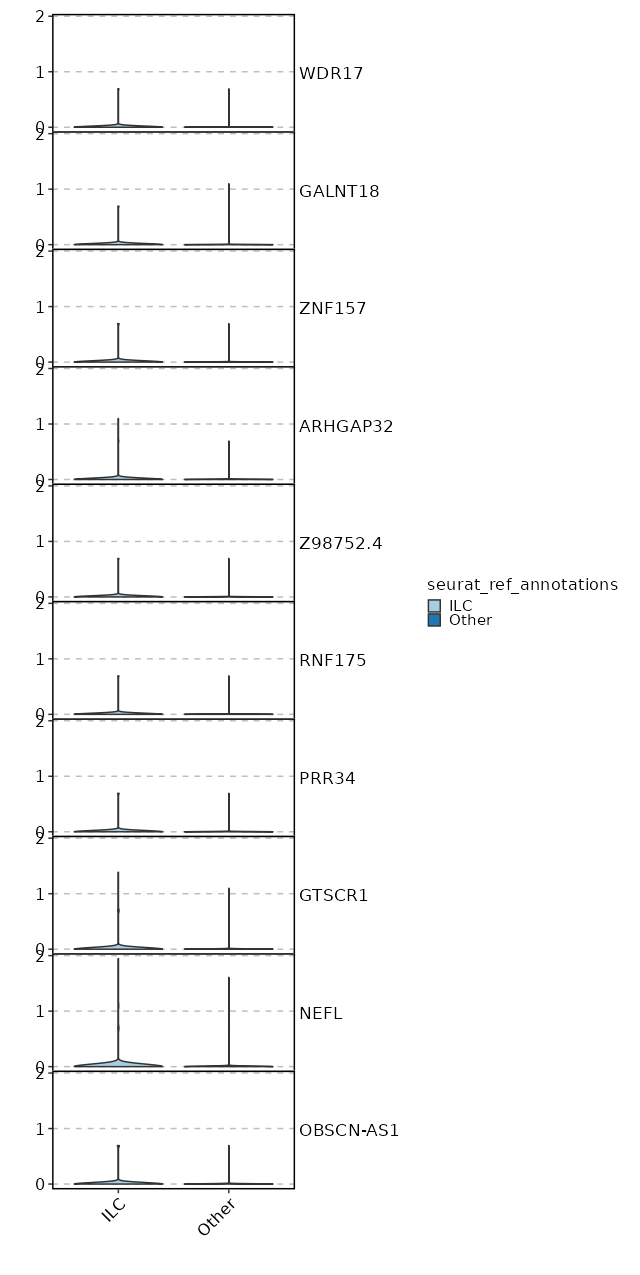
seurat_ref_annotations: ILC Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
2.157e-24 1.1108 0.466 0.232 7.235e-20 CCR7 ILC 1.621e-19 0.5282 0.979 0.984 0 EEF1B2 ILC 2.897e-19 0.4098 0.994 1 0 RPL32 ILC 1.097e-18 1.1579 0.534 0.336 0 KLF2 ILC 3.202e-18 0.4069 0.991 0.999 0 RPS3A ILC 3.546e-18 0.8296 0.543 0.325 0 TCF7 ILC 3.476e-17 0.3029 0.997 1 0 RPL30 ILC 4.032e-17 0.426 1 1 0 RPS12 ILC 0 0.3734 0.997 0.999 0 RPS13 ILC 0 1.2409 0.251 0.116 0 C1orf162 ILC 0 0.3229 1 1 0 RPL10 ILC 0 0.3889 0.985 0.996 0 RPL9 ILC 0 1.2403 0.204 0.086 0 ACTN1 ILC 0 1.7233 0.142 0.053 0 LINC01891 ILC 0 0.9155 0.428 0.27 0 LEF1 ILC 0 0.3114 1 1 0 RPS27A ILC 0 0.3204 0.997 1 0 RPS14 ILC 0 1.22 0.218 0.102 0 TRABD2A ILC 0 0.3817 1 1 0 RPS8 ILC 0 0.8734 0.378 0.223 0 PLAC8 ILC
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
Enrichment Analysis Myc Targets V1 10/200 0 0 16.2303 317.0208 RPL22;RPL18;RPL34;LDHA;COX5A;RPS6;PPIA;RPS5;RPS3;EEF1B2 1 MSigDB_Hallmark_2020 Glycolysis 5/200 0.0009 0.011 7.3322 51.6191 TXN;ENO1;TPI1;LDHA;PPIA 2 MSigDB_Hallmark_2020 Hypoxia 4/200 0.0065 0.0324 5.7522 28.9913 ENO1;PLAC8;TPI1;LDHA 3 MSigDB_Hallmark_2020 mTORC1 Signaling 4/200 0.0065 0.0324 5.7522 28.9913 ENO1;TPI1;LDHA;PPIA 3 MSigDB_Hallmark_2020 Allograft Rejection 4/200 0.0065 0.0324 5.7522 28.9913 RPL9;IFNGR2;RPS3A;LTB 3 MSigDB_Hallmark_2020 Wnt-beta Catenin Signaling 2/42 0.0106 0.0413 13.8097 62.8333 TCF7;LEF1 6 MSigDB_Hallmark_2020 Apical Surface 2/44 0.0116 0.0413 13.1508 58.6596 MAL;LYPD3 7 MSigDB_Hallmark_2020 Reactive Oxygen Species Pathway 2/49 0.0142 0.0444 11.7488 49.9886 TXN;JUNB 8 MSigDB_Hallmark_2020 TGF-beta Signaling 2/54 0.0171 0.0474 10.6165 43.213 IFNGR2;JUNB 9 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 3/199 0.0376 0.0732 4.2534 13.9563 SELL;ITGA6;LTB 10 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 3/200 0.0381 0.0732 4.2316 13.8317 JUNB;IFNGR2;KLF2 11 MSigDB_Hallmark_2020 Inflammatory Response 3/200 0.0381 0.0732 4.2316 13.8317 CCR7;IFNGR2;SELL 11 MSigDB_Hallmark_2020 Oxidative Phosphorylation 3/200 0.0381 0.0732 4.2316 13.8317 LDHA;COX5A;ATP5MC3 11 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 2/87 0.0413 0.0737 6.484 20.6661 IFNGR2;LTB 14 MSigDB_Hallmark_2020 Fatty Acid Metabolism 2/158 0.1161 0.1934 3.5203 7.5815 HSP90AA1;LDHA 15 MSigDB_Hallmark_2020 Apical Junction 2/200 0.1692 0.2488 2.7677 4.9176 ACTN1;LDLRAP1 16 MSigDB_Hallmark_2020 p53 Pathway 2/200 0.1692 0.2488 2.7677 4.9176 RPL18;RPS12 16 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.3026 0.4039 2.8296 3.3821 SELL 18 MSigDB_Hallmark_2020 Androgen Response 1/100 0.3104 0.4039 2.7435 3.2098 ACTN1 19 MSigDB_Hallmark_2020 PI3K/AKT/mTOR Signaling 1/105 0.3231 0.4039 2.6109 2.9498 THEM4 20 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
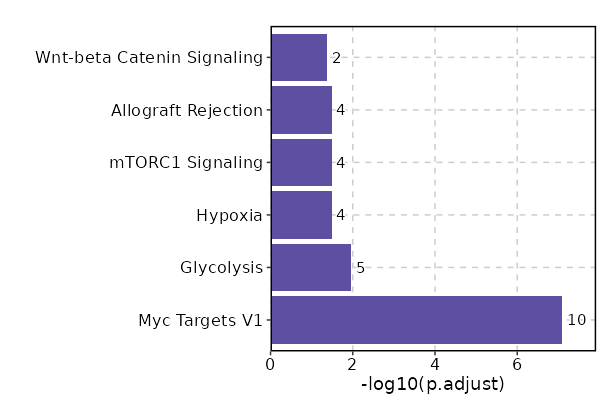
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
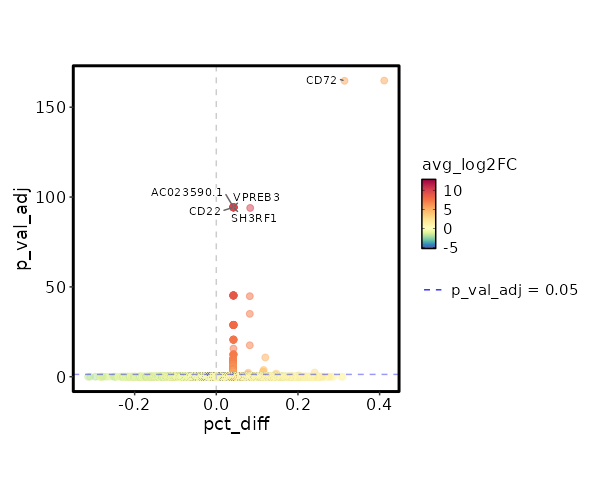
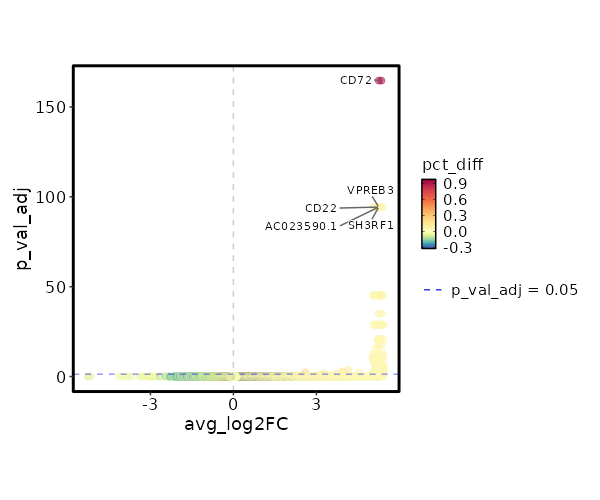
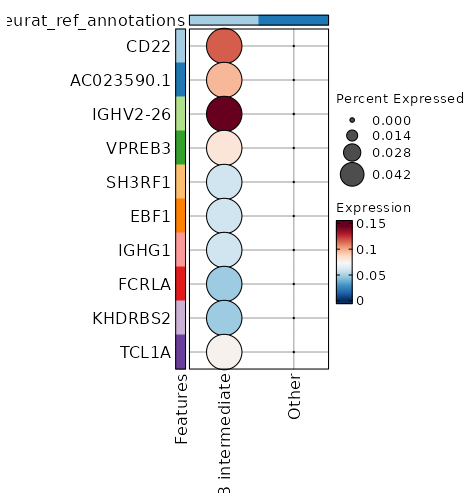
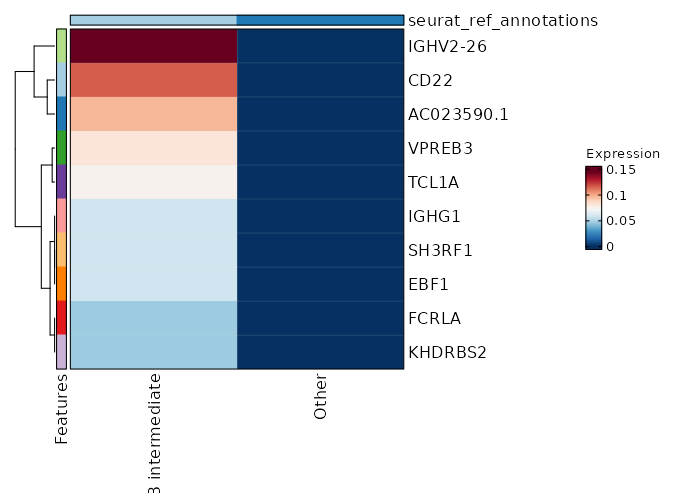
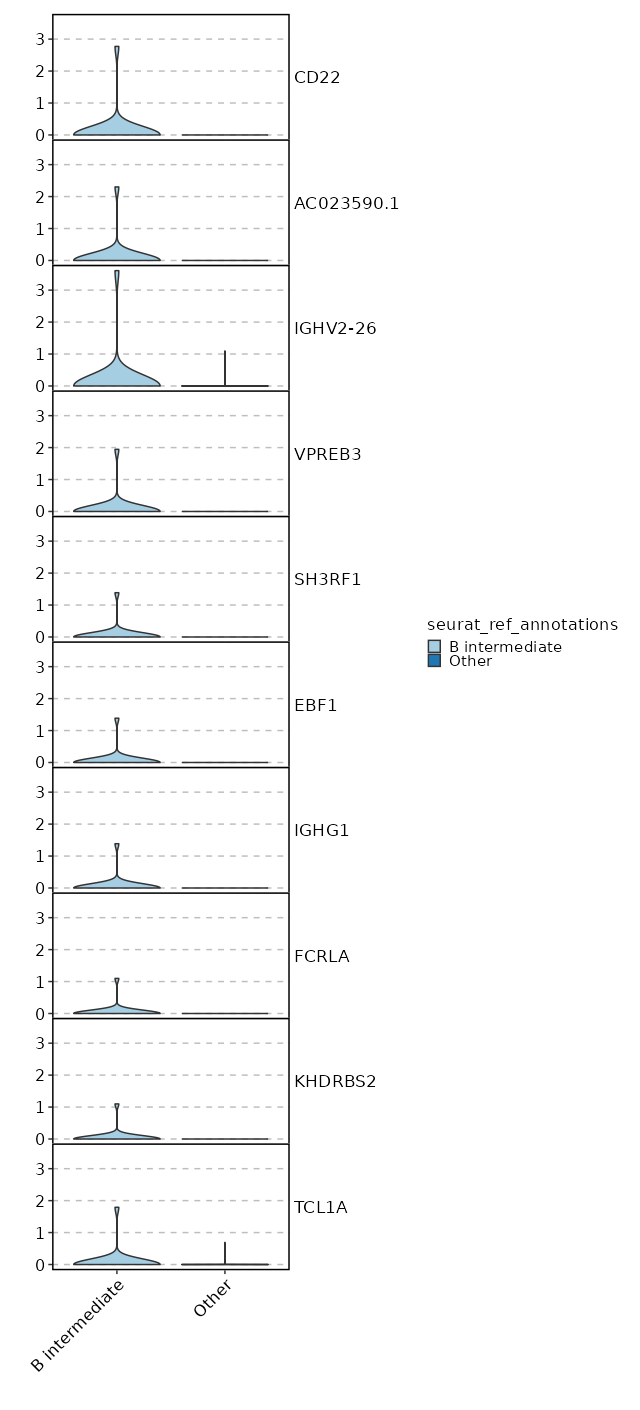
seurat_ref_annotations: B intermediate Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
