c1
Diagnosis: Colitis
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
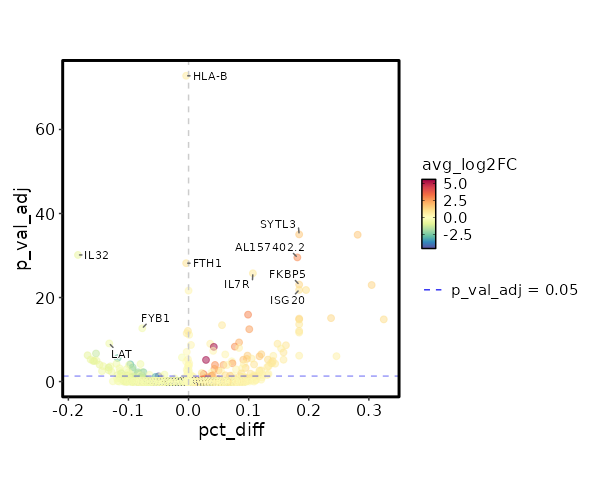
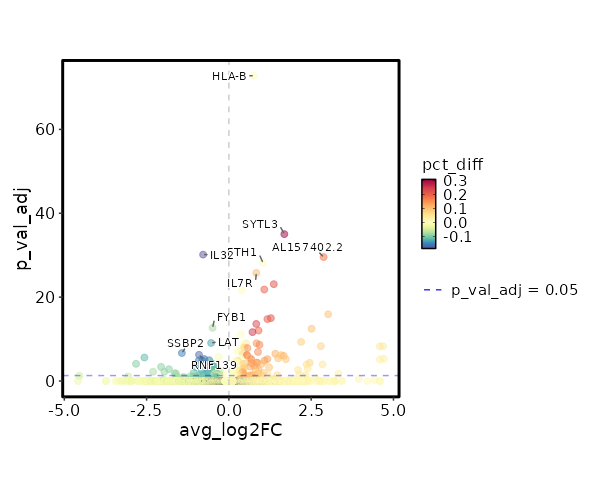
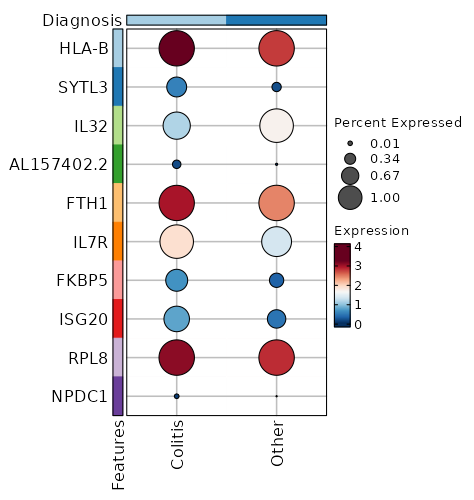
| 4.874e-78 | 0.7561 | 0.995 | 0.999 | 1.635e-73 | HLA-B | Colitis |
| 2.796e-40 | 1.6848 | 0.55 | 0.243 | 9.376e-36 | SYTL3 | Colitis |
| 2.181e-35 | -0.7834 | 0.761 | 0.945 | 7.315e-31 | IL32 | Colitis |
| 8.329e-35 | 2.8782 | 0.218 | 0.037 | 2.793e-30 | AL157402.2 | Colitis |
| 1.969e-33 | 1.037 | 0.995 | 0.999 | 6.605e-29 | FTH1 | Colitis |
| 5.041e-31 | 0.8329 | 0.944 | 0.837 | 1.691e-26 | IL7R | Colitis |
| 2.413e-28 | 1.3644 | 0.608 | 0.386 | 8.093e-24 | FKBP5 | Colitis |
| 4.52e-27 | 1.0756 | 0.712 | 0.506 | 1.516e-22 | ISG20 | Colitis |
| 6.723e-27 | 0.3937 | 1 | 1 | 2.255e-22 | RPL8 | Colitis |
| 3.581e-21 | 3.0168 | 0.113 | 0.014 | 1.201e-16 | NPDC1 | Colitis |
| 3.075e-20 | 1.2753 | 0.574 | 0.353 | 0 | CREM | Colitis |
| 5.132e-20 | 1.1695 | 0.518 | 0.31 | 0 | PLP2 | Colitis |
| 7.244e-19 | 0.8321 | 0.608 | 0.371 | 0 | CD55 | Colitis |
| 6.118e-18 | -0.4955 | 0.863 | 0.94 | 0 | FYB1 | Colitis |
| 1.028e-17 | 2.512 | 0.126 | 0.025 | 0 | HLA-DRB1 | Colitis |
| 2.743e-17 | 0.901 | 0.624 | 0.427 | 0 | LITAF | Colitis |
| 6.506e-17 | 0.7175 | 0.748 | 0.492 | 0 | KLF2 | Colitis |
| 2.281e-16 | 0.3586 | 0.995 | 0.994 | 0 | BTG1 | Colitis |
| 0 | 2.1948 | 0.104 | 0.02 | 0 | ZBTB16 | Colitis |
| 0 | -0.5401 | 0.718 | 0.85 | 0 | LAT | Colitis |
Downloading entire data
Enrichment Analysis
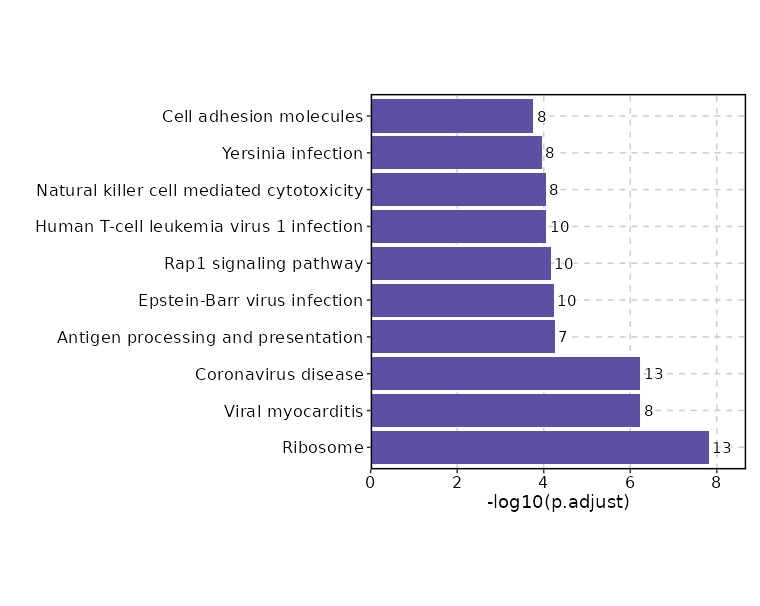
| Ribosome | 13/158 | 0 | 0 | 13.9177 | 322.6026 | RPS9;RPS6;RPL28;RPL10;RPL14;RPS27;RPS26;RPS29;RPL3;RPL8;RPS19;MRPL23;RPLP0 | 1 | KEGG_2021_Human |
| Viral myocarditis | 8/60 | 0 | 0 | 23.0862 | 428.0224 | HLA-DQB1;ACTB;HLA-A;HLA-B;HLA-DRB1;ITGB2;ITGAL;CD55 | 2 | KEGG_2021_Human |
| Coronavirus disease | 13/232 | 0 | 0 | 9.1803 | 169.4655 | RPL28;RPL10;RPL14;RPS27;RPS26;RPS29;PIK3R1;RPL3;RPL8;RPS19;RPLP0;RPS9;RPS6 | 3 | KEGG_2021_Human |
| Antigen processing and presentation | 7/78 | 0 | 0.0001 | 14.6694 | 199.5508 | HLA-DQB1;B2M;HLA-A;HLA-B;PSME2;PSME1;HLA-DRB1 | 4 | KEGG_2021_Human |
| Epstein-Barr virus infection | 10/202 | 0 | 0.0001 | 7.8798 | 105.0454 | VIM;B2M;HLA-A;HLA-B;PIK3R1;HLA-DRB1;HLA-DQB1;CCND3;TNFAIP3;ITGAL | 5 | KEGG_2021_Human |
| Rap1 signaling pathway | 10/210 | 0 | 0.0001 | 7.5615 | 98.1575 | ACTB;FYB1;PIK3R1;RASGRP2;LCP2;LAT;EVL;ITGB2;ITGB1;ITGAL | 6 | KEGG_2021_Human |
| Human T-cell leukemia virus 1 infection | 10/219 | 0 | 0.0001 | 7.2326 | 91.1721 | B2M;HLA-A;HLA-B;PIK3R1;HLA-DRB1;HLA-DQB1;CCND3;LCK;ITGB2;ITGAL | 7 | KEGG_2021_Human |
| Natural killer cell mediated cytotoxicity | 8/131 | 0 | 0.0001 | 9.7251 | 120.9853 | HLA-A;HLA-B;PIK3R1;LCP2;LAT;LCK;ITGB2;ITGAL | 8 | KEGG_2021_Human |
| Yersinia infection | 8/137 | 0 | 0.0001 | 9.2699 | 112.2377 | ACTB;FYB1;PIK3R1;PXN;LCP2;LAT;LCK;ITGB1 | 9 | KEGG_2021_Human |
| Cell adhesion molecules | 8/148 | 0 | 0.0002 | 8.5368 | 98.5027 | HLA-A;HLA-B;SELL;HLA-DRB1;HLA-DQB1;ITGB2;ITGB1;ITGAL | 10 | KEGG_2021_Human |
| Phagosome | 8/152 | 0 | 0.0002 | 8.298 | 94.1284 | ACTB;HLA-A;HLA-B;HLA-DRB1;TUBB;HLA-DQB1;ITGB2;ITGB1 | 11 | KEGG_2021_Human |
| Leukocyte transendothelial migration | 7/114 | 0 | 0.0002 | 9.7162 | 107.5264 | ACTB;PIK3R1;PXN;ITGB2;ITGB1;ITGAL;CXCR4 | 12 | KEGG_2021_Human |
| Human immunodeficiency virus 1 infection | 8/212 | 0.0001 | 0.0018 | 5.8396 | 52.4344 | B2M;HLA-A;HLA-B;PIK3R1;CXCR4;PXN;TNFRSF1B;SAMHD1 | 13 | KEGG_2021_Human |
