seurat_clusters
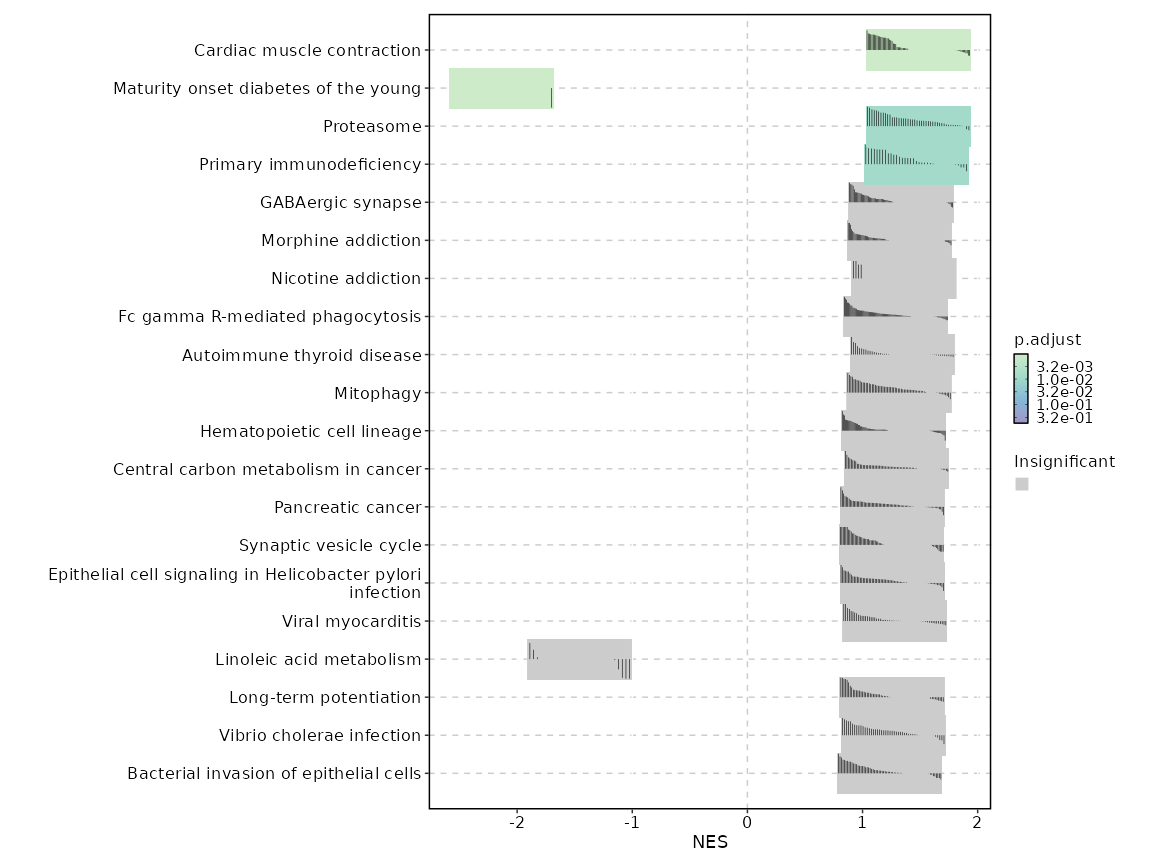
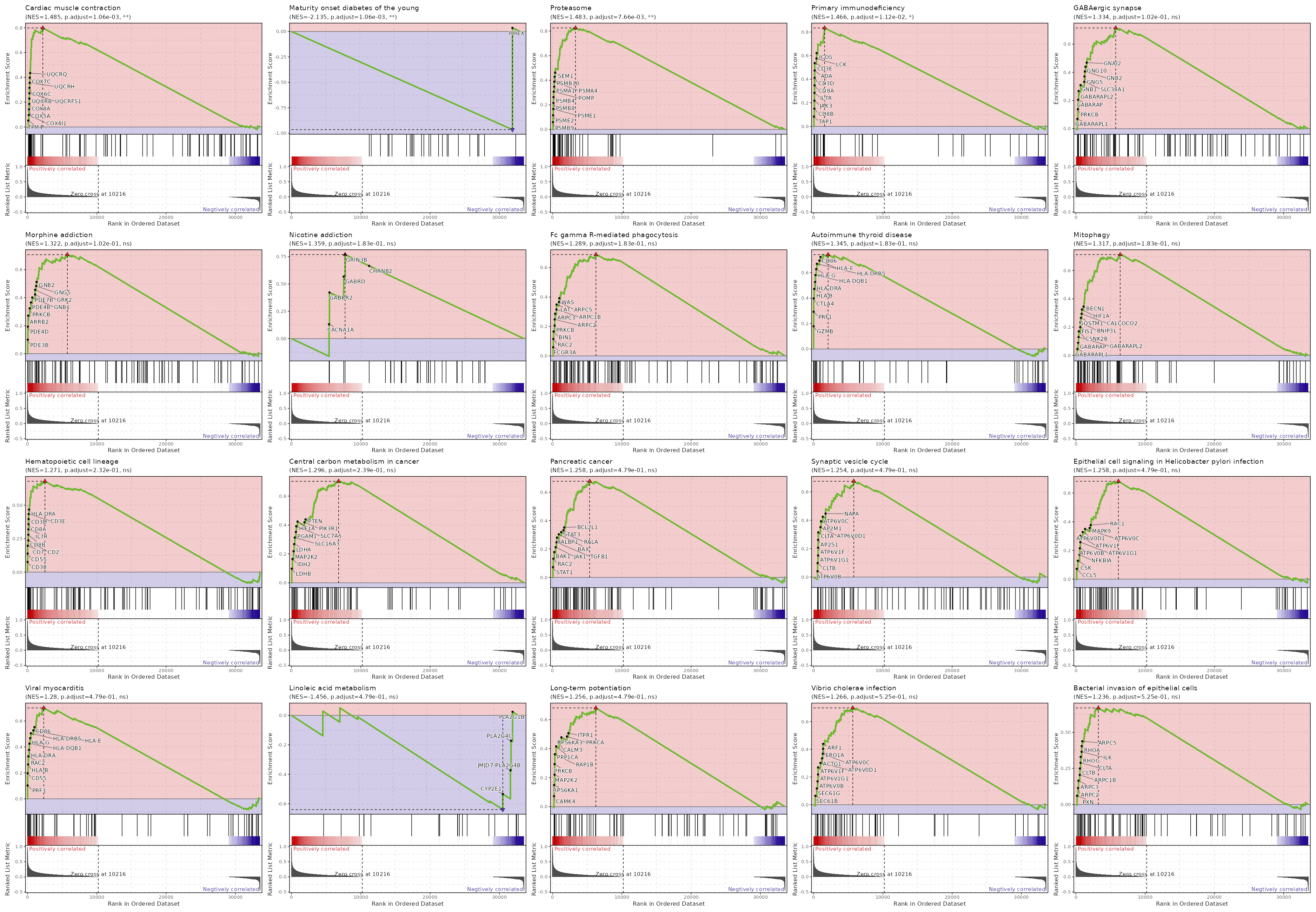
c6
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Cardiac muscle contraction | 0 | 0.0011 | 0.6105 | 0.7984 | 1.4846 | 83 | TPM4,COX4I1,COX5A,COX8A,UQCRB,UQCRFS1,COX6C,UQCRH,COX7C,UQCRQ,COX7A2,COX5B,UQCR11,COX7A2L,UQCR10,COX7B,COX6B1,COX6A1,UQCRC2,ATP1B3,CYC1,UQCRC1,UQCRHL,CACNA2D4 |
| Maturity onset diabetes of the young | 0 | 0.0011 | 0.5933 | -0.9648 | -2.1352 | 26 | HHEX |
| Proteasome | 0.0001 | 0.0077 | 0.5384 | 0.8256 | 1.4832 | 46 | PSMB9,PSME2,PSME1,PSMB8,PSMB4,POMP,PSMA1,PSMA4,PSMB10,SEM1,PSMA5,PSME4,PSMD8,PSMB3,PSMA2,PSMB2,PSMD7,PSMC1,PSMB7,PSMA6,PSMA7,PSMB6,PSMF1,ADRM1,PSMA3,PSMD6,PSMD9,PSMD4,PSMD11,PSMC2,PSMC6,PSMB1 |
| Primary immunodeficiency | 0.0002 | 0.0112 | 0.5188 | 0.8351 | 1.4665 | 37 | TAP1,CD8B,JAK3,IL7R,CD8A,CD3D,ADA,CD3E,LCK,ICOS,IL2RG,ORAI1,ZAP70,CD79A,RFX5,RFXANK,CIITA,TNFRSF13C |