6.063e-170 5.3764 1 0.028 2.034e-165 CD72 B intermediate 1.498e-99 12.8094 0.042 0 5.024e-95 CD22 B intermediate 1.498e-99 12.1313 0.042 0 5.024e-95 AC023590.1 B intermediate 1.498e-99 11.6167 0.042 0 5.024e-95 VPREB3 B intermediate 1.498e-99 10.8094 0.042 0 5.024e-95 SH3RF1 B intermediate 1.498e-99 10.8094 0.042 0 5.024e-95 EBF1 B intermediate 1.498e-99 10.8094 0.042 0 5.024e-95 IGHG1 B intermediate 1.498e-99 10.3943 0.042 0 5.024e-95 FCRLA B intermediate 1.498e-99 10.3943 0.042 0 5.024e-95 KHDRBS2 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 AL591846.2 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 CYS1 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 KCNH8 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 LINC02202 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 SHANK2 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 HTR3A B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 MYL2 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 SERPINA9 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 AL139020.1 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 CD19 B intermediate 1.498e-99 9.8094 0.042 0 5.024e-95 AP001059.2 B intermediate
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
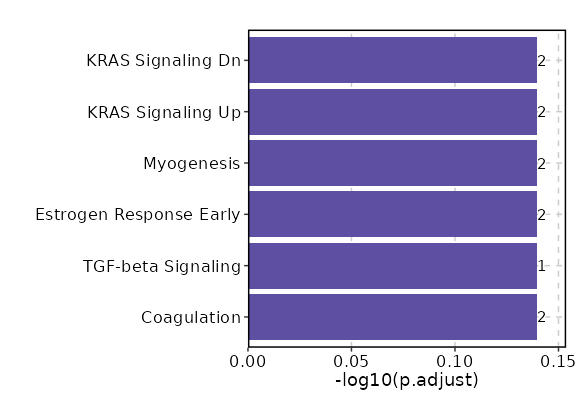
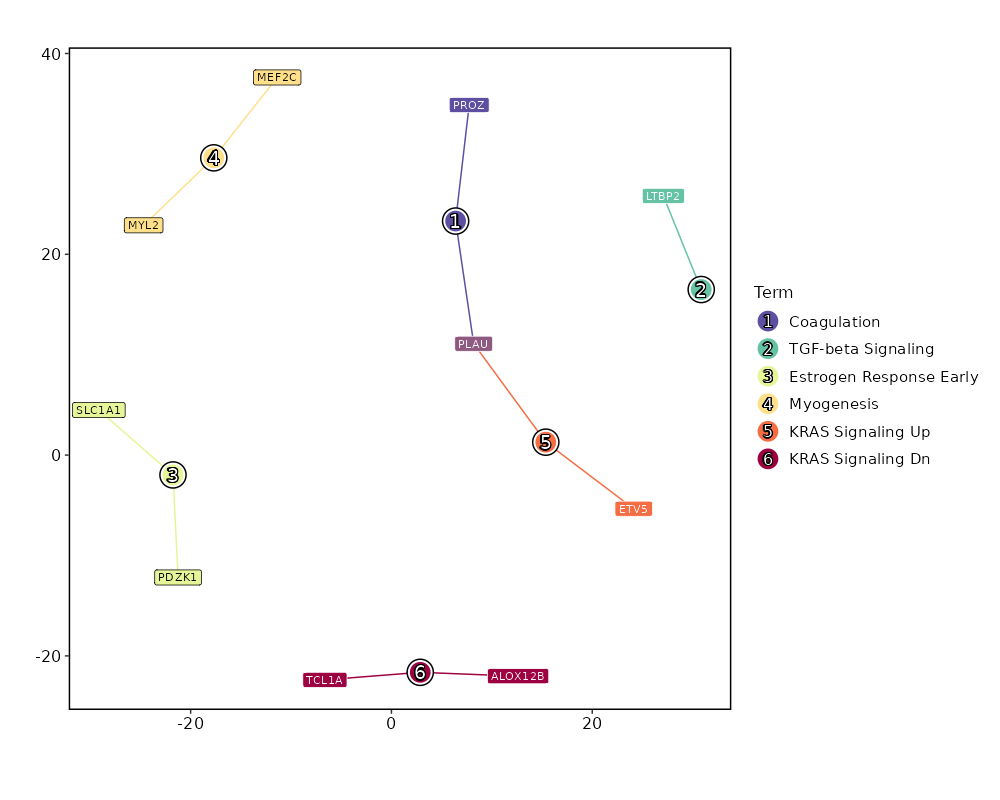
Enrichment Analysis Coagulation 2/138 0.2211 0.7249 2.3035 3.4764 PLAU;PROZ 1 MSigDB_Hallmark_2020 TGF-beta Signaling 1/54 0.2933 0.7249 2.9444 3.6114 LTBP2 2 MSigDB_Hallmark_2020 Estrogen Response Early 2/200 0.3669 0.7249 1.5772 1.5815 PDZK1;SLC1A1 3 MSigDB_Hallmark_2020 Myogenesis 2/200 0.3669 0.7249 1.5772 1.5815 MYL2;MEF2C 3 MSigDB_Hallmark_2020 KRAS Signaling Up 2/200 0.3669 0.7249 1.5772 1.5815 PLAU;ETV5 3 MSigDB_Hallmark_2020 KRAS Signaling Dn 2/200 0.3669 0.7249 1.5772 1.5815 ALOX12B;TCL1A 3 MSigDB_Hallmark_2020 Pperoxisome 1/104 0.488 0.7249 1.5113 1.0842 ABCB4 7 MSigDB_Hallmark_2020 Bile Acid Metabolism 1/112 0.5138 0.7249 1.4018 0.9335 KLF1 8 MSigDB_Hallmark_2020 Spermatogenesis 1/135 0.5809 0.7249 1.1598 0.6299 SLC2A5 9 MSigDB_Hallmark_2020 UV Response Dn 1/144 0.6046 0.7249 1.0863 0.5466 LTBP1 10 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 1/200 0.7249 0.7249 0.7784 0.2505 PLAU 11 MSigDB_Hallmark_2020 Hypoxia 1/200 0.7249 0.7249 0.7784 0.2505 SLC2A5 11 MSigDB_Hallmark_2020 Estrogen Response Late 1/200 0.7249 0.7249 0.7784 0.2505 PDZK1 11 MSigDB_Hallmark_2020 Apical Junction 1/200 0.7249 0.7249 0.7784 0.2505 NRTN 11 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.7249 0.7249 0.7784 0.2505 CYBB 11 MSigDB_Hallmark_2020 p53 Pathway 1/200 0.7249 0.7249 0.7784 0.2505 OSGIN1 11 MSigDB_Hallmark_2020 heme Metabolism 1/200 0.7249 0.7249 0.7784 0.2505 KLF1 11 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
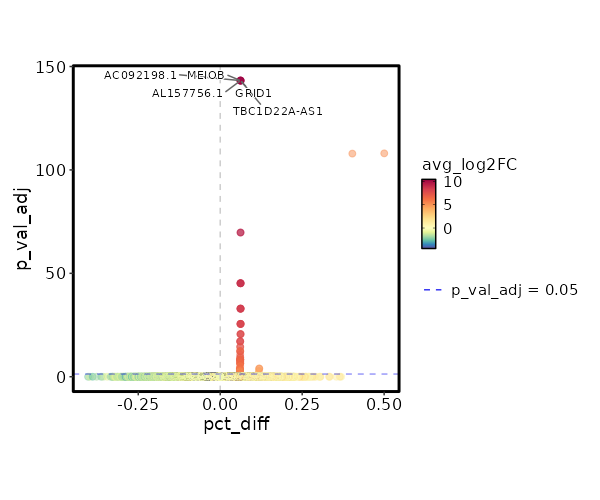
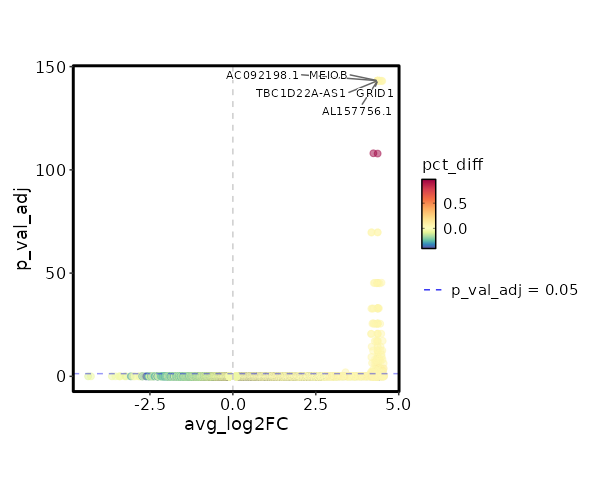
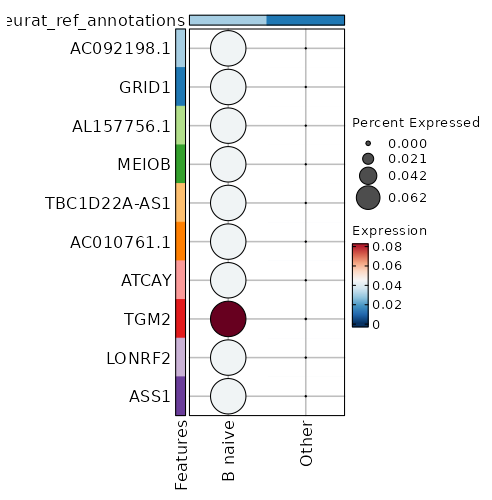
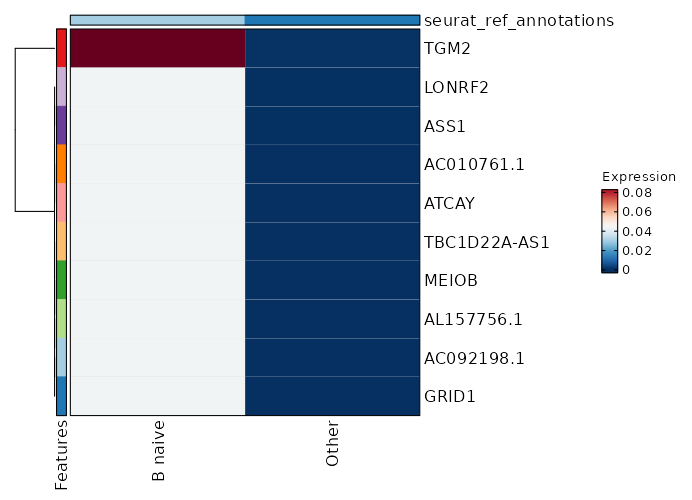
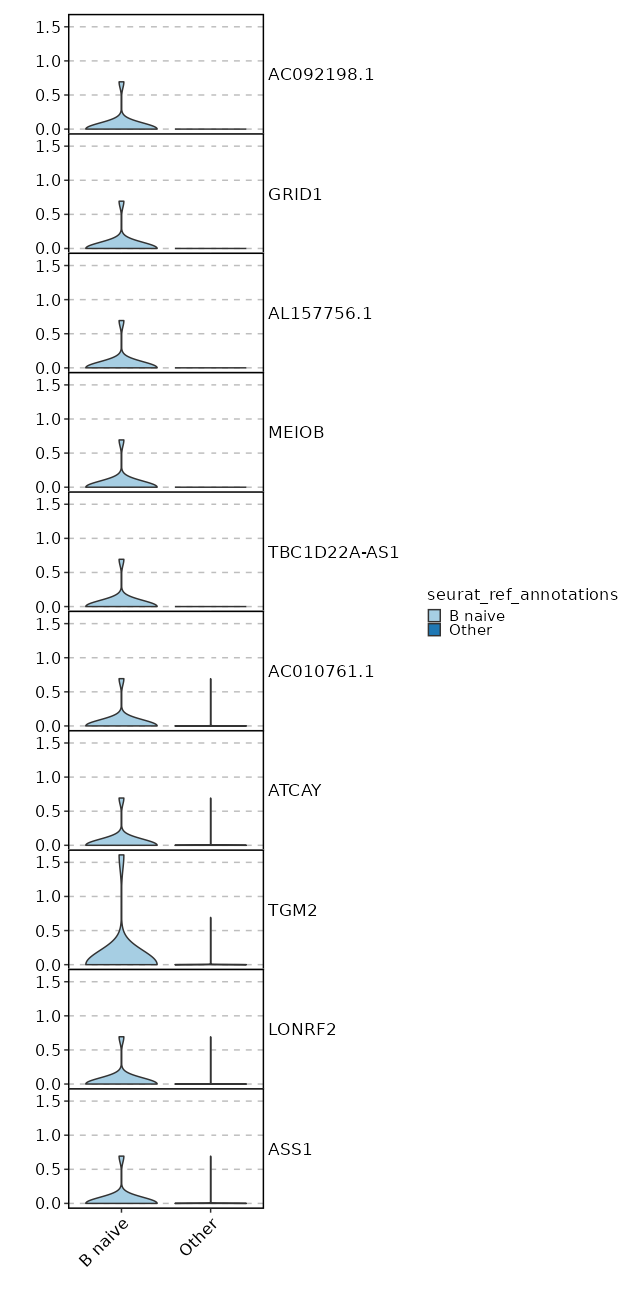
seurat_ref_annotations: B naive Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