| Allograft rejection | 4/38 | 0.0001 | 0.0018 | 17.1505 | 152.0548 | HLA-A;HLA-B;HLA-DQB1;HLA-DRB1 | 14 | KEGG_2021_Human |
| Graft-versus-host disease | 4/42 | 0.0002 | 0.0025 | 15.3421 | 129.967 | HLA-A;HLA-B;HLA-DQB1;HLA-DRB1 | 15 | KEGG_2021_Human |
| Type I diabetes mellitus | 4/43 | 0.0002 | 0.0026 | 14.948 | 125.2496 | HLA-A;HLA-B;HLA-DQB1;HLA-DRB1 | 16 | KEGG_2021_Human |
| Th1 and Th2 cell differentiation | 5/92 | 0.0005 | 0.005 | 8.4176 | 64.4641 | HLA-DQB1;LAT;STAT4;LCK;HLA-DRB1 | 17 | KEGG_2021_Human |
| Autoimmune thyroid disease | 4/53 | 0.0005 | 0.0052 | 11.8914 | 90.0066 | HLA-DQB1;HLA-A;HLA-B;HLA-DRB1 | 18 | KEGG_2021_Human |
| Regulation of actin cytoskeleton | 7/218 | 0.0009 | 0.0083 | 4.9012 | 34.5029 | ACTB;PIK3R1;CXCR4;PXN;ITGB2;ITGB1;ITGAL | 19 | KEGG_2021_Human |
| Human cytomegalovirus infection | 7/225 | 0.0011 | 0.0095 | 4.7422 | 32.5107 | B2M;HLA-A;HLA-B;PIK3R1;PTGER2;CXCR4;PXN | 20 | KEGG_2021_Human |
Downloading entire data
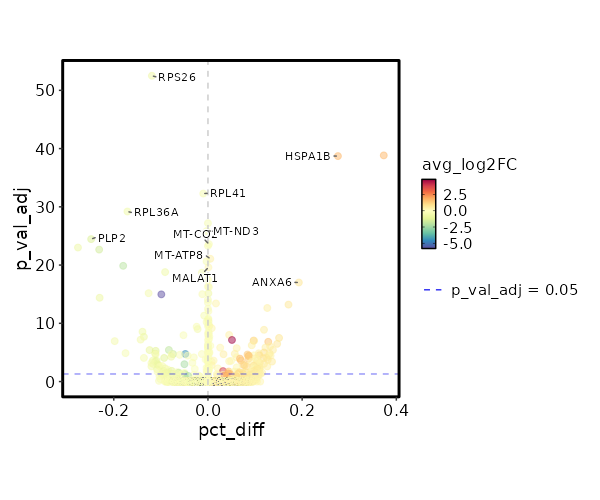
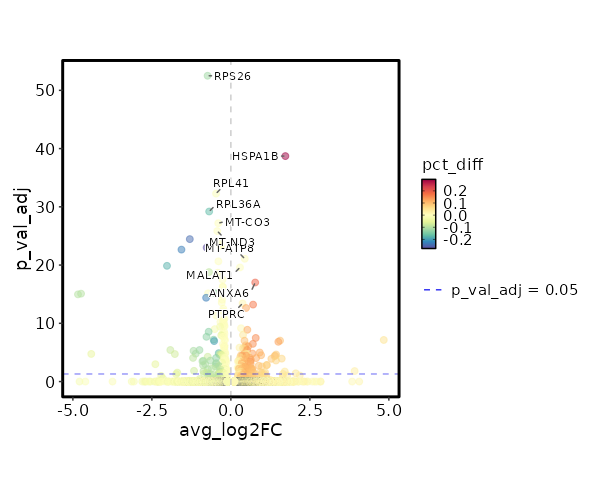
Diagnosis: Control
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
| 9.389e-58 | -0.7363 | 0.879 | 0.998 | 3.149e-53 | RPS26 | Control |
| 5.799e-44 | 1.7253 | 0.45 | 0.155 | 1.945e-39 | HSPA1B | Control |
| 1.574e-37 | -0.4683 | 0.985 | 0.995 | 5.278e-33 | RPL41 | Control |
| 1.898e-34 | -0.687 | 0.74 | 0.911 | 6.366e-30 | RPL36A | Control |
| 1.915e-32 | -0.3988 | 0.996 | 0.996 | 6.424e-28 | MT-CO3 | Control |
| 4.307e-31 | -0.4298 | 0.987 | 0.99 | 1.444e-26 | MT-ND3 | Control |
| 1.044e-29 | -1.2998 | 0.226 | 0.474 | 3.501e-25 | PLP2 | Control |
| 7.529e-29 | -0.3393 | 0.999 | 0.997 | 2.525e-24 | MT-CO2 | Control |
| 1.499e-28 | -0.3124 | 1 | 1 | 5.027e-24 | RPL28 | Control |
| 2.926e-28 | -0.7664 | 0.417 | 0.693 | 9.814e-24 | RPS4Y1 | Control |
| 6.621e-28 | -1.5638 | 0.192 | 0.423 | 2.221e-23 | MT2A | Control |
| 2.475e-26 | 0.4397 | 0.996 | 0.991 | 8.301e-22 | MT-ATP8 | Control |
| 6.777e-26 | -0.3946 | 0.994 | 0.997 | 2.273e-21 | MT-ATP6 | Control |
| 4.022e-25 | -2.0217 | 0.068 | 0.248 | 1.349e-20 | KANSL1-AS1 | Control |
| 7.505e-25 | 0.2888 | 1 | 0.999 | 2.517e-20 | MALAT1 | Control |
| 4.755e-24 | -0.6925 | 0.837 | 0.928 | 1.595e-19 | TSC22D3 | Control |
| 6.095e-24 | -0.4126 | 0.983 | 0.996 | 2.044e-19 | TMSB10 | Control |
| 9.591e-23 | -0.2642 | 1 | 1 | 3.217e-18 | RPS27 | Control |
| 2.935e-22 | 0.7702 | 0.669 | 0.476 | 9.844e-18 | ANXA6 | Control |
| 1.48e-21 | -0.2565 | 1 | 1 | 4.963e-17 | RPS28 | Control |
Downloading entire data
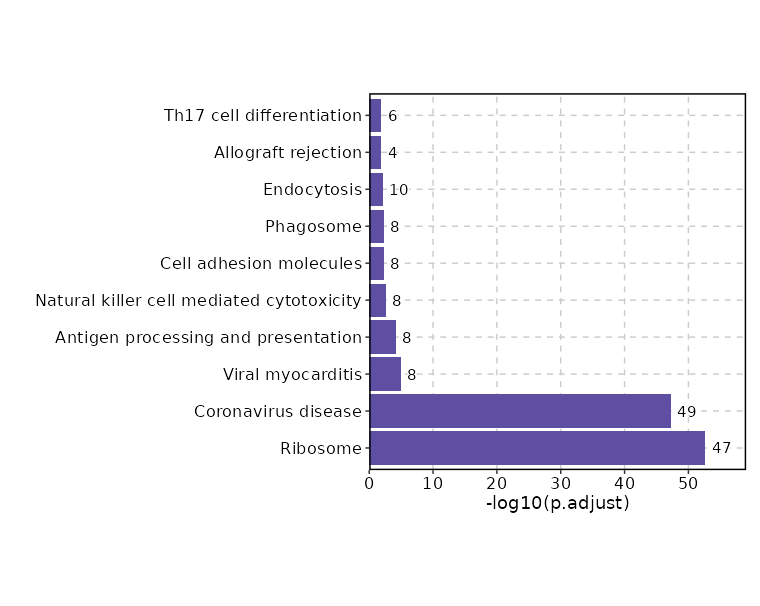
Enrichment Analysis
| Ribosome | 47/158 | 1.374e-55 | 2.185e-53 | 50.1884 | 6340.028 | RPL22;RPS8;RPS9;RPL24;RPS6;FAU;RPL26;RPL28;RPL29;RPL10;RPL12;RPS4Y1;RPL36AL;RPL15;RPL14;RPS2;RPL18;RPL41;RPS27;RPS26;RPS29;RPS28;RPS21;RPS23;RPS25;RPS24;RPL6;RPL3;RPL31;RPL30;RPL34;RPL8;RPS15;RPS18;RPS19;RPL37;RPL36;RPL39;RPL37A;RPL36A;RPS15A;RPL35A;RPL23A;RPLP0;RPLP1;RPL18A;RPL7A | 1 | KEGG_2021_Human |
| Coronavirus disease | 49/232 | 7.378e-50 | 5.865e-48 | 32.0071 | 3620.9853 | RPL22;RPL24;RPL26;RPL28;RPL29;RPL10;RPL12;RPS4Y1;RPL36AL;RPL15;RPL14;RPL18;RPL41;RPS27;RPS26;RPS29;RPS28;RPS21;RPS23;RPS25;RPS24;RPL6;RPL3;RPL31;RPL30;RPL34;RPL8;RPS15;RPS18;RPS19;RPL37;RPL36;RPL39;RPS15A;STAT1;RPLP0;RPLP1;RPL18A;RPS8;RPS9;RPS6;FAU;RPS2;NFKBIA;RPL37A;RPL36A;RPL35A;RPL23A;RPL7A | 2 | KEGG_2021_Human |