| GABAergic synapse | 0.0027 | 0.1018 | 0.4317 | 0.7151 | 1.3342 | 89 | GABARAPL1,PRKCB,GABARAP,GABARAPL2,GNB1,SLC38A1,GNG5,GNB2,GNG10,GNAI2,ADCY3,PRKCA,ADCY7,ABAT,GNGT2,GNG2,GLUL,GLS,GNB4,NSF,TRAK2,ADCY4,GNG4,CACNA1A,GABRR2,CACNA1C,ADCY9,CACNA1F,GNG7 |
| Morphine addiction | 0.0032 | 0.1018 | 0.4317 | 0.7075 | 1.3217 | 91 | PDE3B,PDE4D,ARRB2,PRKCB,PDE4B,GNB1,PDE7B,GRK2,GNG5,GNB2,GNG10,GNAI2,ADCY3,PDE7A,PRKCA,ADCY7,GNGT2,GNG2,PDE4A,GNAS,GNB4,ADCY4,GRK3,GNG4,CACNA1A,GABRR2,ADCY9,GNG7 |
| Nicotine addiction | 0.0073 | 0.183 | 0.407 | 0.7692 | 1.3594 | 40 | CACNA1A,GABRR2,GABRD,GRIN3B,CHRNB2 |
| Fc gamma R-mediated phagocytosis | 0.0084 | 0.183 | 0.3807 | 0.6886 | 1.2888 | 96 | FCGR3A,RAC2,BIN1,PRKCB,ARPC2,ARPC3,ARPC1B,LAT,ARPC5,WAS,FCGR2B,INPP5D,CFL1,DNM2,PIK3R1,FCGR3B,RPS6KB2,PRKCA,RAC1,LIMK1,INPPL1,ARPC4,ACTR2,MAP2K1,VAV1,ARF6,PIK3R3,CFL2,SPHK2,PLPP1,PLCG1,CDC42,GAB2,PIP5K1C,PIP5K1A,ARPC1A,AKT1,PIK3CD,PIP5K1B,PLPP2,PLCG2,CRK,MAPK1,ARPC5L |
| Autoimmune thyroid disease | 0.009 | 0.183 | 0.3807 | 0.7418 | 1.3447 | 50 | GZMB,PRF1,CTLA4,HLA-B,HLA-DRA,HLA-DQB1,HLA-G,HLA-DRB5,HLA-E,CD86,HLA-A,CD28 |