1.902e-148 10.3954 0.062 0 6.379e-144 AC092198.1 B naive 1.902e-148 10.3954 0.062 0 6.379e-144 GRID1 B naive 1.902e-148 10.3954 0.062 0 6.379e-144 AL157756.1 B naive 1.902e-148 10.3954 0.062 0 6.379e-144 MEIOB B naive 1.902e-148 10.3954 0.062 0 6.379e-144 TBC1D22A-AS1 B naive 3.846e-113 5.173 1 0.029 1.29e-108 CD72 B naive 5.545e-75 9.3954 0.062 0 1.86e-70 AC010761.1 B naive 5.545e-75 9.3954 0.062 0 1.86e-70 ATCAY B naive 1.861e-50 8.8104 0.062 0 6.243e-46 LONRF2 B naive 1.861e-50 8.8104 0.062 0 6.243e-46 ASS1 B naive 1.861e-50 8.8104 0.062 0 6.243e-46 NCAM1-AS1 B naive 1.861e-50 8.8104 0.062 0 6.243e-46 AL162274.1 B naive 1.861e-50 8.8104 0.062 0 6.243e-46 ADAMTS7 B naive 3.557e-38 8.3954 0.062 0 1.193e-33 COBLL1 B naive 3.557e-38 8.3954 0.062 0 1.193e-33 AC027271.1 B naive 3.557e-38 8.3954 0.062 0 1.193e-33 KCNRG B naive 3.557e-38 8.3954 0.062 0 1.193e-33 AKAP6 B naive 3.557e-38 8.3954 0.062 0 1.193e-33 WFIKKN1 B naive 8.525e-31 8.0735 0.062 0 2.859e-26 C1orf220 B naive 8.525e-31 8.0735 0.062 0 2.859e-26 LINC01303 B naive
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
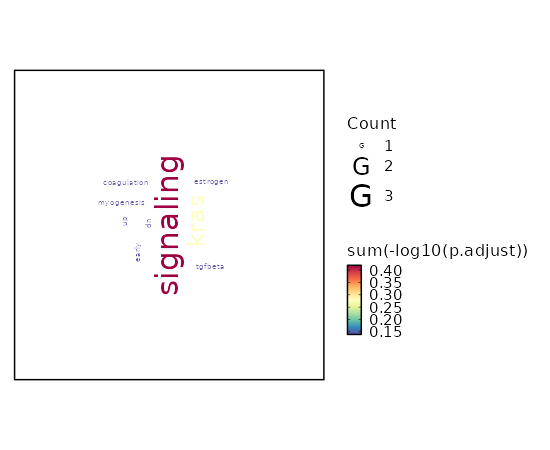
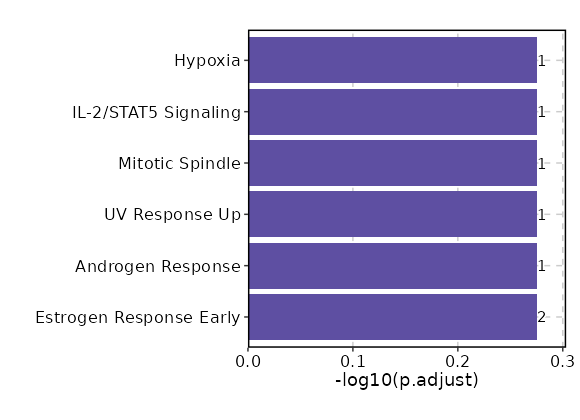
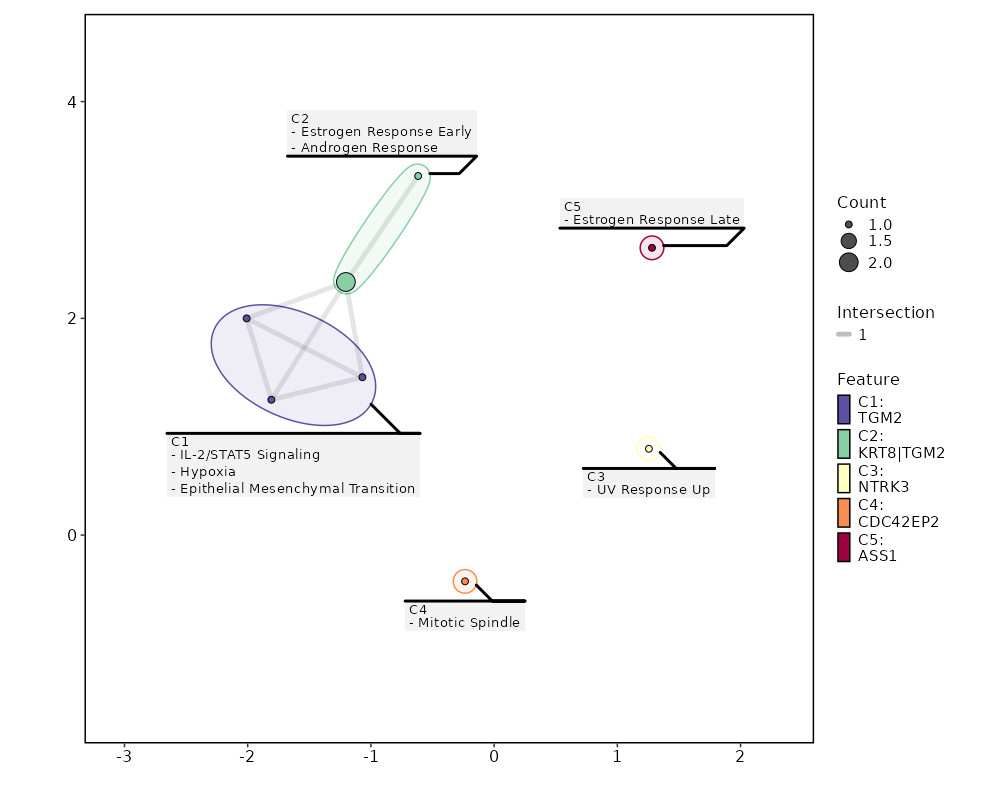
Enrichment Analysis Estrogen Response Early 2/200 0.1727 0.5301 2.7296 4.7932 KRT8;TGM2 1 MSigDB_Hallmark_2020 Androgen Response 1/100 0.3138 0.5301 2.7063 3.1362 KRT8 2 MSigDB_Hallmark_2020 UV Response Up 1/158 0.449 0.5301 1.7015 1.3626 NTRK3 3 MSigDB_Hallmark_2020 Mitotic Spindle 1/199 0.5283 0.5301 1.3464 0.8591 CDC42EP2 4 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 1/199 0.5283 0.5301 1.3464 0.8591 TGM2 4 MSigDB_Hallmark_2020 Hypoxia 1/200 0.5301 0.5301 1.3395 0.8503 TGM2 6 MSigDB_Hallmark_2020 Estrogen Response Late 1/200 0.5301 0.5301 1.3395 0.8503 ASS1 6 MSigDB_Hallmark_2020 Epithelial Mesenchymal Transition 1/200 0.5301 0.5301 1.3395 0.8503 TGM2 6 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
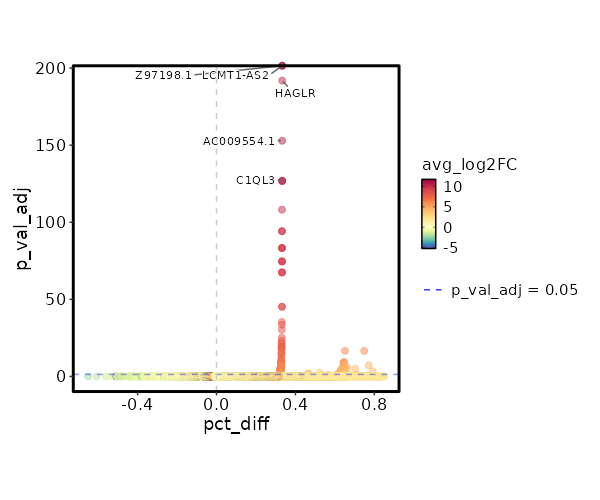
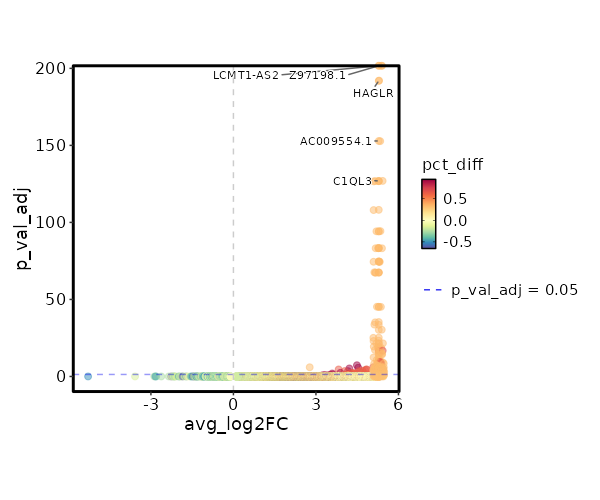
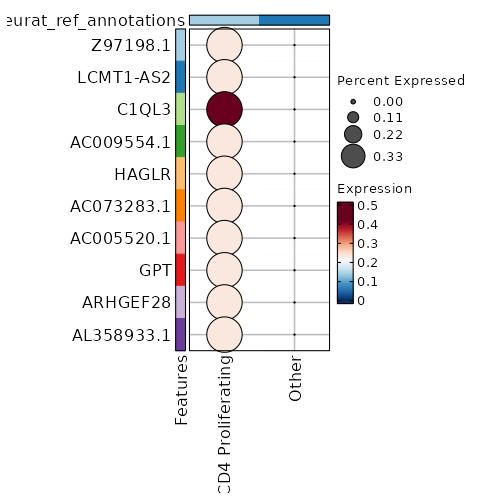
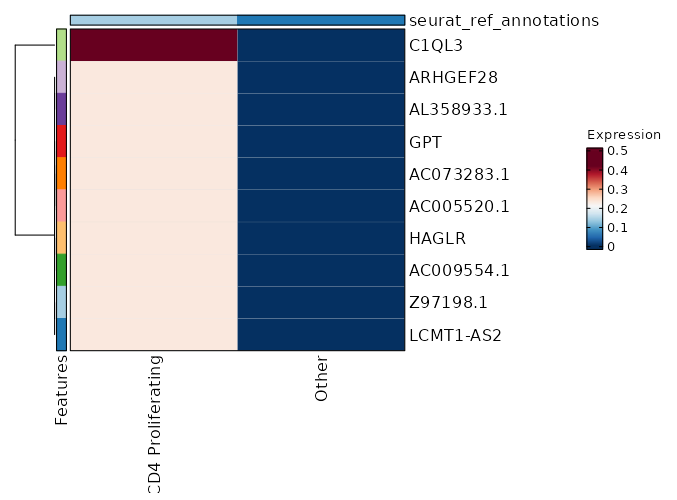
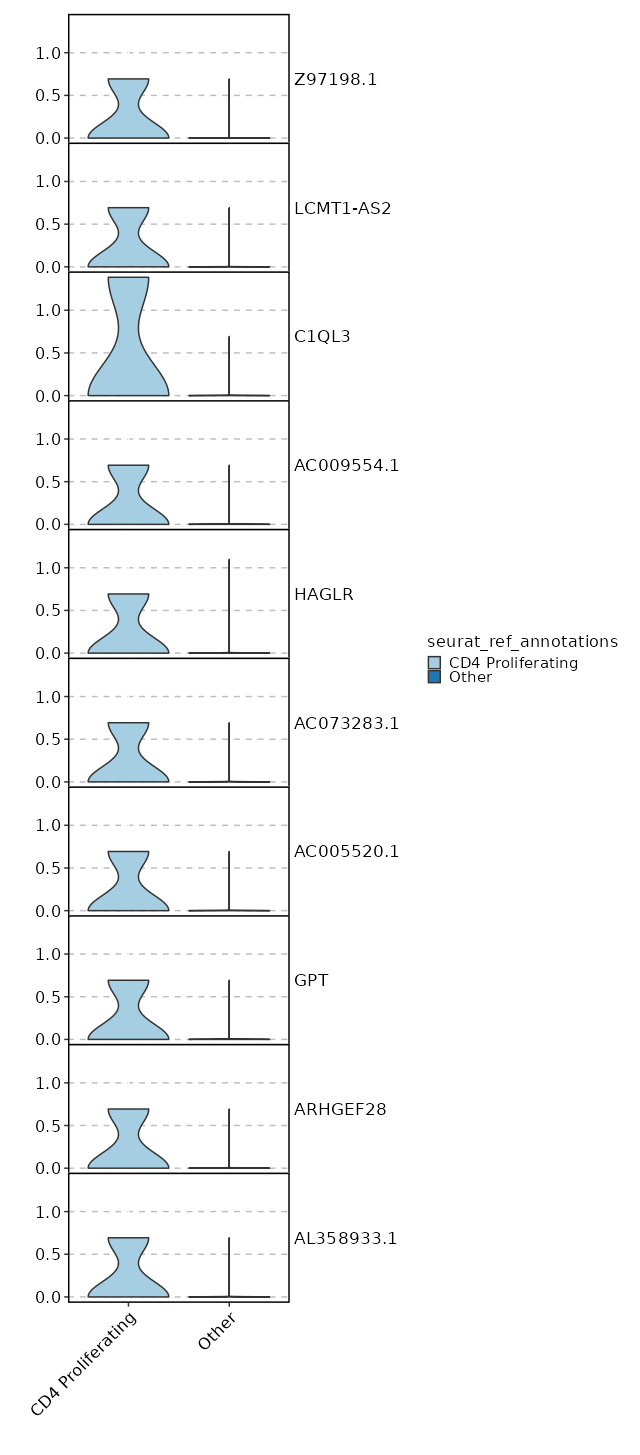
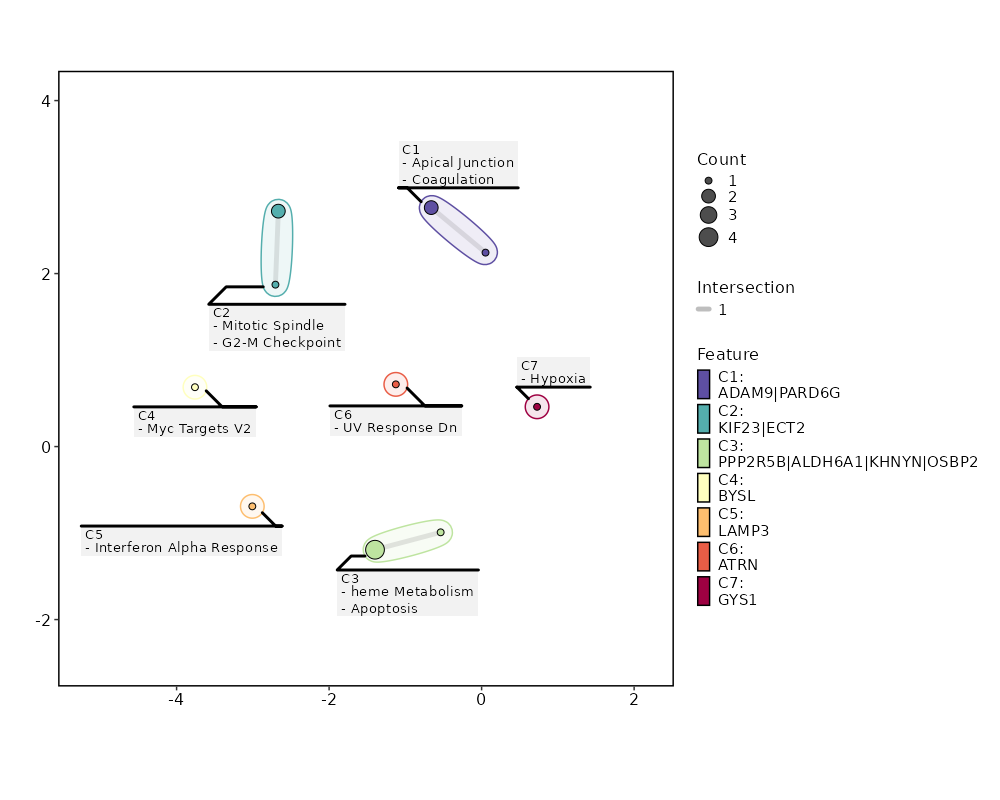
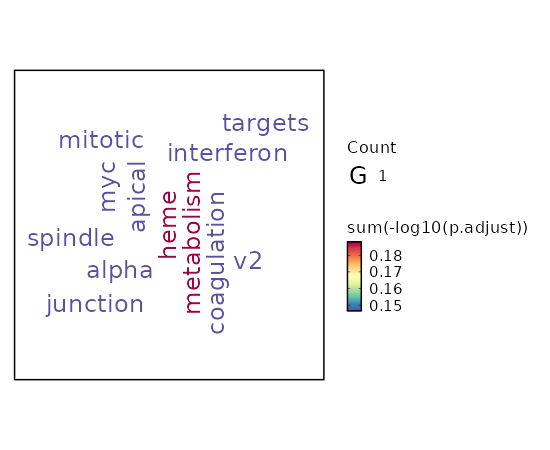
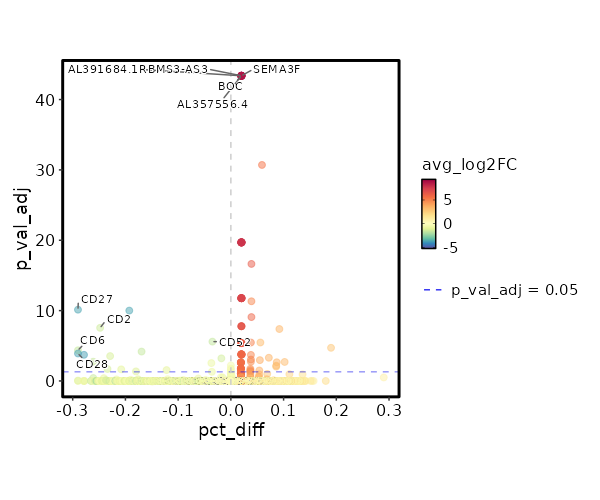
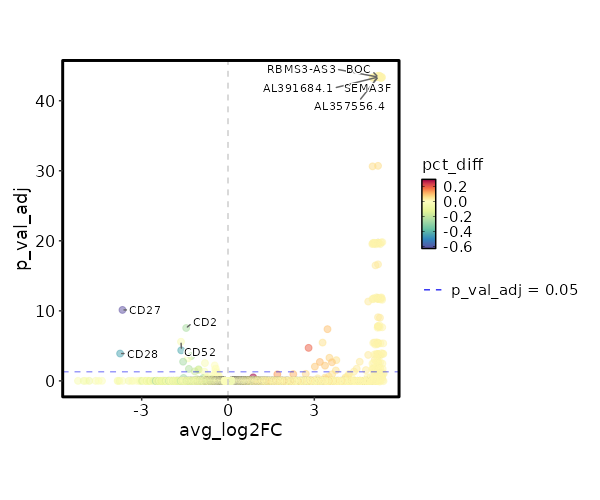
seurat_ref_annotations: CD4 Proliferating Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
0 11.8122 0.333 0 0 Z97198.1 CD4 Proliferating 0 11.8122 0.333 0 0 LCMT1-AS2 CD4 Proliferating 3.886e-197 10.2272 0.333 0 1.303e-192 HAGLR CD4 Proliferating 4.163e-158 10.4902 0.333 0 1.396e-153 AC009554.1 CD4 Proliferating 3.752e-132 11.2272 0.333 0 1.258e-127 C1QL3 CD4 Proliferating 4.954e-132 10.2272 0.333 0 1.661e-127 AC073283.1 CD4 Proliferating 4.954e-132 10.2272 0.333 0 1.661e-127 AC005520.1 CD4 Proliferating 2.116e-113 10.0048 0.333 0.001 7.096e-109 GPT CD4 Proliferating 2.006e-99 9.8122 0.333 0.001 6.727e-95 ARHGEF28 CD4 Proliferating 2.006e-99 9.8122 0.333 0.001 6.727e-95 AL358933.1 CD4 Proliferating 1.5e-88 9.6423 0.333 0.001 5.031e-84 FHAD1 CD4 Proliferating 1.5e-88 9.6423 0.333 0.001 5.031e-84 AC022400.7 CD4 Proliferating 1.557e-88 9.4902 0.333 0.001 5.221e-84 TERT CD4 Proliferating 7.544e-80 9.4902 0.333 0.001 2.53e-75 AC023794.3 CD4 Proliferating 7.544e-80 9.4902 0.333 0.001 2.53e-75 AL132639.3 CD4 Proliferating 7.544e-80 9.4902 0.333 0.001 2.53e-75 IGLON5 CD4 Proliferating 9.977e-73 9.3527 0.333 0.001 3.346e-68 AC105411.1 CD4 Proliferating 9.977e-73 9.3527 0.333 0.001 3.346e-68 AC011448.1 CD4 Proliferating 1.028e-72 9.2272 0.333 0.001 3.449e-68 SH3RF2 CD4 Proliferating 1.861e-50 8.8122 0.333 0.001 6.243e-46 AC069224.1 CD4 Proliferating