| Viral myocarditis | 8/60 | 0 | 0 | 14.8105 | 226.3297 | ACTB;HLA-C;HLA-A;HLA-B;HLA-G;ITGB2;ITGAL;CD55 | 3 | KEGG_2021_Human |
| Antigen processing and presentation | 8/78 | 0 | 0.0001 | 10.9921 | 145.4303 | HSP90AA1;HSPA1A;HSPA1B;HLA-C;HLA-A;HLA-B;HLA-G;HSP90AB1 | 4 | KEGG_2021_Human |
| Natural killer cell mediated cytotoxicity | 8/131 | 0.0001 | 0.0026 | 6.2388 | 58.7137 | HLA-C;HLA-A;HLA-B;HLA-G;LCP2;LCK;ITGB2;ITGAL | 5 | KEGG_2021_Human |
| Cell adhesion molecules | 8/148 | 0.0002 | 0.0051 | 5.4765 | 46.8974 | HLA-C;HLA-A;HLA-B;HLA-G;SELL;PTPRC;ITGB2;ITGAL | 6 | KEGG_2021_Human |
| Phagosome | 8/152 | 0.0002 | 0.0052 | 5.3233 | 44.6149 | ACTB;HLA-C;HLA-A;HLA-B;HLA-G;TUBB;ITGB2;TUBA1B | 7 | KEGG_2021_Human |
| Endocytosis | 10/252 | 0.0004 | 0.0077 | 3.9785 | 31.2642 | HSPA1A;HSPA1B;HLA-C;HLA-A;HLA-B;HLA-G;CAPZA1;SH3KBP1;SMAP2;WIPF1 | 8 | KEGG_2021_Human |
| Allograft rejection | 4/38 | 0.0007 | 0.0123 | 11.1191 | 80.852 | HLA-C;HLA-A;HLA-B;HLA-G | 9 | KEGG_2021_Human |
| Th17 cell differentiation | 6/107 | 0.001 | 0.014 | 5.6496 | 38.9916 | PRKCQ;HSP90AA1;NFKBIA;STAT1;LCK;HSP90AB1 | 10 | KEGG_2021_Human |
| Graft-versus-host disease | 4/42 | 0.001 | 0.014 | 9.9466 | 68.5159 | HLA-C;HLA-A;HLA-B;HLA-G | 11 | KEGG_2021_Human |
| Type I diabetes mellitus | 4/43 | 0.0011 | 0.014 | 9.6911 | 65.8899 | HLA-C;HLA-A;HLA-B;HLA-G | 12 | KEGG_2021_Human |
| Spliceosome | 7/150 | 0.0011 | 0.014 | 4.6679 | 31.6083 | HNRNPK;HSPA1A;HSPA1B;SRSF2;HNRNPU;SNRPD2;FUS | 13 | KEGG_2021_Human |
| Epstein-Barr virus infection | 8/202 | 0.0015 | 0.0169 | 3.9413 | 25.6576 | HLA-C;HLA-A;HLA-B;HLA-G;STAT1;NFKBIA;TNFAIP3;ITGAL | 14 | KEGG_2021_Human |
| Autoimmune thyroid disease | 4/53 | 0.0024 | 0.0245 | 7.7094 | 46.3932 | HLA-C;HLA-A;HLA-B;HLA-G | 15 | KEGG_2021_Human |
| Human T-cell leukemia virus 1 infection | 8/219 | 0.0025 | 0.0245 | 3.6206 | 21.7427 | HLA-C;HLA-A;HLA-B;HLA-G;NFKBIA;LCK;ITGB2;ITGAL | 16 | KEGG_2021_Human |
| Legionellosis | 4/57 | 0.0032 | 0.0297 | 7.1261 | 40.9854 | NFKBIA;HSPA1A;HSPA1B;ITGB2 | 17 | KEGG_2021_Human |
| Yersinia infection | 6/137 | 0.0035 | 0.0311 | 4.3492 | 24.5701 | ACTB;FYB1;NFKBIA;LCP2;LCK;WIPF1 | 18 | KEGG_2021_Human |
| Kaposi sarcoma-associated herpesvirus infection | 7/193 | 0.0047 | 0.0395 | 3.5809 | 19.1813 | HLA-C;HLA-A;HLA-B;HLA-G;STAT1;UBC;NFKBIA | 19 | KEGG_2021_Human |
| T cell receptor signaling pathway | 5/104 | 0.0051 | 0.0408 | 4.7805 | 25.2092 | PRKCQ;NFKBIA;PTPRC;LCP2;LCK | 20 | KEGG_2021_Human |
Downloading entire data
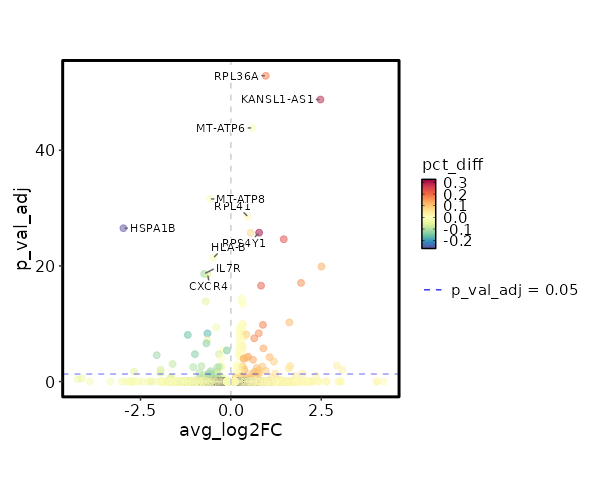
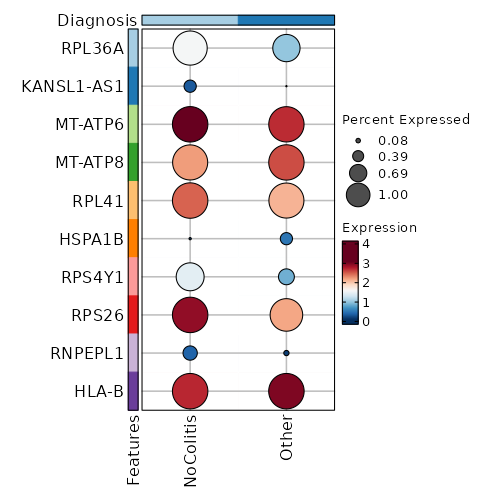
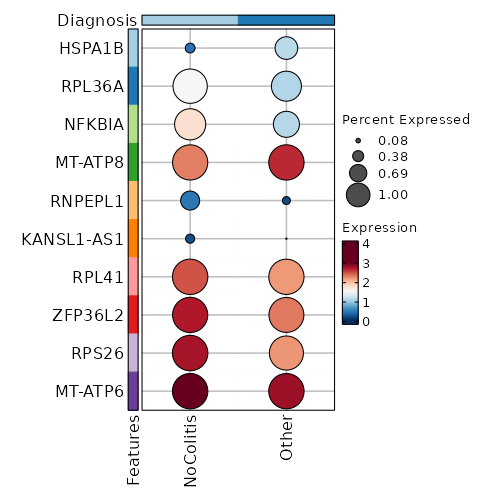
Diagnosis: NoColitis
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
| 3.855e-58 | 0.9667 | 0.963 | 0.777 | 1.293e-53 | RPL36A | NoColitis |
| 4.96e-54 | 2.4782 | 0.371 | 0.079 | 1.663e-49 | KANSL1-AS1 | NoColitis |
| 3.998e-49 | 0.5895 | 0.998 | 0.994 | 1.341e-44 | MT-ATP6 | NoColitis |
| 7.775e-37 | -0.5783 | 0.989 | 0.995 | 2.608e-32 | MT-ATP8 | NoColitis |
| 9.215e-34 | 0.4717 | 0.998 | 0.987 | 3.091e-29 | RPL41 | NoColitis |
| 8.945e-32 | -2.9729 | 0.103 | 0.369 | 3e-27 | HSPA1B | NoColitis |
| 5.324e-31 | 0.781 | 0.795 | 0.47 | 1.785e-26 | RPS4Y1 | NoColitis |
| 5.773e-31 | 0.5475 | 0.998 | 0.921 | 1.936e-26 | RPS26 | NoColitis |
| 7.193e-30 | 1.4621 | 0.426 | 0.183 | 2.412e-25 | RNPEPL1 | NoColitis |
| 1.193e-26 | -0.4833 | 0.998 | 0.998 | 4.002e-22 | HLA-B | NoColitis |