| Mitophagy | 0.0096 | 0.183 | 0.3807 | 0.713 | 1.3169 | 68 | GABARAPL1,GABARAP,GABARAPL2,CSNK2B,FIS1,BNIP3L,SQSTM1,CALCOCO2,HIF1A,BECN1,RRAS2,ATG5,TFEB,UBA52,USP15,TBC1D17,BCL2L1,ATG9B,MAPK9,UBB,ATF4,OPTN,ATG9A,AMBRA1,MFN1,TP53,MAPK8,TBC1D15,TOMM7,KRAS,RELA,PGAM5,RAB7A,EIF2AK3,E2F1,CSNK2A2,TFE3,SP1,CSNK2A1,ULK1,PINK1,NRAS,USP8,TBK1,RRAS |
| Hematopoietic cell lineage | 0.0134 | 0.2322 | 0.3807 | 0.6788 | 1.2706 | 96 | CD38,CD55,CD7,CD2,CD8B,IL7R,CD8A,CD3D,CD3E,HLA-DRA,HLA-DQB1,ITGA1,CD44,HLA-DRB5,ITGA4,CD59,CSF2,TFRC,IL11RA,IL7,FLT3LG,CD3G,ITGA2 |
| Central carbon metabolism in cancer | 0.015 | 0.2386 | 0.3807 | 0.7027 | 1.2961 | 70 | LDHB,IDH2,MAP2K2,LDHA,SLC16A3,PGAM1,SLC7A5,HIF1A,PIK3R1,PTEN,G6PD,HK1,MAP2K1,TP53,MYC,NTRK1,SLC1A5,PIK3R3,KRAS,HK2,ERBB2,GLS,HKDC1,IDH1,SIRT6,TIGAR,PDK1,PFKL,FGFR1,AKT1,PIK3CD,PDGFRB,PFKM,NRAS,MAPK1,SIRT3,PDHA1,PFKP,FGFR2 |