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
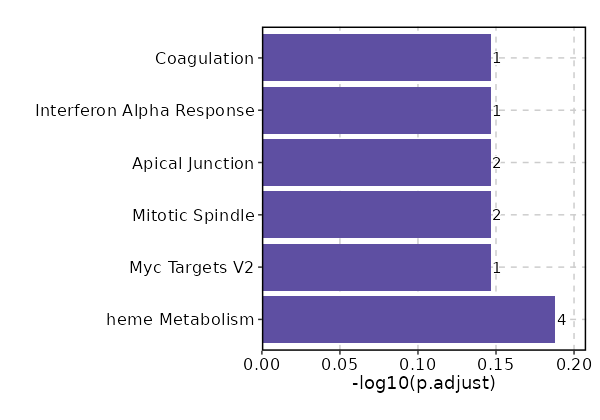
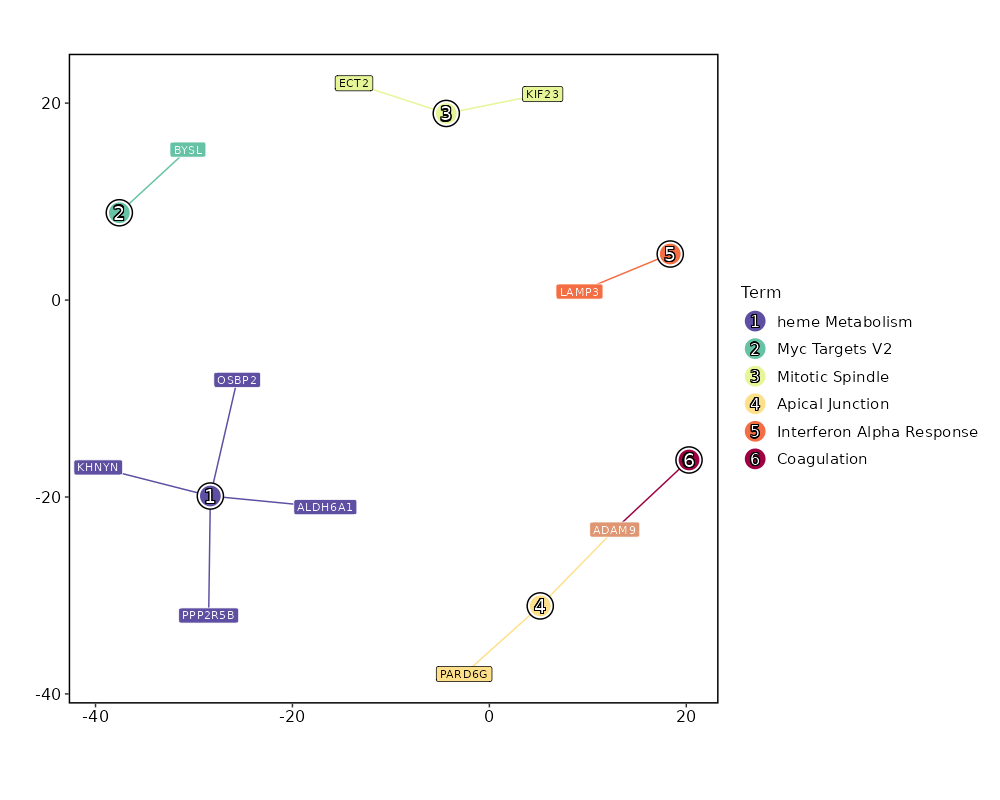
Enrichment Analysis heme Metabolism 4/200 0.036 0.6486 3.3469 11.1231 ALDH6A1;OSBP2;PPP2R5B;KHNYN 1 MSigDB_Hallmark_2020 Myc Targets V2 1/58 0.3032 0.7135 2.8268 3.3736 BYSL 2 MSigDB_Hallmark_2020 Mitotic Spindle 2/199 0.3502 0.7135 1.6376 1.7181 KIF23;ECT2 3 MSigDB_Hallmark_2020 Apical Junction 2/200 0.3525 0.7135 1.6292 1.699 PARD6G;ADAM9 4 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.4538 0.7135 1.6751 1.3236 LAMP3 5 MSigDB_Hallmark_2020 Coagulation 1/138 0.5774 0.7135 1.1714 0.6434 ADAM9 6 MSigDB_Hallmark_2020 UV Response Dn 1/144 0.5929 0.7135 1.1219 0.5864 ATRN 7 MSigDB_Hallmark_2020 Apoptosis 1/161 0.6341 0.7135 1.0018 0.4564 PPP2R5B 8 MSigDB_Hallmark_2020 Hypoxia 1/200 0.7135 0.7135 0.8039 0.2713 GYS1 9 MSigDB_Hallmark_2020 G2-M Checkpoint 1/200 0.7135 0.7135 0.8039 0.2713 KIF23 9 MSigDB_Hallmark_2020 Myogenesis 1/200 0.7135 0.7135 0.8039 0.2713 COL6A3 9 MSigDB_Hallmark_2020 Complement 1/200 0.7135 0.7135 0.8039 0.2713 ADAM9 9 MSigDB_Hallmark_2020 E2F Targets 1/200 0.7135 0.7135 0.8039 0.2713 WDR90 9 MSigDB_Hallmark_2020 Epithelial Mesenchymal Transition 1/200 0.7135 0.7135 0.8039 0.2713 COL6A3 9 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.7135 0.7135 0.8039 0.2713 LAMP3 9 MSigDB_Hallmark_2020 Oxidative Phosphorylation 1/200 0.7135 0.7135 0.8039 0.2713 ALDH6A1 9 MSigDB_Hallmark_2020 Glycolysis 1/200 0.7135 0.7135 0.8039 0.2713 GYS1 9 MSigDB_Hallmark_2020 Allograft Rejection 1/200 0.7135 0.7135 0.8039 0.2713 IL12A 9 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
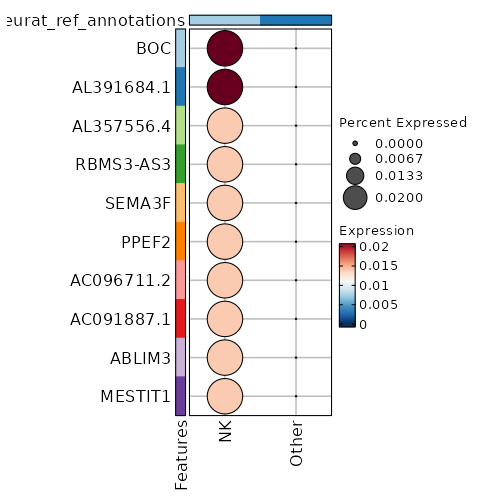
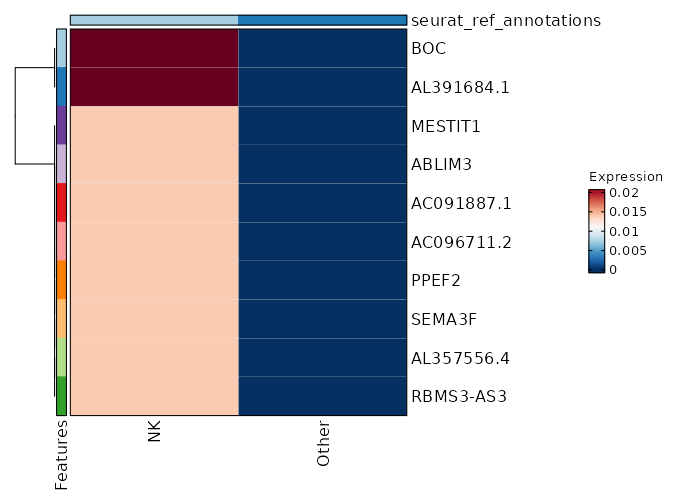
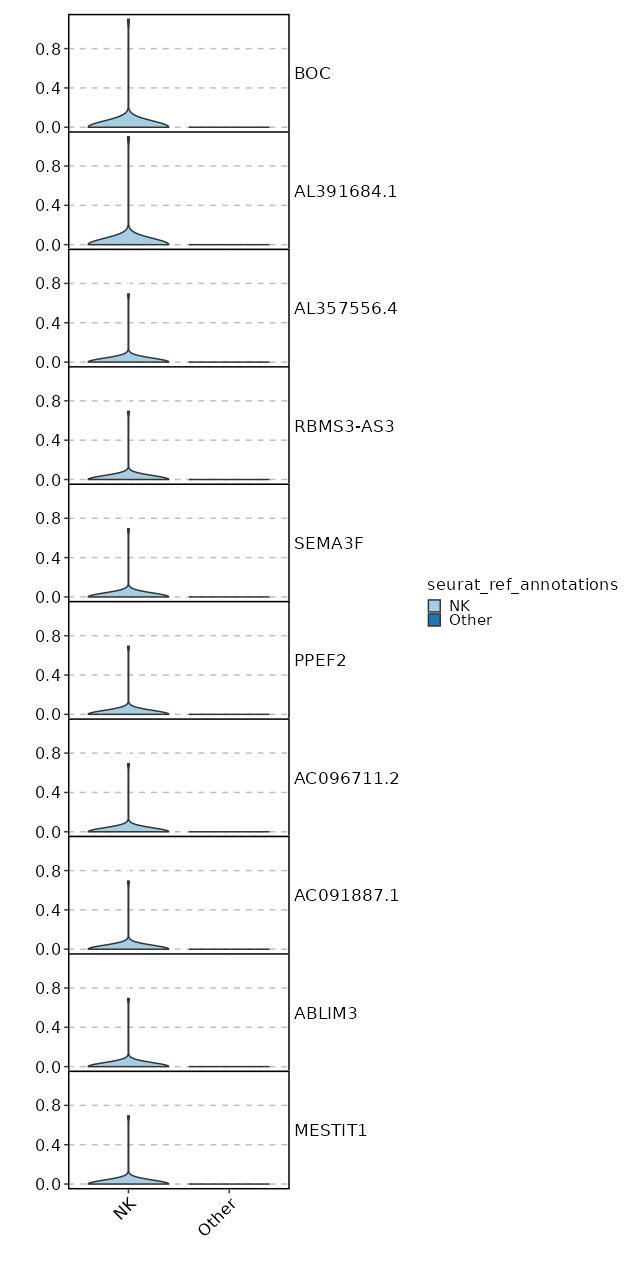
seurat_ref_annotations: NK Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
1.245e-48 9.3319 0.02 0 4.174e-44 BOC NK 1.245e-48 9.3319 0.02 0 4.174e-44 AL391684.1 NK 1.245e-48 8.747 0.02 0 4.174e-44 AL357556.4 NK 1.245e-48 8.747 0.02 0 4.174e-44 RBMS3-AS3 NK 1.245e-48 8.747 0.02 0 4.174e-44 SEMA3F NK 1.245e-48 8.747 0.02 0 4.174e-44 PPEF2 NK 1.245e-48 8.747 0.02 0 4.174e-44 AC096711.2 NK 1.245e-48 8.747 0.02 0 4.174e-44 AC091887.1 NK 1.245e-48 8.747 0.02 0 4.174e-44 ABLIM3 NK 1.245e-48 8.747 0.02 0 4.174e-44 MESTIT1 NK 1.245e-48 8.747 0.02 0 4.174e-44 SMIM10L2B NK 1.245e-48 8.747 0.02 0 4.174e-44 AC137834.2 NK 1.245e-48 8.747 0.02 0 4.174e-44 JPH4 NK 1.245e-48 8.747 0.02 0 4.174e-44 AC007673.1 NK 5.901e-36 5.4389 0.06 0.001 1.979e-31 TCN1 NK 5.959e-25 8.3319 0.02 0 1.998e-20 MARCKS NK 6.019e-25 7.747 0.02 0 2.019e-20 FOXP1-AS1 NK 6.019e-25 7.747 0.02 0 2.019e-20 AC080013.1 NK 6.019e-25 7.747 0.02 0 2.019e-20 AP000777.3 NK 6.019e-25 7.747 0.02 0 2.019e-20 ARHGAP20 NK
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
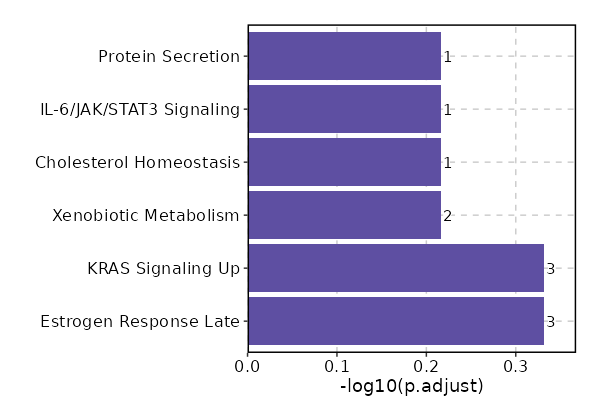
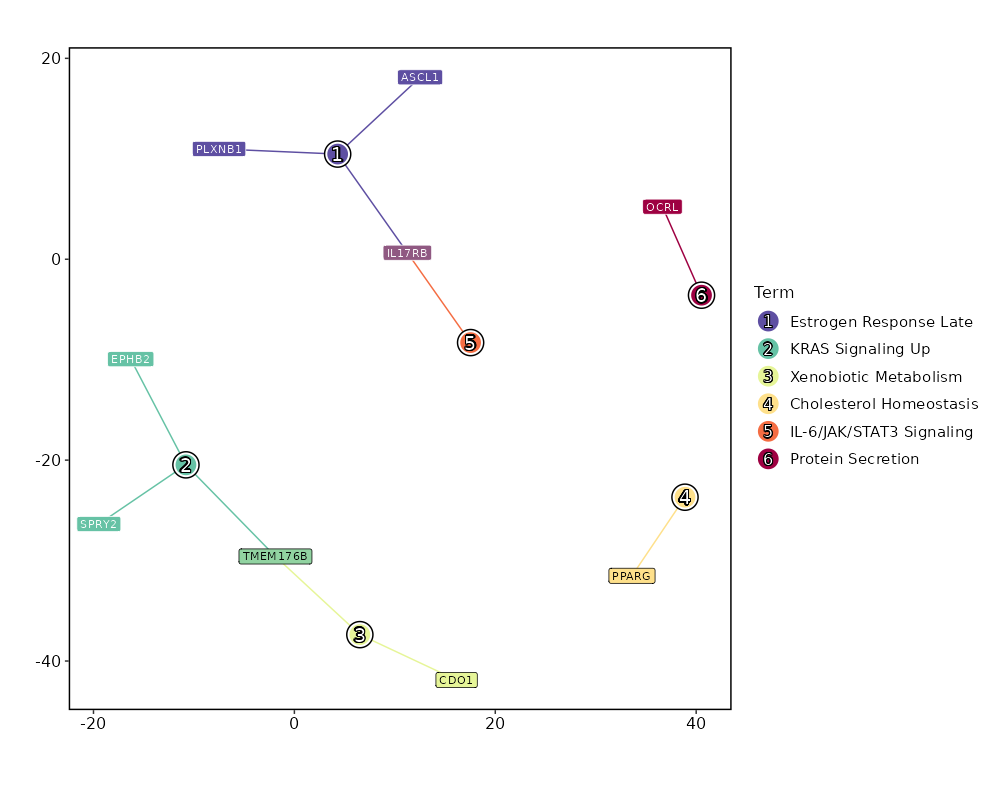
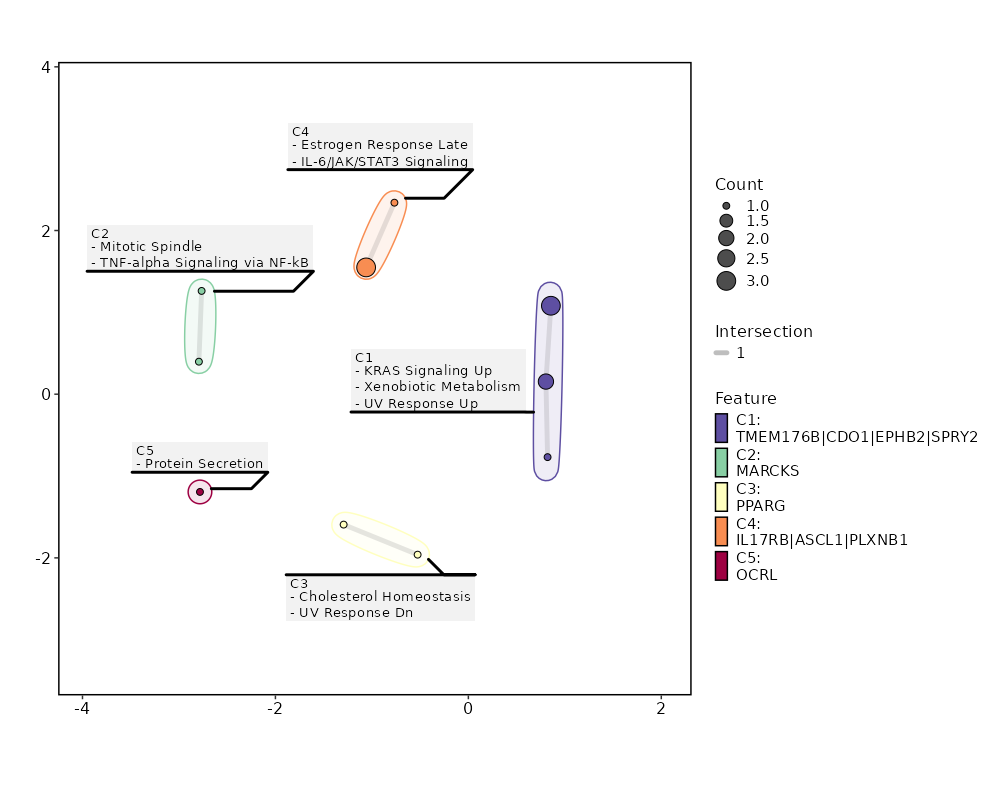