| 3.78e-25 | 2.5076 | 0.164 | 0.029 | 1.268e-20 | RAB34 | NoColitis |
| 7.022e-24 | -0.7387 | 0.779 | 0.899 | 2.355e-19 | IL7R | NoColitis |
| 9.143e-24 | -0.642 | 0.86 | 0.917 | 3.066e-19 | CXCR4 | NoColitis |
| 2.587e-22 | 1.9406 | 0.242 | 0.078 | 8.677e-18 | CHI3L2 | NoColitis |
| 7.808e-22 | 0.8346 | 0.633 | 0.416 | 2.619e-17 | GZMM | NoColitis |
| 1.183e-19 | 0.2977 | 1 | 1 | 0 | RPS28 | NoColitis |
| 3.925e-19 | -0.6925 | 0.876 | 0.928 | 0 | HSP90AA1 | NoColitis |
| 4.399e-19 | 0.3107 | 1 | 1 | 0 | RPL35A | NoColitis |
| 1.381e-18 | 0.3373 | 0.996 | 0.996 | 0 | MT-CO3 | NoColitis |
| 0 | 1.6186 | 0.185 | 0.063 | 0 | MRPL23 | NoColitis |
Downloading entire data
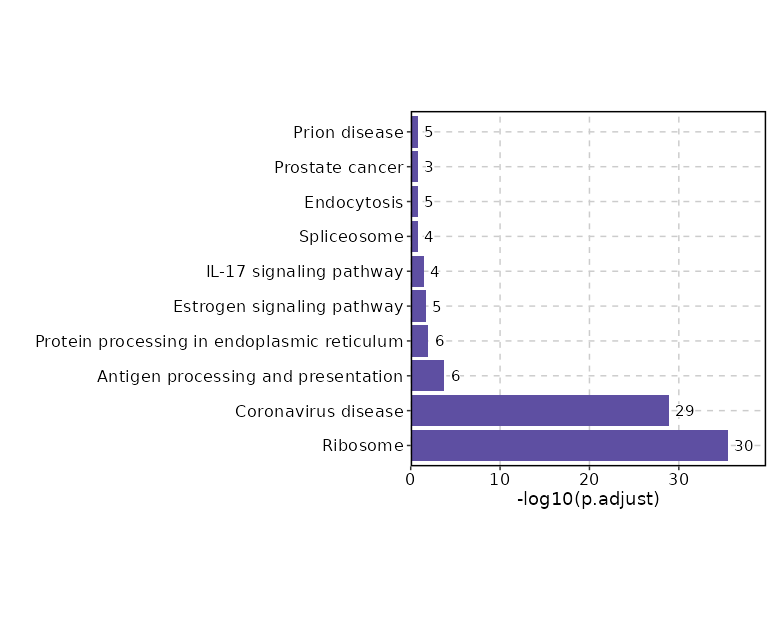
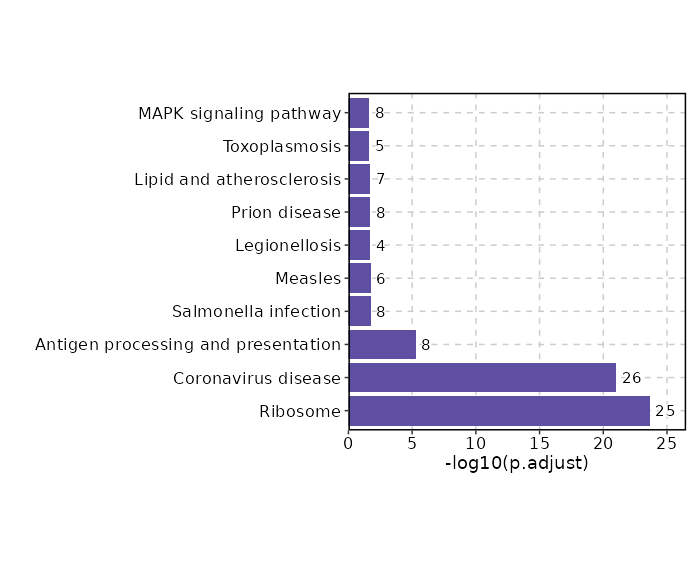
Enrichment Analysis
| Ribosome | 30/158 | 3.166e-38 | 3.45e-36 | 57.1788 | 4937.1574 | RPL22;RPS8;RPL24;RPL26;RPL28;RPS4Y1;RPL36AL;RPL41;RPS27;RPS26;RPS28;RPS21;RPS23;RPS25;RPS24;RPL31;RPL32;RPL34;RPL37;RPL39;RPS10;RPS14;RPS13;MRPL23;RPL37A;RPL36A;RPS15A;RPL35A;RPL23A;RPLP1 | 1 | KEGG_2021_Human |
| Coronavirus disease | 29/232 | 2.267e-31 | 1.235e-29 | 34.2962 | 2420 | RPL22;RPL24;RPL26;RPL28;RPS4Y1;RPL36AL;RPL41;RPS27;RPS26;RPS28;RPS21;RPS23;RPS25;RPS24;RPL31;RPL32;RPL34;RPL37;RPL39;RPS10;RPS14;RPS13;RPS15A;RPLP1;RPS8;RPL37A;RPL36A;RPL35A;RPL23A | 2 | KEGG_2021_Human |
| Antigen processing and presentation | 6/78 | 0 | 0.0002 | 15.7278 | 192.8324 | HSP90AA1;HSPA1A;HSPA1B;HLA-C;HLA-B;HSP90AB1 | 3 | KEGG_2021_Human |
| Protein processing in endoplasmic reticulum | 6/171 | 0.0004 | 0.0105 | 6.8308 | 53.6857 | HSPA1A;HSPA1B;PPP1R15A;HSP90AA1;DNAJB1;HSP90AB1 | 4 | KEGG_2021_Human |
| Estrogen signaling pathway | 5/137 | 0.001 | 0.0219 | 7.0601 | 48.7359 | HSPA1A;FKBP5;HSPA1B;HSP90AA1;HSP90AB1 | 5 | KEGG_2021_Human |
| IL-17 signaling pathway | 4/94 | 0.0019 | 0.0339 | 8.2239 | 51.6732 | HSP90AA1;JUND;FOSB;HSP90AB1 | 6 | KEGG_2021_Human |
| Spliceosome | 4/150 | 0.0098 | 0.1526 | 5.0552 | 23.3834 | HSPA1A;HSPA1B;SRSF2;SNRPD2 | 7 | KEGG_2021_Human |
| Endocytosis | 5/252 | 0.0133 | 0.1712 | 3.7511 | 16.2104 | HSPA1A;HSPA1B;HLA-C;HLA-B;CXCR4 | 8 | KEGG_2021_Human |
| Prostate cancer | 3/97 | 0.0168 | 0.1712 | 5.8496 | 23.8958 | HSP90AA1;HSP90AB1;GSTP1 | 9 | KEGG_2021_Human |
| Prion disease | 5/273 | 0.0182 | 0.1712 | 3.4534 | 13.8441 | HSPA1A;HSPA1B;ATP5F1E;NDUFA11;TUBA1B | 10 | KEGG_2021_Human |
| Allograft rejection | 2/38 | 0.0188 | 0.1712 | 10.1188 | 40.188 | HLA-C;HLA-B | 11 | KEGG_2021_Human |
| Primary immunodeficiency | 2/38 | 0.0188 | 0.1712 | 10.1188 | 40.188 | PTPRC;IL7R | 11 | KEGG_2021_Human |
| Graft-versus-host disease | 2/42 | 0.0227 | 0.1851 | 9.105 | 34.4465 | HLA-C;HLA-B | 13 | KEGG_2021_Human |
| Type I diabetes mellitus | 2/43 | 0.0238 | 0.1851 | 8.8825 | 33.2133 | HLA-C;HLA-B | 14 | KEGG_2021_Human |
| Human immunodeficiency virus 1 infection | 4/212 | 0.0305 | 0.2173 | 3.5372 | 12.3399 | HLA-C;HLA-B;GNG10;CXCR4 | 15 | KEGG_2021_Human |
| Lipid and atherosclerosis | 4/215 | 0.0319 | 0.2173 | 3.4864 | 12.0087 | HSPA1A;HSPA1B;HSP90AA1;HSP90AB1 | 16 | KEGG_2021_Human |
| Autoimmune thyroid disease | 2/53 | 0.035 | 0.2173 | 7.1373 | 23.9255 | HLA-C;HLA-B | 17 | KEGG_2021_Human |
| Human cytomegalovirus infection | 4/225 | 0.0368 | 0.2173 | 3.3269 | 10.9881 | HLA-C;HLA-B;GNG10;CXCR4 | 18 | KEGG_2021_Human |
| Oxidative phosphorylation | 3/133 | 0.0379 | 0.2173 | 4.222 | 13.8131 | ATP5F1E;ATP5MG;NDUFA11 | 19 | KEGG_2021_Human |