| Pancreatic cancer | 0.0367 | 0.4789 | 0.2378 | 0.678 | 1.2576 | 76 | STAT1,RAC2,BAK1,JAK1,TGFB1,RALBP1,BAX,RALA,STAT3,BCL2L1,E2F2,PIK3R1,MAPK9,RPS6KB2,CDKN2A,RAC1,SMAD4,RB1,MAP2K1,GADD45A,TP53,ARHGEF6,MAPK8,PIK3R3,KRAS,BAD,RELA,ERBB2,IKBKB,TGFB3,ARAF,SMAD2,CDC42,E2F1,DDB2,RALGDS,NFKB1,AKT1,SMAD3,PIK3CD |
| Synaptic vesicle cycle | 0.0376 | 0.4789 | 0.2344 | 0.6746 | 1.2544 | 78 | ATP6V0B,CLTB,ATP6V1G1,ATP6V1F,AP2S1,CLTA,ATP6V0D1,AP2M1,ATP6V0C,NAPA,DNM2,ATP6V1D,ATP6V1A,AP2A1,TCIRG1,ATP6V0A1,ATP6V1B2,ATP6V1G2,AP2A2,NSF,ATP6V0E1,ATP6V1H,ATP6V0A2,CACNA1A,ATP6V1C2,SLC1A7,STXBP1 |
| Epithelial cell signaling in Helicobacter pylori infection | 0.0391 | 0.4789 | 0.2311 | 0.6821 | 1.258 | 70 | CCL5,CSK,NFKBIA,ATP6V0B,ATP6V1G1,ATP6V1F,ATP6V0D1,ATP6V0C,MAPK9,RAC1,ADAM10,ATP6V1D,ATP6V1A,JAM3,TCIRG1,MAPK8,ATP6V0A1,MAPK11,ATP6V1B2,RELA,CXCR1,ATP6AP1,IKBKB,PLCG1,ATP6V1G2,MAPK12,CXCL3,CDC42,ATP6V0E1,ATP6V1H,HBEGF,NFKB1,ATP6V0A2,ATP6V1C2,IKBKG,PLCG2 |
| Viral myocarditis | 0.0436 | 0.4789 | 0.2193 | 0.7008 | 1.2804 | 57 | PRF1,CD55,HLA-B,RAC2,HLA-DRA,HLA-DQB1,HLA-G,HLA-DRB5,HLA-E,CD86,ACTG1,FYN,EIF4G3,HLA-A,BID,CD28,RAC1,CASP8 |