Enrichment Analysis Estrogen Response Late 3/200 0.0665 0.4658 3.335 9.0378 ASCL1;IL17RB;PLXNB1 1 MSigDB_Hallmark_2020 KRAS Signaling Up 3/200 0.0665 0.4658 3.335 9.0378 TMEM176B;SPRY2;EPHB2 1 MSigDB_Hallmark_2020 Xenobiotic Metabolism 2/200 0.2383 0.6081 2.1877 3.1376 CDO1;TMEM176B 3 MSigDB_Hallmark_2020 Cholesterol Homeostasis 1/74 0.2922 0.6081 2.9532 3.6338 PPARG 4 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.3339 0.6081 2.5052 2.7477 IL17RB 5 MSigDB_Hallmark_2020 Protein Secretion 1/96 0.3614 0.6081 2.2668 2.307 OCRL 6 MSigDB_Hallmark_2020 UV Response Dn 1/144 0.4901 0.6081 1.5023 1.0713 PPARG 7 MSigDB_Hallmark_2020 UV Response Up 1/158 0.5226 0.6081 1.3673 0.8874 CDO1 8 MSigDB_Hallmark_2020 Mitotic Spindle 1/199 0.6063 0.6081 1.082 0.5414 MARCKS 9 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 1/200 0.6081 0.6081 1.0765 0.5354 MARCKS 10 MSigDB_Hallmark_2020 G2-M Checkpoint 1/200 0.6081 0.6081 1.0765 0.5354 MARCKS 10 MSigDB_Hallmark_2020 Adipogenesis 1/200 0.6081 0.6081 1.0765 0.5354 PPARG 10 MSigDB_Hallmark_2020 Estrogen Response Early 1/200 0.6081 0.6081 1.0765 0.5354 IL17RB 10 MSigDB_Hallmark_2020 KRAS Signaling Dn 1/200 0.6081 0.6081 1.0765 0.5354 KCNQ2 10 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
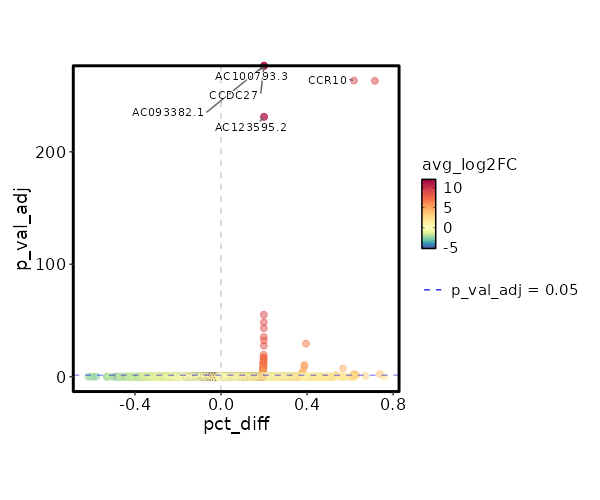
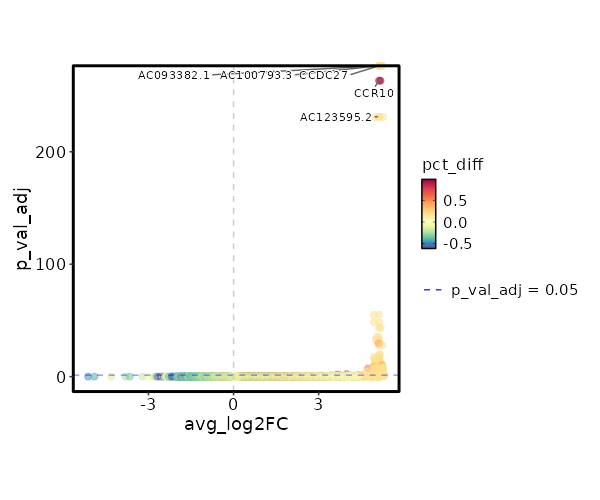
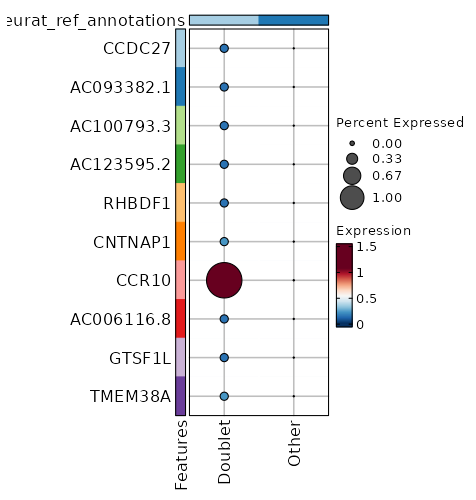
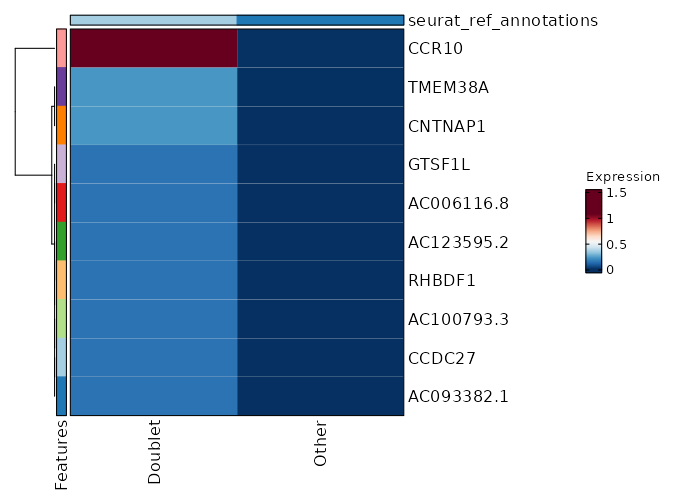
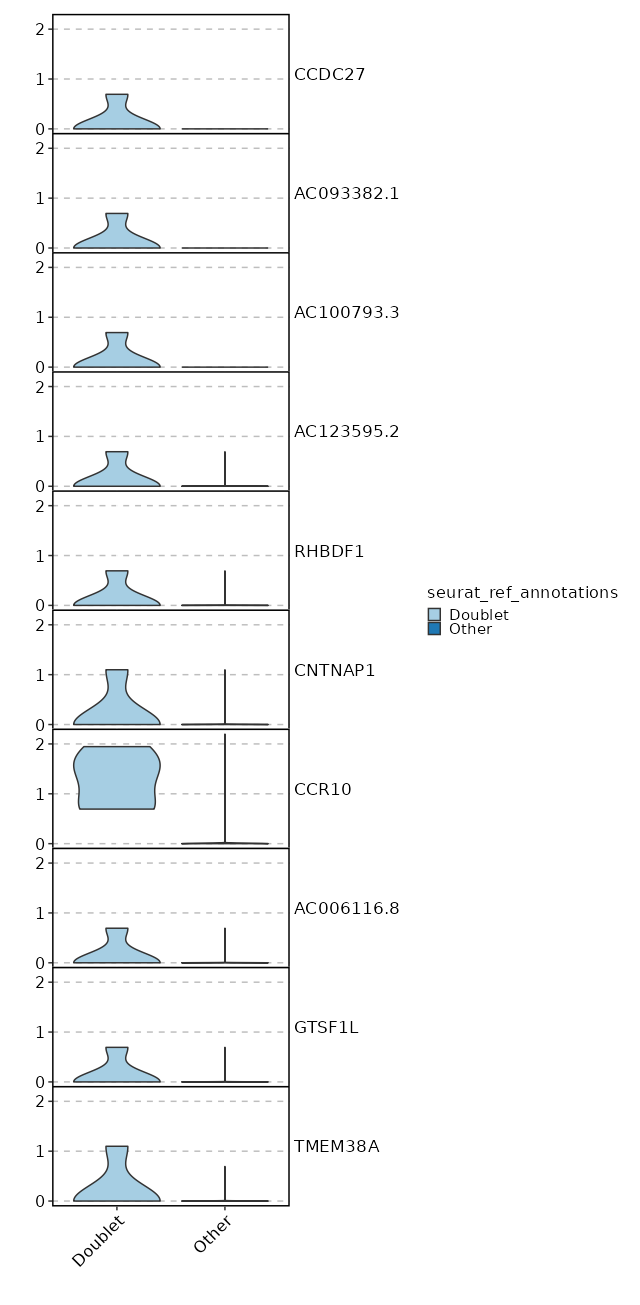
seurat_ref_annotations: Doublet Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
0 12.0749 0.2 0 0 CCDC27 Doublet 0 12.0749 0.2 0 0 AC093382.1 Doublet 0 12.0749 0.2 0 0 AC100793.3 Doublet 1.444e-268 9.2421 1 0.004 4.841e-264 CCR10 Doublet 2.565e-236 11.0749 0.2 0 8.603e-232 AC123595.2 Doublet 2.565e-236 11.0749 0.2 0 8.603e-232 RHBDF1 Doublet 2.629e-60 9.49 0.2 0.001 8.816e-56 CNTNAP1 Doublet 1.049e-53 8.905 0.2 0.001 3.517e-49 AC006116.8 Doublet 1.785e-48 8.753 0.2 0.001 5.986e-44 GTSF1L Doublet 1.268e-40 8.49 0.2 0.001 4.254e-36 AC008115.1 Doublet 1.334e-37 8.3745 0.2 0.001 4.475e-33 C11orf45 Doublet 1.044e-34 6.8785 0.4 0.005 3.503e-30 SLC3A1 Doublet 7.629e-33 8.753 0.2 0.001 2.558e-28 TMEM38A Doublet 6.887e-25 7.753 0.2 0.002 2.31e-20 AC005920.1 Doublet 1.012e-22 7.3201 0.2 0.002 3.392e-18 CCR3 Doublet 8.416e-22 7.5514 0.2 0.002 2.822e-17 DEPDC4 Doublet 6.073e-21 7.49 0.2 0.002 2.037e-16 AC090948.1 Doublet 2.024e-19 7.2676 0.2 0.002 0 SERP2 Doublet 4.071e-18 7.217 0.2 0.003 0 S100A13 Doublet 5.464e-17 7.1681 0.2 0.003 0 PKP3 Doublet
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
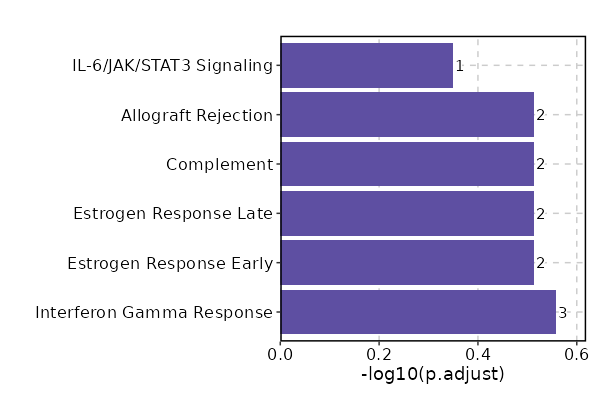
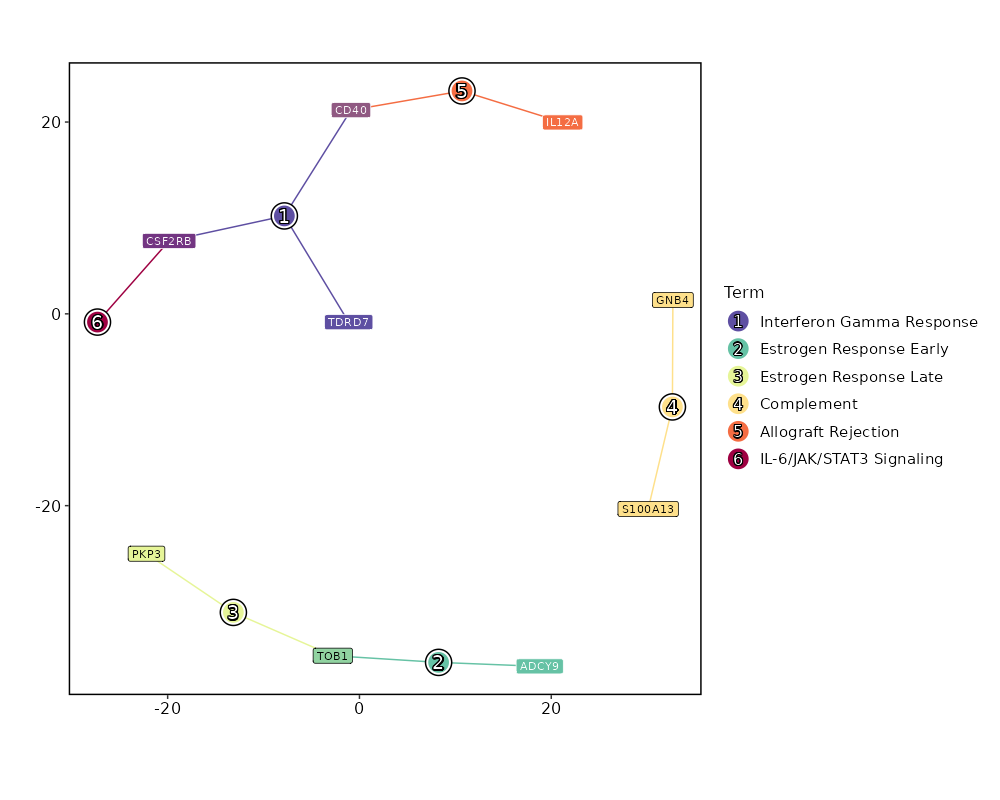
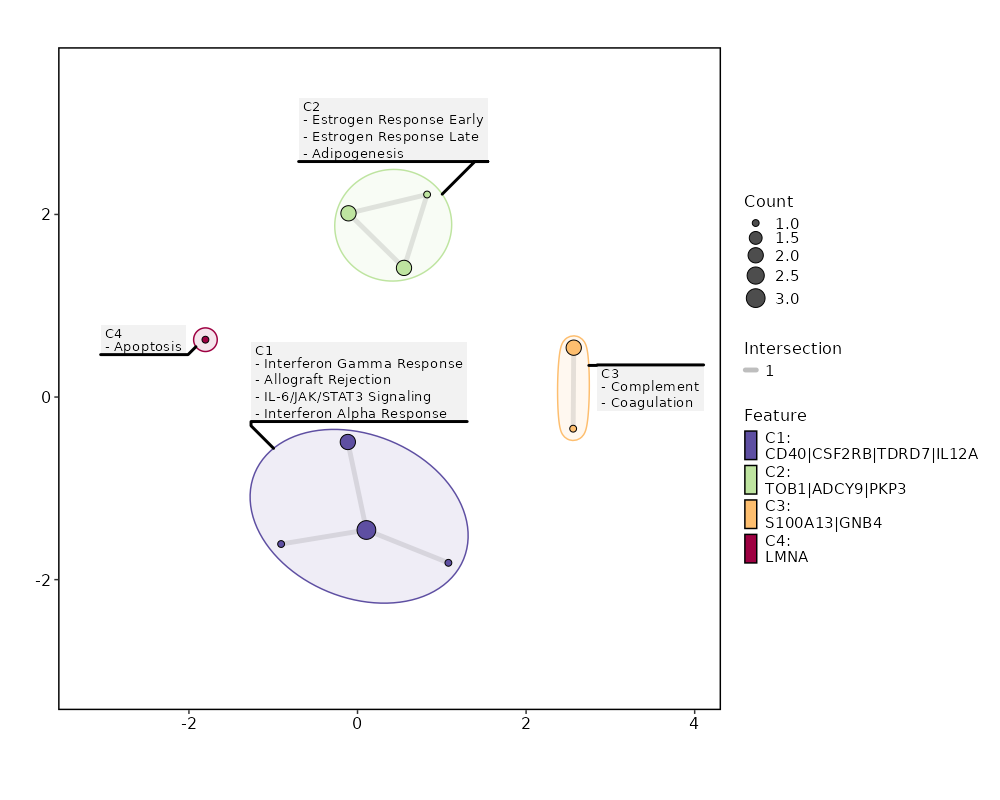
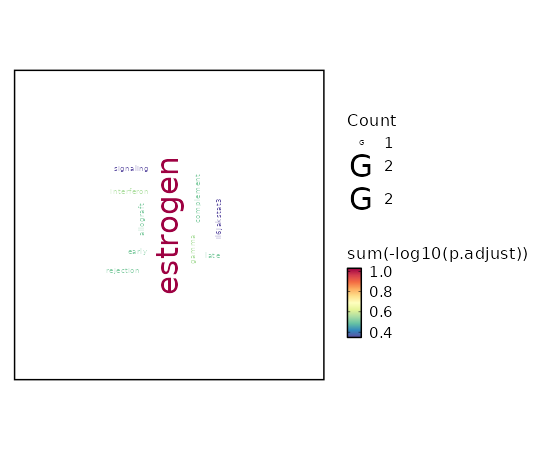
Enrichment Analysis Interferon Gamma Response 3/200 0.0213 0.2764 5.3691 20.6766 CD40;TDRD7;CSF2RB 1 MSigDB_Hallmark_2020 Estrogen Response Early 2/200 0.1177 0.3061 3.4987 7.4847 ADCY9;TOB1 2 MSigDB_Hallmark_2020 Estrogen Response Late 2/200 0.1177 0.3061 3.4987 7.4847 TOB1;PKP3 2 MSigDB_Hallmark_2020 Complement 2/200 0.1177 0.3061 3.4987 7.4847 S100A13;GNB4 2 MSigDB_Hallmark_2020 Allograft Rejection 2/200 0.1177 0.3061 3.4987 7.4847 IL12A;CD40 2 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.2271 0.4478 3.9806 5.9009 CSF2RB 