| Legionellosis | 2/57 | 0.04 | 0.2173 | 6.6168 | 21.3034 | HSPA1A;HSPA1B | 20 | KEGG_2021_Human |
Downloading entire data
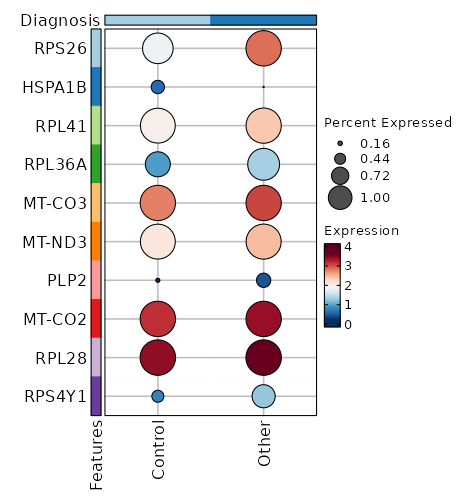
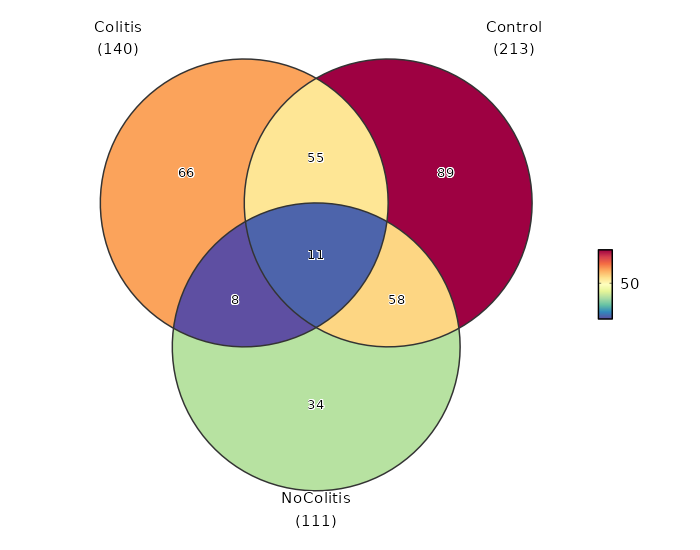
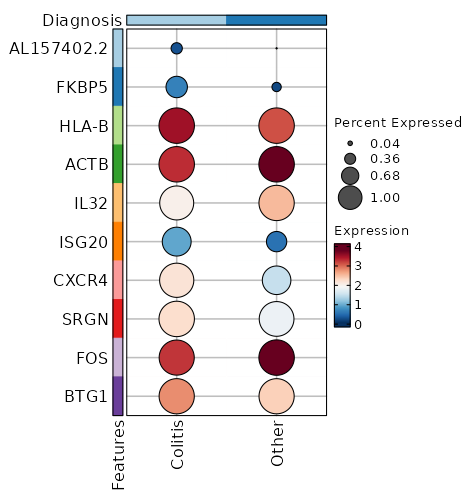
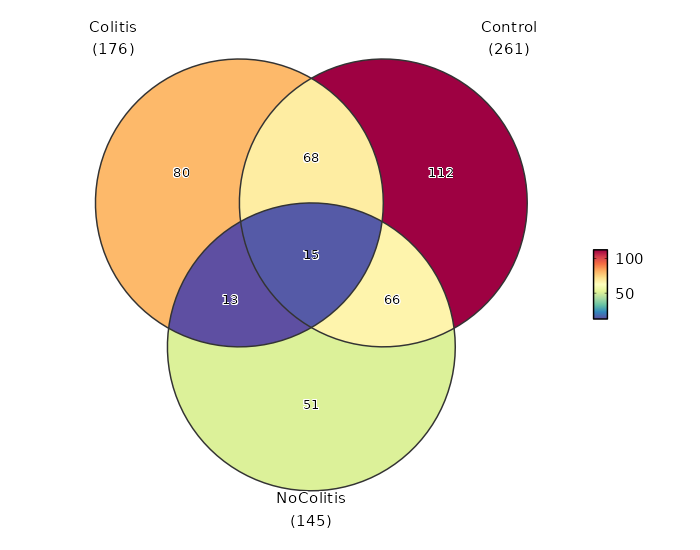
Diagnosis (Overlaps)
Image loading error!
c2
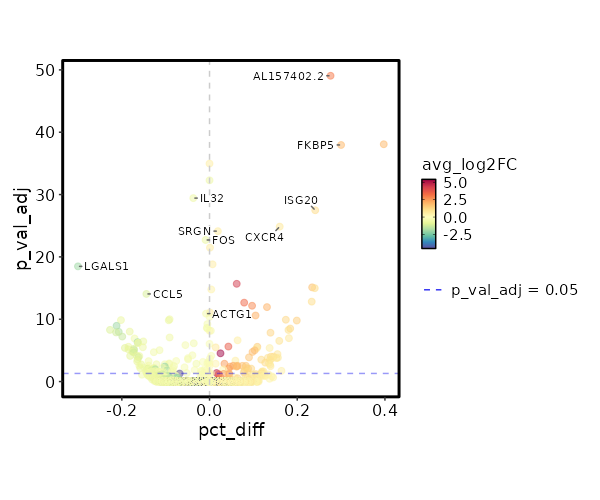
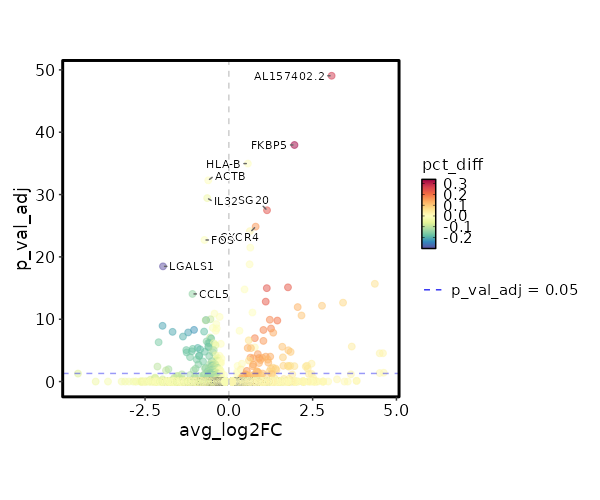
Diagnosis: Colitis
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
| 2.62e-54 | 3.0666 | 0.32 | 0.044 | 8.786e-50 | AL157402.2 | Colitis |
| 3.338e-43 | 1.9577 | 0.605 | 0.267 | 1.119e-38 | FKBP5 | Colitis |
| 3.198e-40 | 0.567 | 1 | 1 | 1.073e-35 | HLA-B | Colitis |
| 1.552e-37 | -0.6141 | 1 | 1 | 5.203e-33 | ACTB | Colitis |
| 1.104e-34 | -0.6454 | 0.959 | 0.996 | 3.703e-30 | IL32 | Colitis |
| 9.82e-33 | 1.1389 | 0.815 | 0.574 | 3.294e-28 | ISG20 | Colitis |
| 4.153e-30 | 0.7999 | 0.966 | 0.806 | 1.393e-25 | CXCR4 | Colitis |
| 2.222e-29 | 0.6284 | 1 | 0.981 | 7.451e-25 | SRGN | Colitis |
| 5.778e-28 | -0.727 | 0.991 | 1 | 1.938e-23 | FOS | Colitis |
| 9.822e-27 | 0.6403 | 0.994 | 0.993 | 3.294e-22 | BTG1 | Colitis |
| 4.622e-24 | 0.6194 | 0.994 | 0.987 | 1.55e-19 | ZFP36L2 | Colitis |
| 9.833e-24 | -1.9645 | 0.197 | 0.497 | 3.298e-19 | LGALS1 | Colitis |
| 6.206e-21 | 4.3559 | 0.063 | 0.001 | 2.082e-16 | IGKV3D-20 | Colitis |
| 2.279e-20 | 1.764 | 0.464 | 0.23 | 7.644e-16 | KLF2 | Colitis |
| 3.094e-20 | 1.1323 | 0.592 | 0.352 | 0 | SMAP2 | Colitis |
| 4.952e-20 | 0.4682 | 0.991 | 0.987 | 0 | IL7R | Colitis |
| 2.618e-19 | -1.0858 | 0.636 | 0.78 | 0 | CCL5 | Colitis |
| 4.484e-18 | 1.1006 | 0.552 | 0.319 | 0 | STAT1 | Colitis |
| 6.54e-18 | 3.4072 | 0.088 | 0.009 | 0 | NPDC1 | Colitis |
| 2.058e-17 | 2.7791 | 0.116 | 0.019 | 0 | SELL | Colitis |
Downloading entire data
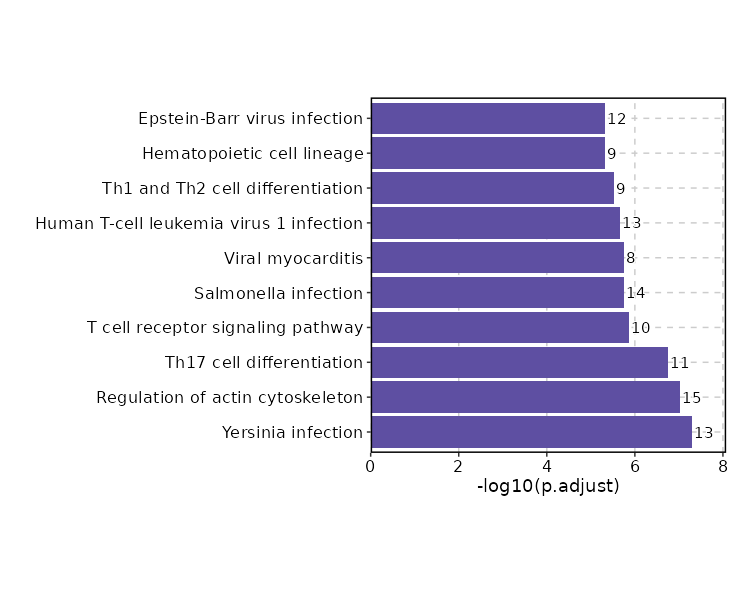
Enrichment Analysis
| Yersinia infection | 13/137 | 0 | 0 | 12.6707 | 280.0482 | CD4;TNF;ACTB;FYB1;ARPC5;PXN;ACTG1;ACTR3;NFKBIA;LCP2;LAT;FOS;LCK | 1 | KEGG_2021_Human |
| Regulation of actin cytoskeleton | 15/218 | 0 | 0 | 9.0051 | 187.0314 | PFN1;CYFIP2;MYL12B;ACTB;MYL12A;TMSB4X;CXCR4;PPP1CA;ARPC5;PXN;ACTG1;ITGA1;ACTR3;ITGAE;ITGB2 | 2 | KEGG_2021_Human |
| Th17 cell differentiation | 11/107 | 0 | 0 | 13.7 | 270.4508 | CD4;HLA-DRB1;HSP90AA1;NFKBIA;IRF4;LAT;CD3G;CD3E;STAT1;FOS;LCK | 3 | KEGG_2021_Human |
| T cell receptor signaling pathway | 10/104 | 0 | 0 | 12.6442 | 220.0956 | CD4;TNF;NFKBIA;LCP2;LAT;CD3G;CD3E;FOS;LCK;CD40LG | 4 | KEGG_2021_Human |
| Salmonella infection | 14/249 | 0 | 0 | 7.2037 | 121.7008 | PFN1;CYFIP2;MYL12B;ACTB;MYL12A;RIPK2;FOS;TNF;ARPC5;ACTG1;ACTR3;HSP90AA1;NFKBIA;GAPDH | 5 | KEGG_2021_Human |
| Viral myocarditis | 8/60 | 0 | 0 | 18.1062 | 303.2913 | ACTB;HLA-C;HLA-B;CD40LG;HLA-DRB1;ITGB2;ACTG1;CD55 | 6 | KEGG_2021_Human |
| Human T-cell leukemia virus 1 infection | 13/219 | 0 | 0 | 7.5953 | 124.5871 | CD4;EGR1;HLA-C;HLA-B;HLA-DRB1;CD3G;CD3E;FOS;TNF;CCND3;NFKBIA;LCK;ITGB2 | 7 | KEGG_2021_Human |
| Th1 and Th2 cell differentiation | 9/92 | 0 | 0 | 12.8179 | 204.2961 | NFKBIA;CD4;LAT;CD3G;CD3E;STAT1;FOS;LCK;HLA-DRB1 | 8 | KEGG_2021_Human |
| Hematopoietic cell lineage | 9/99 | 0 | 0 | 11.8168 | 180.8153 | CD2;CD4;TNF;HLA-DRB1;IL7R;ITGA1;CD3G;CD3E;CD55 | 9 | KEGG_2021_Human |
| Epstein-Barr virus infection | 12/202 | 0 | 0 | 7.5612 | 115.2443 | VIM;HLA-C;HLA-B;HLA-DRB1;CD3G;CD3E;STAT1;TNF;CCND3;TAPBP;NFKBIA;TNFAIP3 | 10 | KEGG_2021_Human |