6 MSigDB_Hallmark_2020 Interferon Alpha Response 1/97 0.2497 0.4478 3.5641 4.9454 TDRD7 7 MSigDB_Hallmark_2020 Coagulation 1/138 0.3358 0.4478 2.4923 2.72 S100A13 8 MSigDB_Hallmark_2020 Apoptosis 1/161 0.3797 0.4478 2.1316 2.0641 LMNA 9 MSigDB_Hallmark_2020 Adipogenesis 1/200 0.4478 0.4478 1.7104 1.3742 TOB1 10 MSigDB_Hallmark_2020 Myogenesis 1/200 0.4478 0.4478 1.7104 1.3742 ADCY9 10 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.4478 0.4478 1.7104 1.3742 CD40 10 MSigDB_Hallmark_2020 p53 Pathway 1/200 0.4478 0.4478 1.7104 1.3742 TOB1 10 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
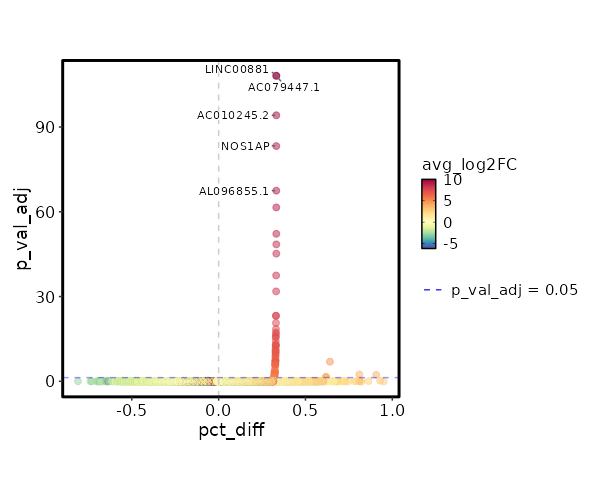
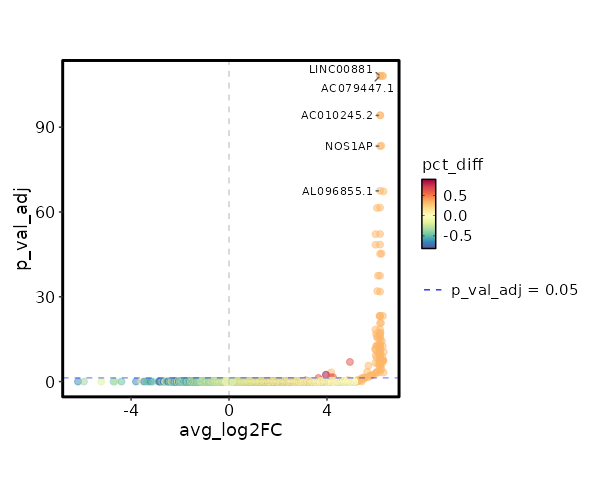
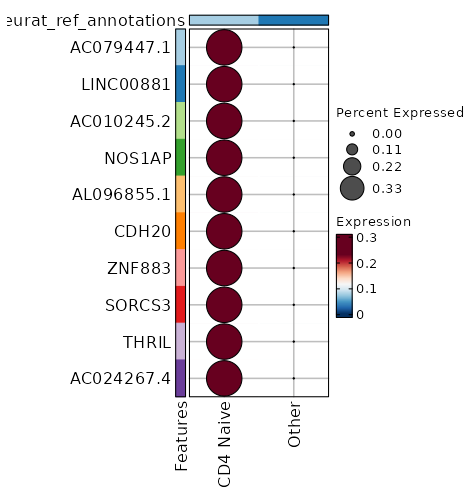
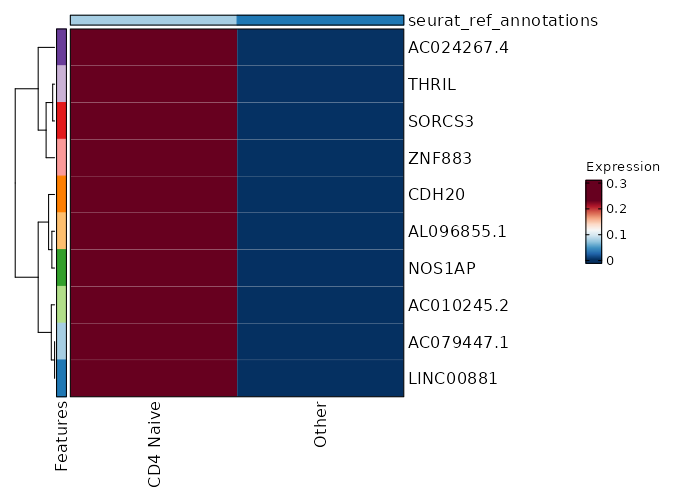
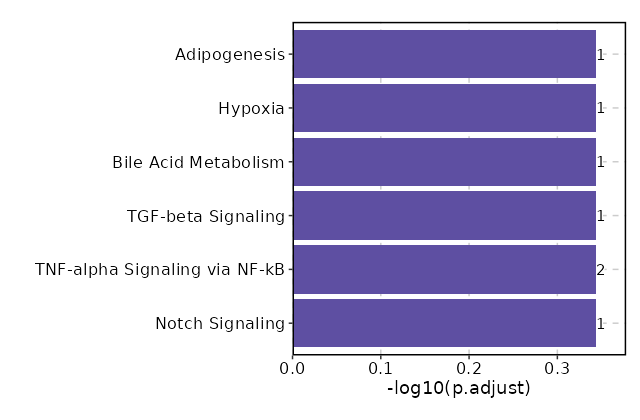
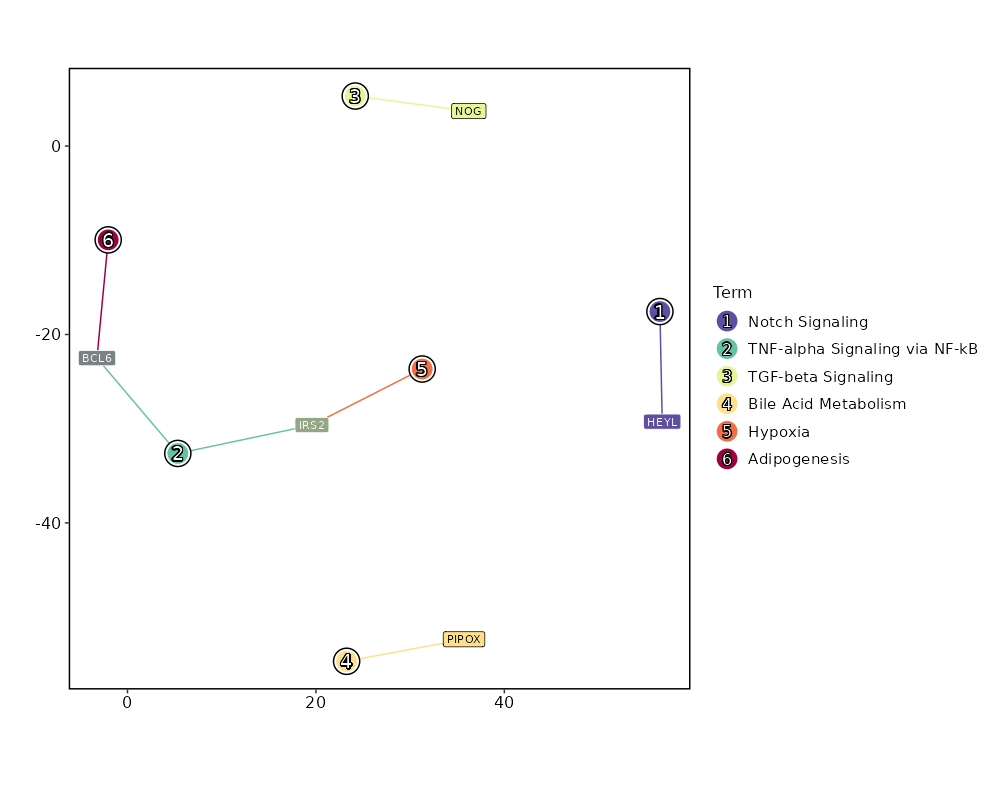
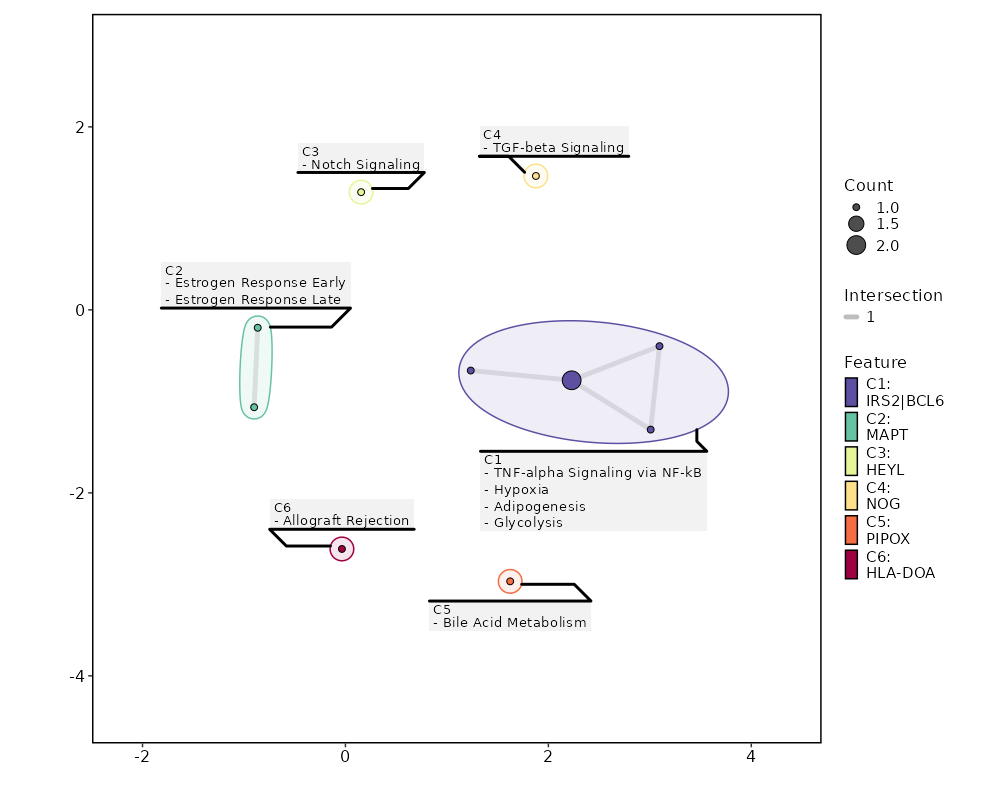
seurat_ref_annotations: CD4 Naive Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
2.116e-113 10.0048 0.333 0.001 7.096e-109 AC079447.1 CD4 Naive 2.116e-113 10.0048 0.333 0.001 7.096e-109 LINC00881 CD4 Naive 2.006e-99 9.8122 0.333 0.001 6.727e-95 AC010245.2 CD4 Naive 1.616e-88 9.3527 0.333 0.001 5.418e-84 NOS1AP CD4 Naive 9.977e-73 9.3527 0.333 0.001 3.346e-68 AL096855.1 CD4 Naive 8.612e-67 9.2272 0.333 0.001 2.888e-62 CDH20 CD4 Naive 1.849e-57 9.0048 0.333 0.001 6.203e-53 ZNF883 CD4 Naive 1.026e-53 8.7247 0.333 0.001 3.44e-49 SORCS3 CD4 Naive 1.901e-50 8.7247 0.333 0.001 6.374e-46 THRIL CD4 Naive 1.035e-42 8.5642 0.333 0.002 3.472e-38 AC024267.4 CD4 Naive 4.494e-37 8.3527 0.333 0.002 1.507e-32 FAM19A1 CD4 Naive 1.915e-28 7.9542 0.333 0.003 6.422e-24 AL035701.1 CD4 Naive 1.937e-28 7.9053 0.333 0.003 6.495e-24 FAM167A CD4 Naive 6.806e-26 7.7678 0.333 0.003 2.282e-21 C5 CD4 Naive 9.133e-24 7.4546 0.333 0.003 3.063e-19 GTSCR1 CD4 Naive 1.439e-22 7.6027 0.333 0.003 4.828e-18 MORN4 CD4 Naive 5.242e-22 7.5642 0.333 0.003 1.758e-17 ADGRG6 CD4 Naive 5.731e-21 7.4902 0.333 0.004 1.922e-16 PIPOX CD4 Naive 5.778e-21 7.4546 0.333 0.004 1.938e-16 RGS12 CD4 Naive 1.379e-19 7.3527 0.333 0.004 0 AP000866.1 CD4 Naive
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
Enrichment Analysis Notch Signaling 1/32 0.0917 0.4533 10.8852 26.0031 HEYL 1 MSigDB_Hallmark_2020 TNF-alpha Signaling via NF-kB 2/200 0.121 0.4533 3.4382 7.2605 IRS2;BCL6 2 MSigDB_Hallmark_2020 TGF-beta Signaling 1/54 0.1499 0.4533 6.3598 12.0674 NOG 3 MSigDB_Hallmark_2020 Bile Acid Metabolism 1/112 0.2864 0.4533 3.0278 3.7858 PIPOX 4 MSigDB_Hallmark_2020 Hypoxia 1/200 0.4533 0.4533 1.6814 1.3302 IRS2 5 MSigDB_Hallmark_2020 Adipogenesis 1/200 0.4533 0.4533 1.6814 1.3302 BCL6 5 MSigDB_Hallmark_2020 Estrogen Response Early 1/200 0.4533 0.4533 1.6814 1.3302 MAPT 5 MSigDB_Hallmark_2020 Estrogen Response Late 1/200 0.4533 0.4533 1.6814 1.3302 MAPT 5 MSigDB_Hallmark_2020 Glycolysis 1/200 0.4533 0.4533 1.6814 1.3302 IRS2 5 MSigDB_Hallmark_2020 Allograft Rejection 1/200 0.4533 0.4533 1.6814 1.3302 HLA-DOA 5 MSigDB_Hallmark_2020 KRAS Signaling Dn 1/200 0.4533 0.4533 1.6814 1.3302 C5 5 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
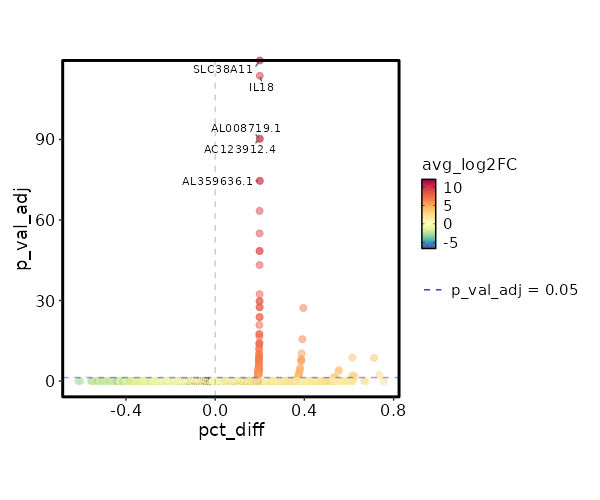
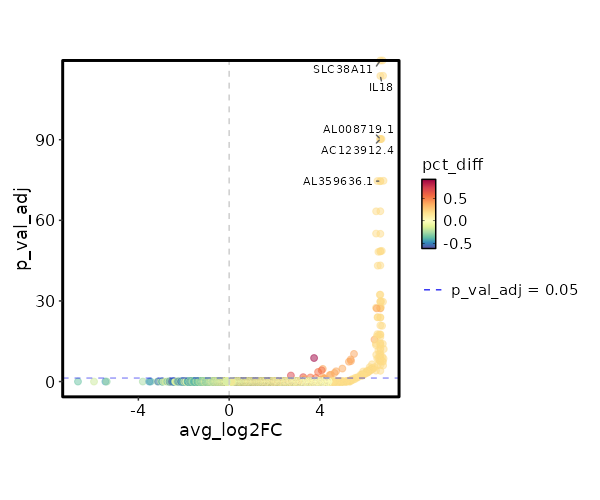
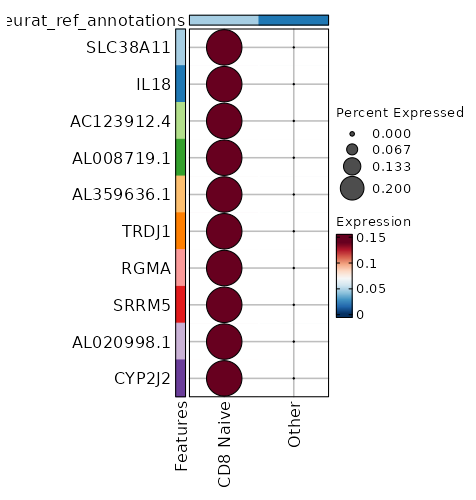
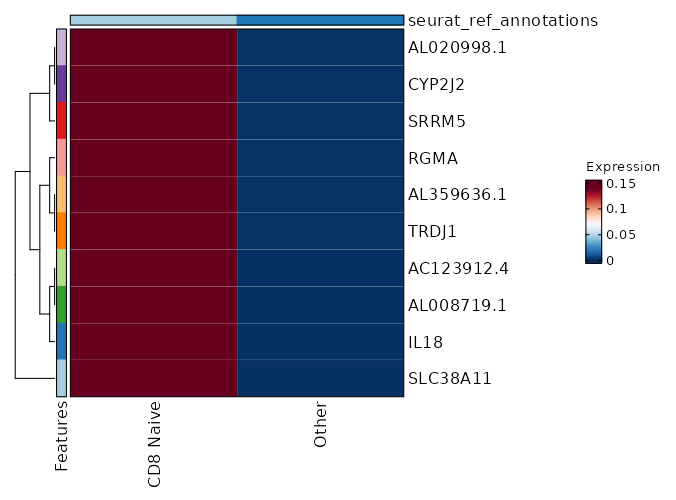
seurat_ref_annotations: CD8 Naive Markers Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05 & avg_log2FC > 0'.
0 12.0749 0.2 0 0 SLC38A11 CD8 Naive 5.534e-119 10.0749 0.2 0 1.856e-114 IL18 CD8 Naive 1.691e-95 9.753 0.2 0 5.671e-91 AC123912.4 CD8 Naive 1.691e-95 9.753 0.2 0 5.671e-91 AL008719.1 CD8 Naive 7.799e-80 9.49 0.2 0 2.616e-75 AL359636.1 CD8 Naive 7.799e-80 9.49 0.2 0 2.616e-75 TRDJ1 CD8 Naive 1.218e-68 9.2676 0.2 0.001 4.086e-64 RGMA CD8 Naive 3.055e-60 9.0749 0.2 0.001 1.024e-55 SRRM5 CD8 Naive 1.049e-53 8.905 0.2 0.001 3.517e-49 AL020998.1 CD8 Naive 1.049e-53 8.905 0.2 0.001 3.517e-49 CYP2J2 CD8 Naive 1.785e-48 8.753 0.2 0.001 5.986e-44 JMJD1C-AS1 CD8 Naive 1.334e-37 8.3745 0.2 0.001 4.475e-33 TSPAN33 CD8 Naive 5.209e-35 8.2676 0.2 0.001 1.747e-30 NPBWR1 CD8 Naive 5.209e-35 8.2676 0.2 0.001 1.747e-30 FGF11 CD8 Naive 9.198e-33 8.1681 0.2 0.001 3.085e-28 ANKRD53 CD8 Naive 9.198e-33 8.1681 0.2 0.001 3.085e-28 SLC4A11 CD8 Naive 1.846e-32 6.7773 0.4 0.005 6.189e-28 NANOS3 CD8 Naive 4.081e-29 7.7895 0.2 0.001 1.369e-24 TRBV5-3 CD8 Naive 4.646e-29 7.9875 0.2 0.001 1.558e-24 PROSER2 CD8 Naive 3.921e-26 7.827 0.2 0.002 1.315e-21 AP001496.1 CD8 Naive
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
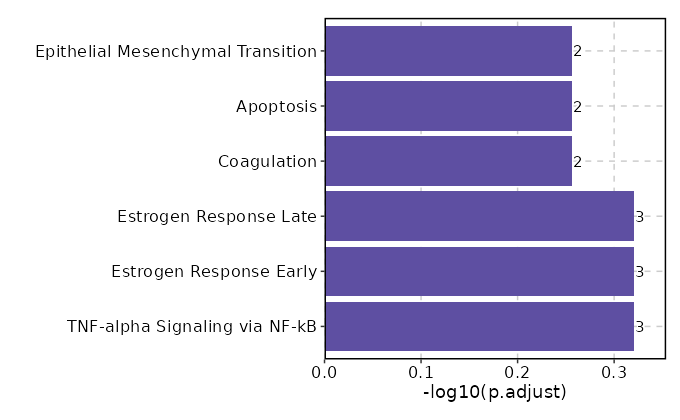
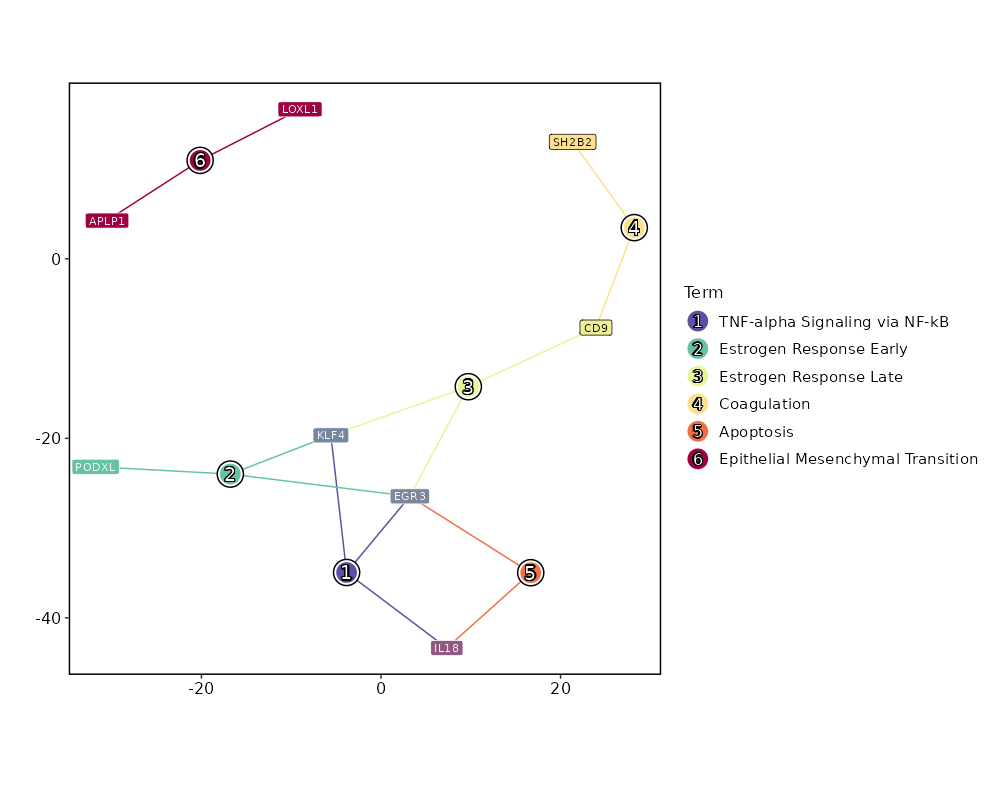
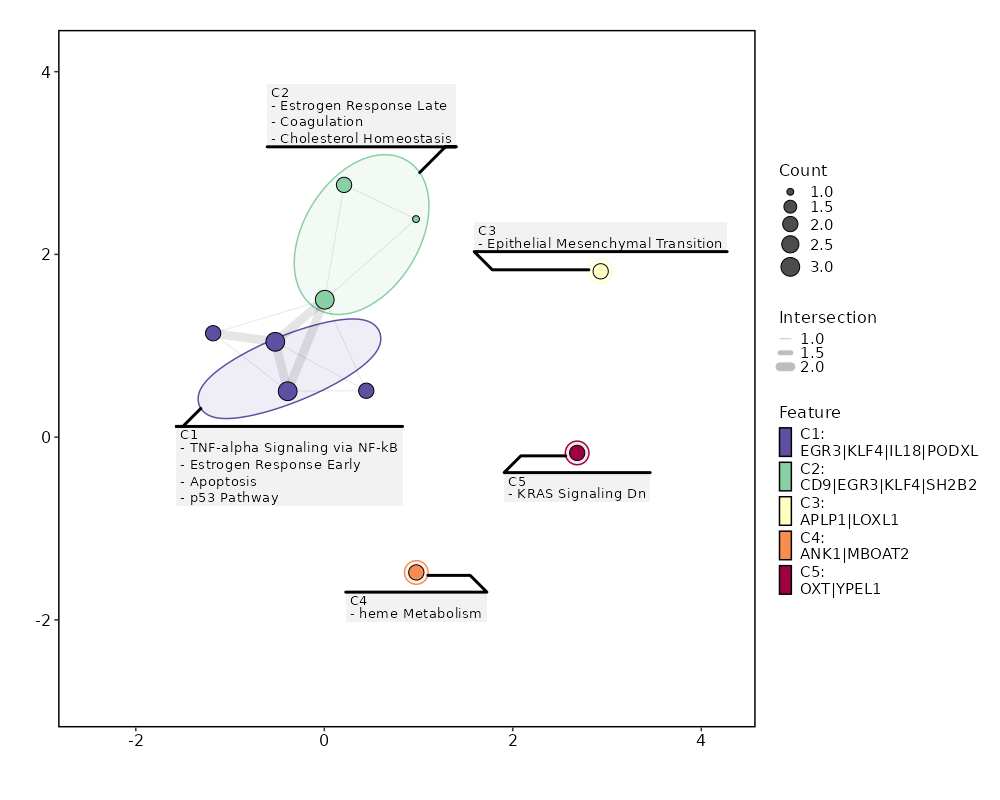
Enrichment Analysis TNF-alpha Signaling via NF-kB 3/200 0.0717 0.4782 3.227 8.5028 EGR3;IL18;KLF4 1 MSigDB_Hallmark_2020 Estrogen Response Early 3/200 0.0717 0.4782 3.227 8.5028 EGR3;PODXL;KLF4 1 MSigDB_Hallmark_2020 Estrogen Response Late 3/200 0.0717 0.4782 3.227 8.5028 CD9;EGR3;KLF4 1 MSigDB_Hallmark_2020 Coagulation 2/138 0.1422 0.5542 3.0926 6.0331 CD9;SH2B2 4 MSigDB_Hallmark_2020 Apoptosis 2/161 0.1809 0.5542 2.6422 4.5172 EGR3;IL18 5 MSigDB_Hallmark_2020 Epithelial Mesenchymal Transition 2/200 0.2494 0.5542 2.1176 2.9406 APLP1;LOXL1 6 MSigDB_Hallmark_2020 p53 Pathway 2/200 0.2494 0.5542 2.1176 2.9406 KLF4;ITGB4 6 MSigDB_Hallmark_2020 heme Metabolism 2/200 0.2494 0.5542 2.1176 2.9406 ANK1;MBOAT2 6 MSigDB_Hallmark_2020 KRAS Signaling Dn 2/200 0.2494 0.5542 2.1176 2.9406 YPEL1;OXT 6 MSigDB_Hallmark_2020 Cholesterol Homeostasis 1/74 0.3 0.6 2.8596 3.4426 CD9 10 MSigDB_Hallmark_2020 IL-6/JAK/STAT3 Signaling 1/87 0.3426 0.6177 2.4257 2.5982 CD9 11 MSigDB_Hallmark_2020 Protein Secretion 1/96 0.3706 0.6177 2.1949 2.1786 OCRL 12 MSigDB_Hallmark_2020 IL-2/STAT5 Signaling 1/199 0.618 0.6198 1.0476 0.5042 APLP1 13 MSigDB_Hallmark_2020 Myogenesis 1/200 0.6198 0.6198 1.0423 0.4985 ITGB4 14 MSigDB_Hallmark_2020 Apical Junction 1/200 0.6198 0.6198 1.0423 0.4985 ITGB4 14 MSigDB_Hallmark_2020 mTORC1 Signaling 1/200 0.6198 0.6198 1.0423 0.4985 CD9 14 MSigDB_Hallmark_2020 Inflammatory Response 1/200 0.6198 0.6198 1.0423 0.4985 IL18 14 MSigDB_Hallmark_2020 Xenobiotic Metabolism 1/200 0.6198 0.6198 1.0423 0.4985 CYP2J2 14 MSigDB_Hallmark_2020 Allograft Rejection 1/200 0.6198 0.6198 1.0423 0.4985 IL18 14 MSigDB_Hallmark_2020 KRAS Signaling Up 1/200 0.6198 0.6198 1.0423 0.4985 KLF4 14 MSigDB_Hallmark_2020
You must use "Save as" or "Save link as" from the context menu (by right-clicking the button) to download the entire data.