| Allograft rejection | 6/38 | 0 | 0 | 21.8294 | 303.1248 | GZMB;HLA-C;HLA-B;CD40LG;HLA-DRB1;TNF | 11 | KEGG_2021_Human |
| PD-L1 expression and PD-1 checkpoint pathway in cancer | 8/89 | 0 | 0 | 11.6067 | 158.4246 | NFKBIA;CD4;LAT;CD3G;CD3E;STAT1;FOS;LCK | 12 | KEGG_2021_Human |
| Pathogenic Escherichia coli infection | 11/197 | 0 | 0 | 7.0387 | 94.7538 | CYFIP2;ACTB;FOS;TNF;ARPC5;ACTG1;ACTR3;NFKBIA;TMBIM6;MYO1B;GAPDH | 13 | KEGG_2021_Human |
| Shigellosis | 12/246 | 0 | 0 | 6.1257 | 80.5551 | PFN1;MYL12B;ACTB;MYL12A;RIPK2;TNF;ARPC5;PXN;ACTG1;ACTR3;NFKBIA;CCL5 | 14 | KEGG_2021_Human |
| Natural killer cell mediated cytotoxicity | 9/131 | 0 | 0 | 8.7032 | 112.5134 | TNF;HLA-C;HLA-B;LCP2;LAT;GZMB;LCK;CD48;ITGB2 | 15 | KEGG_2021_Human |
| Human immunodeficiency virus 1 infection | 11/212 | 0 | 0 | 6.5085 | 82.9684 | CD4;HLA-C;HLA-B;CD3G;CD3E;FOS;CXCR4;TNF;PXN;TAPBP;NFKBIA | 16 | KEGG_2021_Human |
| Antigen processing and presentation | 7/78 | 0 | 0.0001 | 11.5235 | 139.2237 | HSP90AA1;TAPBP;CD4;TNF;HLA-C;HLA-B;HLA-DRB1 | 17 | KEGG_2021_Human |
| Cytokine-cytokine receptor interaction | 12/295 | 0 | 0.0001 | 5.0524 | 57.0752 | CD4;IL16;CD27;IL32;CD40LG;CXCR4;CXCR3;TNF;IL7R;TNFRSF4;TNFRSF18;CCL5 | 18 | KEGG_2021_Human |
| Rap1 signaling pathway | 10/210 | 0 | 0.0002 | 5.9108 | 64.7308 | PFN1;SKAP1;ACTB;FYB1;APBB1IP;ACTG1;LCP2;LAT;EVL;ITGB2 | 19 | KEGG_2021_Human |
| Primary immunodeficiency | 5/38 | 0 | 0.0002 | 17.5359 | 189.9309 | LCK;CD4;CD40LG;IL7R;CD3E | 20 | KEGG_2021_Human |
Downloading entire data
Diagnosis: Control
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
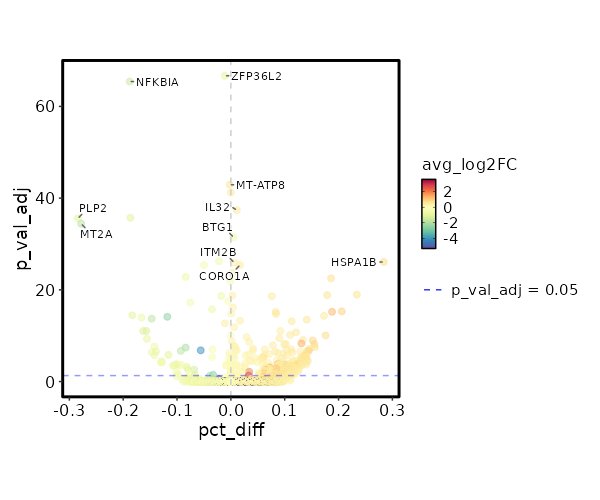
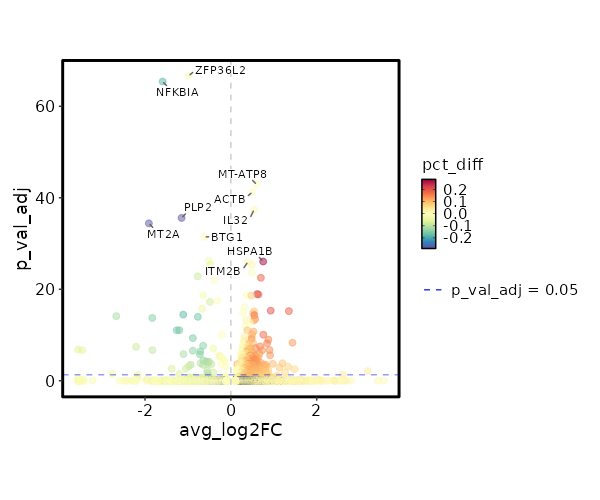
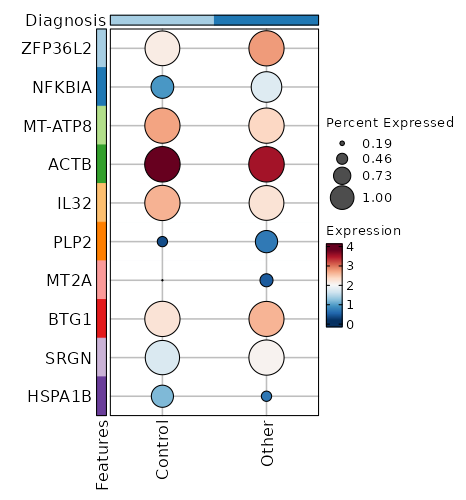
| 6.259e-72 | -0.9793 | 0.983 | 0.994 | 2.099e-67 | ZFP36L2 | Control |
| 1.133e-70 | -1.5899 | 0.695 | 0.883 | 3.798e-66 | NFKBIA | Control |
| 3.606e-48 | 0.5976 | 0.994 | 0.996 | 1.21e-43 | MT-ATP8 | Control |
| 1.854e-46 | 0.4964 | 1 | 1 | 6.217e-42 | ACTB | Control |
| 1.254e-42 | 0.5393 | 0.995 | 0.984 | 4.205e-38 | IL32 | Control |
| 7.613e-41 | -1.1484 | 0.396 | 0.683 | 2.553e-36 | PLP2 | Control |
| 1.165e-39 | -1.9102 | 0.185 | 0.463 | 3.907e-35 | MT2A | Control |
| 1.046e-36 | -0.6105 | 0.995 | 0.99 | 3.509e-32 | BTG1 | Control |
| 1.727e-31 | -0.5118 | 0.973 | 0.995 | 5.791e-27 | SRGN | Control |
| 2.55e-31 | 0.7521 | 0.681 | 0.397 | 8.553e-27 | HSPA1B | Control |
| 3.34e-31 | 0.3944 | 0.991 | 0.985 | 1.12e-26 | ITM2B | Control |
| 1.151e-30 | 0.4909 | 0.983 | 0.966 | 3.861e-26 | CORO1A | Control |
| 1.187e-30 | -0.4792 | 0.95 | 1 | 3.981e-26 | RPS26 | Control |
| 6.31e-29 | 0.4866 | 1 | 0.997 | 2.116e-24 | FOS | Control |
| 4.515e-28 | -0.7741 | 0.791 | 0.875 | 1.514e-23 | CXCR4 | Control |
| 9.308e-28 | 0.6991 | 0.776 | 0.59 | 3.122e-23 | ANXA6 | Control |
| 3.659e-27 | -0.3906 | 0.997 | 0.998 | 1.227e-22 | MT-ND3 | Control |
| 3.463e-24 | 0.6143 | 0.759 | 0.525 | 1.161e-19 | NR4A1 | Control |
| 4.341e-24 | 0.6453 | 0.786 | 0.607 | 1.456e-19 | EIF5A | Control |
| 5.403e-24 | 0.3642 | 1 | 0.997 | 1.812e-19 | PFN1 | Control |
Downloading entire data
Enrichment Analysis
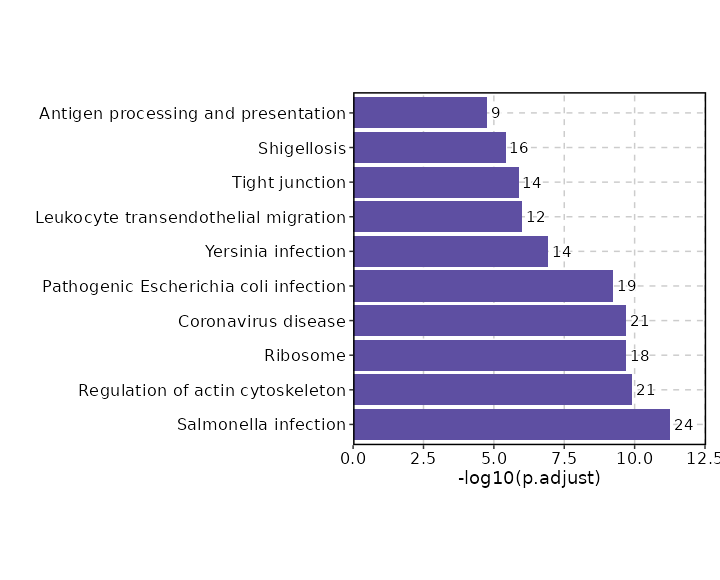
| Salmonella infection | 24/249 | 0 | 0 | 8.7827 | 274.9736 | PFN1;CYFIP2;AHNAK;MYL12B;ACTB;MYL12A;ANXA2;TUBB;PTPRC;FOS;TUBA1B;TNF;HSP90B1;M6PR;ARPC5;ARPC3;ARPC2;ACTG1;ACTR3;ACTR2;NFKBIA;CDC42;GAPDH;HSP90AB1 | 1 | KEGG_2021_Human |