The bar plot shows the top enriched terms in database 'MSigDB_Hallmark_2020', the x-axis shows the -log10 of the adjusted p-values, and the y-axis shows the term names. The number next to each bar indicates the overlap gene count.
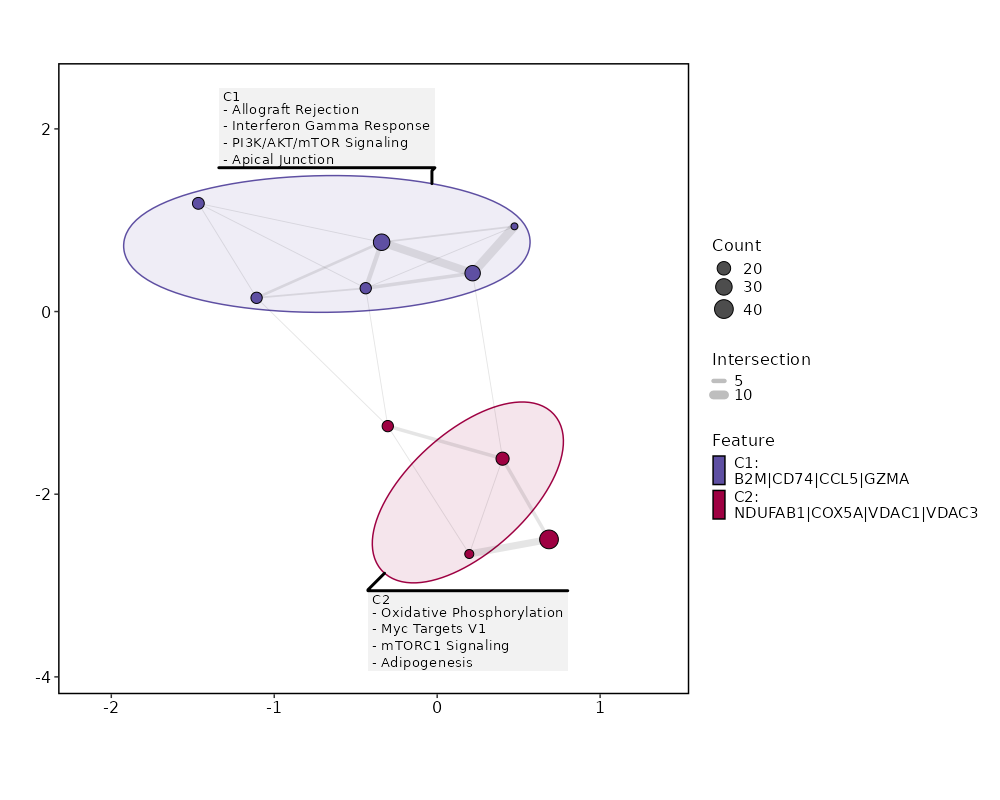
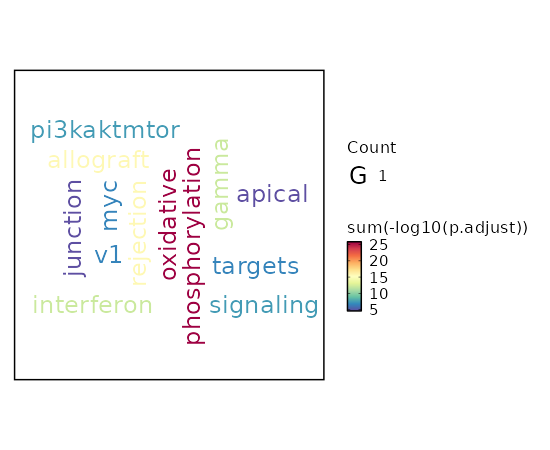
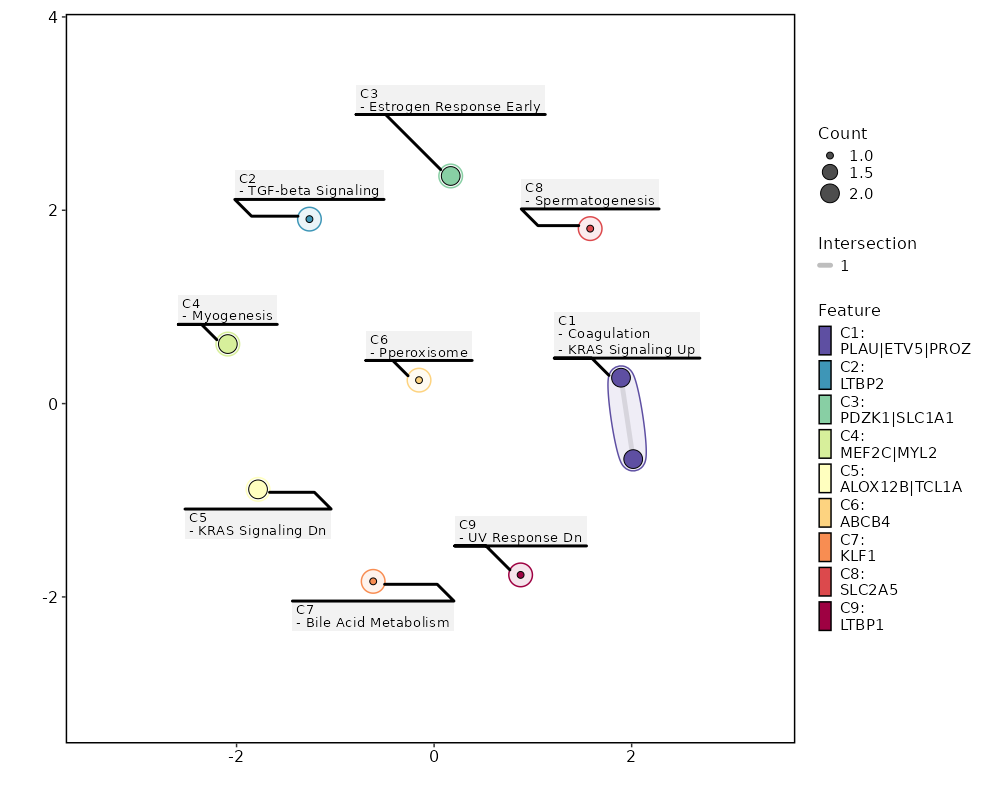
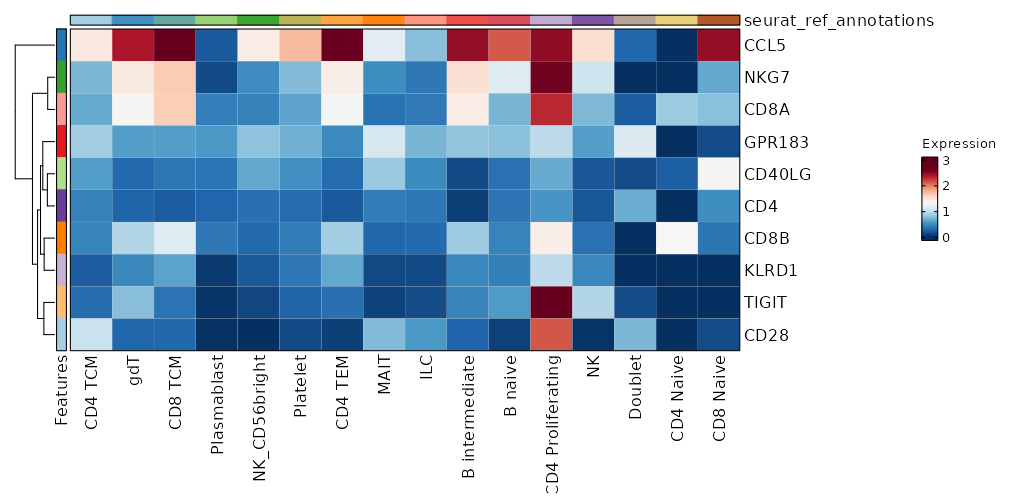
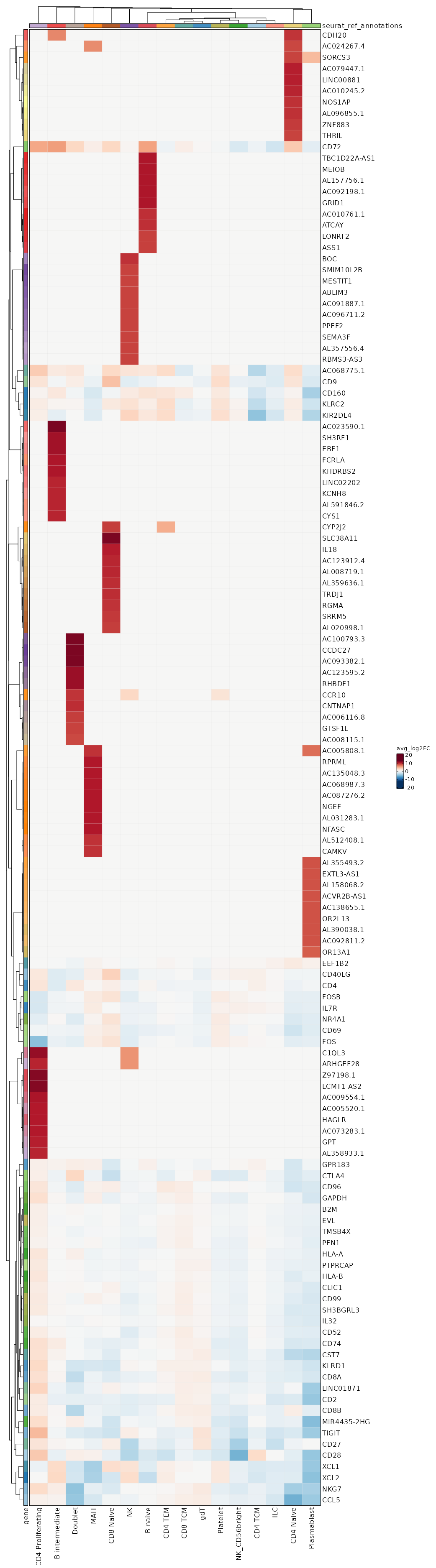
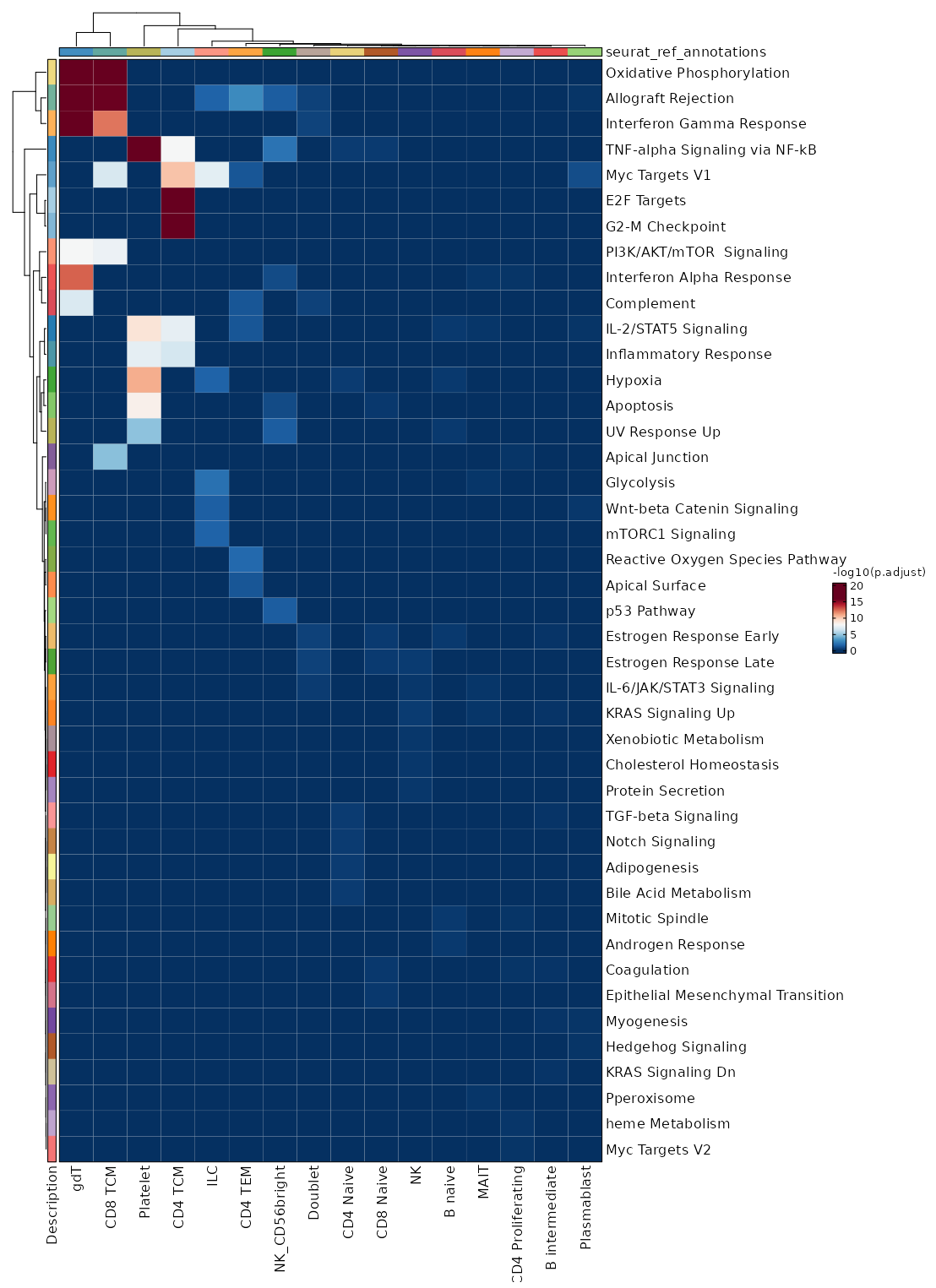
seurat_ref_annotations (All Markers) seurat_ref_annotations (All Enrichments) Heatmap of enriched terms of all clusters