| Regulation of actin cytoskeleton | 21/218 | 0 | 0 | 8.6798 | 238.7993 | PFN1;CYFIP2;MYL12B;ACTB;MYL12A;CXCR4;RAC2;PPP1CA;PPP1CC;CFL1;ARPC5;ARPC3;ARPC2;ACTG1;ACTR3;ACTR2;ITGA4;MSN;CDC42;ITGB2;ITGB7 | 2 | KEGG_2021_Human |
| Ribosome | 18/158 | 0 | 0 | 10.3698 | 275.7577 | RPL23;RPS7;RPSA;RPL28;RPL12;RPS5;RPL41;RPS27;RPS26;RPS29;RPS28;RPL5;RPL6;RPL7;RPL8;MRPL32;RPL36A;RPS4X | 3 | KEGG_2021_Human |
| Coronavirus disease | 21/232 | 0 | 0 | 8.0981 | 213.0218 | RPL23;RPL28;RPL12;RPL41;RPS27;RPS26;RPS29;RPS28;RPL5;RPL6;RPL7;RPL8;STAT1;FOS;RPS7;RPSA;TNF;RPS5;NFKBIA;RPL36A;RPS4X | 4 | KEGG_2021_Human |
| Pathogenic Escherichia coli infection | 19/197 | 0 | 0 | 8.628 | 215.8854 | CYFIP2;ACTB;TUBB;FOS;TUBA1B;PTPN6;TNF;ARPC5;ARPC3;ARPC2;NCL;ACTG1;ACTR3;ACTR2;NFKBIA;TMBIM6;CDC42;GAPDH;MYO1G | 5 | KEGG_2021_Human |
| Yersinia infection | 14/137 | 0 | 0 | 9.0393 | 176.4548 | TNF;ACTB;ARPC5;ARPC3;ARPC2;ACTG1;ACTR3;ACTR2;ITGA4;NFKBIA;LCP2;FOS;CDC42;RAC2 | 6 | KEGG_2021_Human |
| Leukocyte transendothelial migration | 12/114 | 0 | 0 | 9.2781 | 160.0592 | MYL12B;ACTB;MYL12A;ACTG1;RASSF5;ITGA4;MSN;RAP1A;CDC42;ITGB2;CXCR4;RAC2 | 7 | KEGG_2021_Human |
| Tight junction | 14/169 | 0 | 0 | 7.1614 | 120.4471 | MYL6;MYL12B;ACTB;ARPC5;MYL12A;ARPC3;ARPC2;ACTG1;ACTR3;ACTR2;MSN;RAP1A;CDC42;TUBA1B | 8 | KEGG_2021_Human |
| Shigellosis | 16/246 | 0 | 0 | 5.5394 | 86.6445 | PFN1;MYL12B;ACTB;MYL12A;TNF;ARPC5;ARPC3;ARPC2;ACTG1;UBC;UBB;ACTR3;ACTR2;NFKBIA;CDC42;CCL5 | 9 | KEGG_2021_Human |
| Antigen processing and presentation | 9/78 | 0 | 0 | 10.1812 | 142.9876 | HSPA8;TAPBP;HSPA1A;HSPA1B;TNF;PDIA3;HLA-C;KIR3DL2;HSP90AB1 | 10 | KEGG_2021_Human |
| Endocytosis | 15/252 | 0 | 0 | 5.0175 | 68.3454 | HSPA1A;HSPA1B;HLA-C;CXCR4;CAPZA1;ACAP1;ARPC5;ARPC3;ARPC2;ACTR3;ACTR2;HSPA8;CAPZB;CDC42;SMAP2 | 11 | KEGG_2021_Human |
| Measles | 11/139 | 0 | 0 | 6.7413 | 87.911 | HSPA1A;HSPA1B;HSPA8;MSN;NFKBIA;CDK6;TNFAIP3;CD3G;CD3E;STAT1;FOS | 12 | KEGG_2021_Human |
| Rap1 signaling pathway | 13/210 | 0 | 0.0001 | 5.1999 | 64.4981 | PFN1;SKAP1;CALM1;ACTB;RASSF5;APBB1IP;RAP1A;RAC2;ACTG1;LCP2;CDC42;EVL;ITGB2 | 13 | KEGG_2021_Human |
| Fc gamma R-mediated phagocytosis | 9/97 | 0 | 0.0001 | 7.9752 | 97.3055 | CFL1;ARPC5;ARPC3;ARPC2;ACTR3;ACTR2;PTPRC;CDC42;RAC2 | 14 | KEGG_2021_Human |
| Bacterial invasion of epithelial cells | 8/77 | 0 | 0.0001 | 9.0141 | 106.6074 | ACTR3;ACTR2;ACTB;ARPC5;CDC42;ARPC3;ARPC2;ACTG1 | 15 | KEGG_2021_Human |
| T cell receptor signaling pathway | 9/104 | 0 | 0.0001 | 7.385 | 85.8621 | PTPN6;TNF;NFKBIA;PTPRC;LCP2;CD3G;CD3E;FOS;CDC42 | 16 | KEGG_2021_Human |
| Human immunodeficiency virus 1 infection | 12/212 | 0 | 0.0003 | 4.7082 | 50.1528 | CALM1;HLA-C;CD3G;CD3E;FOS;CXCR4;RAC2;TNF;CFL1;PDIA3;TAPBP;NFKBIA | 17 | KEGG_2021_Human |
| Glycolysis / Gluconeogenesis | 7/67 | 0 | 0.0003 | 9.0389 | 95.2444 | TPI1;ALDOA;LDHA;PGAM1;PKM;GAPDH;ENO1 | 18 | KEGG_2021_Human |
| Lipid and atherosclerosis | 12/215 | 0 | 0.0003 | 4.6379 | 48.7609 | HSPA1A;CALM1;HSPA1B;RAP1A;FOS;TNF;HSP90B1;HSPA8;NFKBIA;CDC42;HSP90AB1;CCL5 | 19 | KEGG_2021_Human |
| Platelet activation | 9/124 | 0 | 0.0004 | 6.0944 | 62.2301 | PPP1CA;MYL12B;PPP1CC;ACTB;MYL12A;ACTG1;APBB1IP;LCP2;RAP1A | 20 | KEGG_2021_Human |
Downloading entire data
Diagnosis: NoColitis
Markers
Showing top 100 markers ordered by p_val_adj ascendingly, then abs(avg_log2FC) descendingly. Use 'Download the entire data' button to download all significant markers by 'p_val_adj < 0.05'.
| 1.587e-47 | -1.6867 | 0.305 | 0.652 | 5.321e-43 | HSPA1B | NoColitis |
| 1.497e-45 | 0.7144 | 0.969 | 0.852 | 5.019e-41 | RPL36A | NoColitis |
| 6.637e-42 | 1.0124 | 0.884 | 0.744 | 2.226e-37 | NFKBIA | NoColitis |
| 5.408e-41 | -0.6015 | 0.994 | 0.996 | 1.814e-36 | MT-ATP8 | NoColitis |
| 1.673e-38 | 1.2607 | 0.554 | 0.255 | 5.61e-34 | RNPEPL1 | NoColitis |
| 1.582e-33 | 2.1346 | 0.286 | 0.076 | 5.305e-29 | KANSL1-AS1 | NoColitis |
| 1.256e-32 | 0.3901 | 0.998 | 0.994 | 4.212e-28 | RPL41 | NoColitis |
| 4.665e-27 | 0.5516 | 0.994 | 0.986 | 1.564e-22 | ZFP36L2 | NoColitis |
| 5.116e-26 | 0.4228 | 1 | 0.963 | 1.716e-21 | RPS26 | NoColitis |
| 1.007e-25 | 0.356 | 0.998 | 0.997 | 3.376e-21 | MT-ATP6 | NoColitis |
| 1.094e-25 | -0.7308 | 0.952 | 0.979 | 3.668e-21 | HSP90AA1 | NoColitis |
| 3.563e-25 | 2.7997 | 0.152 | 0.023 | 1.195e-20 | RAB34 | NoColitis |
| 8.127e-25 | 0.7923 | 0.698 | 0.465 | 2.726e-20 | PLP2 | NoColitis |
| 3.366e-24 | 0.4024 | 0.997 | 0.997 | 1.129e-19 | MT-ND3 | NoColitis |
| 2.245e-23 | -0.8281 | 0.863 | 0.925 | 7.528e-19 | DNAJB1 | NoColitis |
| 2.553e-23 | -0.3494 | 0.998 | 1 | 8.563e-19 | RPS5 | NoColitis |
| 3.277e-20 | -0.7475 | 0.853 | 0.904 | 0 | LTB | NoColitis |
| 3.874e-20 | 1.0463 | 0.467 | 0.257 | 0 | MT2A | NoColitis |
| 3.191e-19 | 1.9107 | 0.166 | 0.043 | 0 | C20orf204 | NoColitis |
| 4.045e-19 | -0.4333 | 0.979 | 0.996 | 0 | HSPA8 | NoColitis |
Downloading entire data
Enrichment Analysis
| Ribosome | 25/158 | 1.35e-26 | 2.215e-24 | 30.8929 | 1840.189 | RPL23;RPS7;RPSA;RPL11;RPL13;RPL12;RPS5;RPL41;RPS27;RPS26;RPS29;RPS28;RPS21;RPL5;RPL6;RPL4;RPL34;RPL7;RPL37;RPL39;RPL13A;RPL37A;RPL36A;RPS4X;RPL18A | 1 | KEGG_2021_Human |
| Coronavirus disease | 26/232 | 1.224e-23 | 1.004e-21 | 20.8401 | 1099.4593 | RPL23;RPL11;RPL13;RPL12;RPL41;RPS27;RPS26;RPS29;RPS28;RPS21;RPL5;RPL6;RPL4;RPL34;RPL7;RPL37;RPL39;RPL18A;RPS7;RPSA;RPS5;RPL13A;NFKBIA;RPL37A;RPL36A;RPS4X | 2 | KEGG_2021_Human |
| Antigen processing and presentation | 8/78 | 0 | 0 | 16.5047 | 266.6885 | HSP90AA1;HSPA8;HSPA1A;HSPA1B;KIR3DL2;HLA-B;PSME2;HSP90AB1 | 3 | KEGG_2021_Human |
| Salmonella infection | 8/249 | 0.0005 | 0.0175 | 4.7525 | 36.3718 | S100A10;PTPRC;TUBA1B;ARPC3;HSP90AA1;NFKBIA;CDC42;HSP90AB1 | 4 | KEGG_2021_Human |
| Measles | 6/139 | 0.0005 | 0.0175 | 6.4008 | 48.2403 | HSPA1A;HSPA1B;HSPA8;MSN;NFKBIA;RACK1 | 5 | KEGG_2021_Human |
| Legionellosis | 4/57 | 0.0008 | 0.0203 | 10.5992 | 75.8907 | HSPA8;NFKBIA;HSPA1A;HSPA1B | 6 | KEGG_2021_Human |
| Prion disease | 8/273 | 0.0009 | 0.0203 | 4.3168 | 30.4419 | HSPA1A;HSPA1B;ATP5F1E;NDUFA11;ATP5F1B;TUBA1B;HSPA8;SLC25A6 | 7 | KEGG_2021_Human |
| Lipid and atherosclerosis | 7/215 | 0.001 | 0.0204 | 4.7913 | 33.1304 | HSPA1A;HSPA1B;HSP90AA1;HSPA8;NFKBIA;CDC42;HSP90AB1 | 8 | KEGG_2021_Human |
| Toxoplasmosis | 5/112 | 0.0013 | 0.0228 | 6.5915 | 43.5707 | HSPA1A;HSPA1B;HSPA8;NFKBIA;SOCS1 | 9 | KEGG_2021_Human |
| MAPK signaling pathway | 8/294 | 0.0014 | 0.0228 | 3.9955 | 26.284 | HSPA1A;HSPA1B;JUND;CDC25B;RASGRP2;NR4A1;HSPA8;CDC42 | 10 | KEGG_2021_Human |
| Protein processing in endoplasmic reticulum | 6/171 | 0.0016 | 0.0233 | 5.1511 | 33.2828 | HSPA1A;HSPA1B;HSP90AA1;HSPA8;DNAJB1;HSP90AB1 | 11 | KEGG_2021_Human |
| Endocytosis | 7/252 | 0.0025 | 0.0336 | 4.06 | 24.3981 | HSPA1A;HSPA1B;HLA-B;CAPZA1;ARPC3;HSPA8;CDC42 | 12 | KEGG_2021_Human |
| Estrogen signaling pathway | 5/137 | 0.0032 | 0.041 | 5.3363 | 30.5767 | HSPA1A;HSPA1B;HSP90AA1;HSPA8;HSP90AB1 | 13 | KEGG_2021_Human |
| Viral carcinogenesis | 6/203 | 0.0037 | 0.0431 | 4.3073 | 24.1463 | HNRNPK;SP100;HLA-B;NFKBIA;YWHAZ;CDC42 | 14 | KEGG_2021_Human |
| Pyruvate metabolism | 3/47 | 0.0048 | 0.05 | 9.5123 | 50.801 | MDH2;LDHB;LDHA | 15 | KEGG_2021_Human |
| IL-17 signaling pathway | 4/94 | 0.0049 | 0.05 | 6.2301 | 33.1593 | HSP90AA1;NFKBIA;JUND;HSP90AB1 | 16 | KEGG_2021_Human |
| Cysteine and methionine metabolism | 3/50 | 0.0057 | 0.055 | 8.9038 | 46.0036 | LDHB;LDHA;MDH2 | 17 | KEGG_2021_Human |
| Progesterone-mediated oocyte maturation | 4/100 | 0.0061 | 0.0553 | 5.8389 | 29.8036 | CDC25B;HSP90AA1;PDE3B;HSP90AB1 | 18 | KEGG_2021_Human |
| HIF-1 signaling pathway | 4/109 | 0.0082 | 0.0707 | 5.336 | 25.6361 | ENO1;LDHB;LDHA;TIMP1 | 19 | KEGG_2021_Human |
| Parkinson disease | 6/249 | 0.0097 | 0.0796 | 3.4838 | 16.1485 | ATP5F1E;NDUFA11;ATP5F1B;TUBA1B;UBB;SLC25A6 | 20 | KEGG_2021_Human |
Downloading entire data
Diagnosis (Overlaps)
Image loading error!