| Linoleic acid metabolism | 0.0445 | 0.4789 | 0.3218 | -0.6403 | -1.4561 | 28 | PLA2G1B,PLA2G4C,JMJD7-PLA2G4B,CYP2E1 |
| Long-term potentiation | 0.0451 | 0.4789 | 0.2139 | 0.6811 | 1.2562 | 67 | CAMK4,RPS6KA1,MAP2K2,PRKCB,RAP1B,PPP1CA,CALM3,RPS6KA3,PRKCA,ITPR1,ATF4,EP300,CREBBP,MAP2K1,PPP1CC,KRAS,PLCB2,CALM1,ARAF,PLCB3,CAMK2D,CACNA1C,PPP3CA,PPP3CB,NRAS,MAPK1 |
| Vibrio cholerae infection | 0.0554 | 0.525 | 0.1938 | 0.6982 | 1.2658 | 50 | SEC61B,SEC61G,ATP6V0B,ATP6V1G1,ATP6V1F,ATP6V0D1,ACTG1,ATP6V0C,ERO1A,ARF1,ADCY3,PRKCA,ATP6V1D,ATP6V1A,TCIRG1,KDELR1,ATP6V0A1,SEC61A2,ATP6V1B2,ATP6AP1,PLCG1,GNAS,ATP6V1G2,ATP6V0E1,ATP6V1H,KDELR3,ATP6V0A2,ADCY9,ATP6V1C2 |
| Bacterial invasion of epithelial cells | 0.0616 | 0.525 | 0.1814 | 0.6693 | 1.2357 | 69 | PXN,ARPC2,ARPC3,ARPC1B,CLTB,CLTA,RHOG,ILK,RHOA,ARPC5,ITGB1,ACTG1,HCLS1,DNM2,PIK3R1,MAD2L2,CBL,RAC1,ELMO1,ARPC4,ACTR2,PIK3R3,VCL,CDC42,ARPC1A,PIK3CD,ARHGEF26 |
Downloading entire data
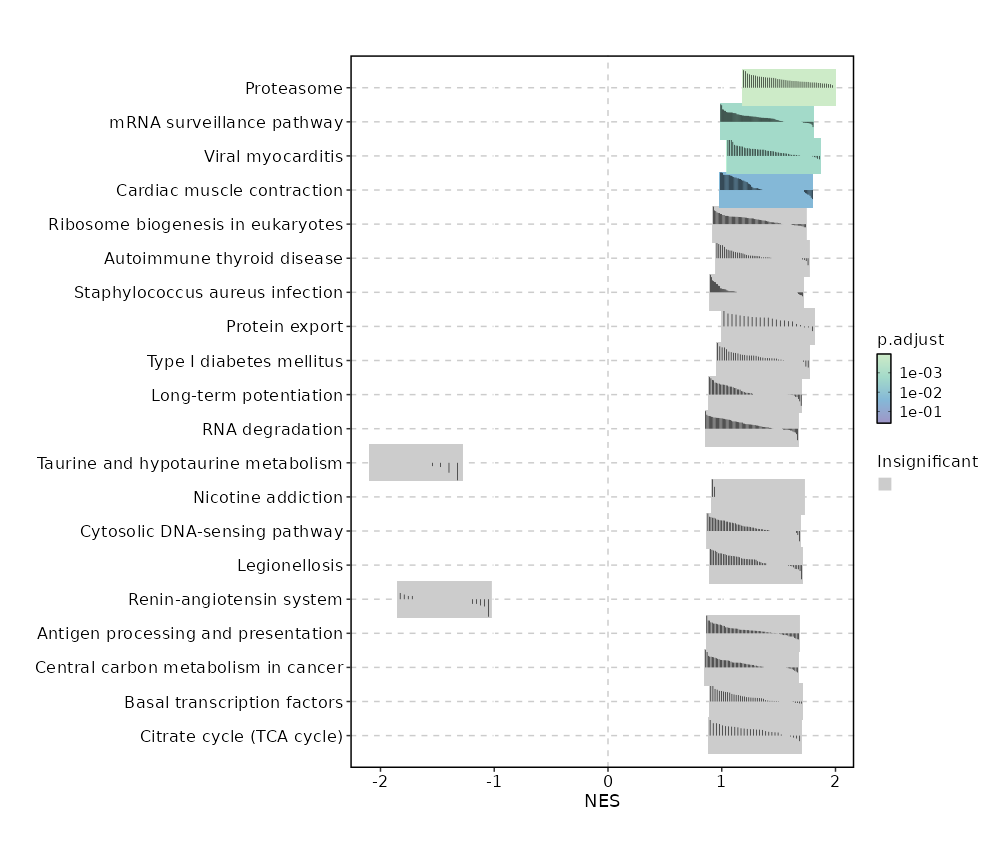
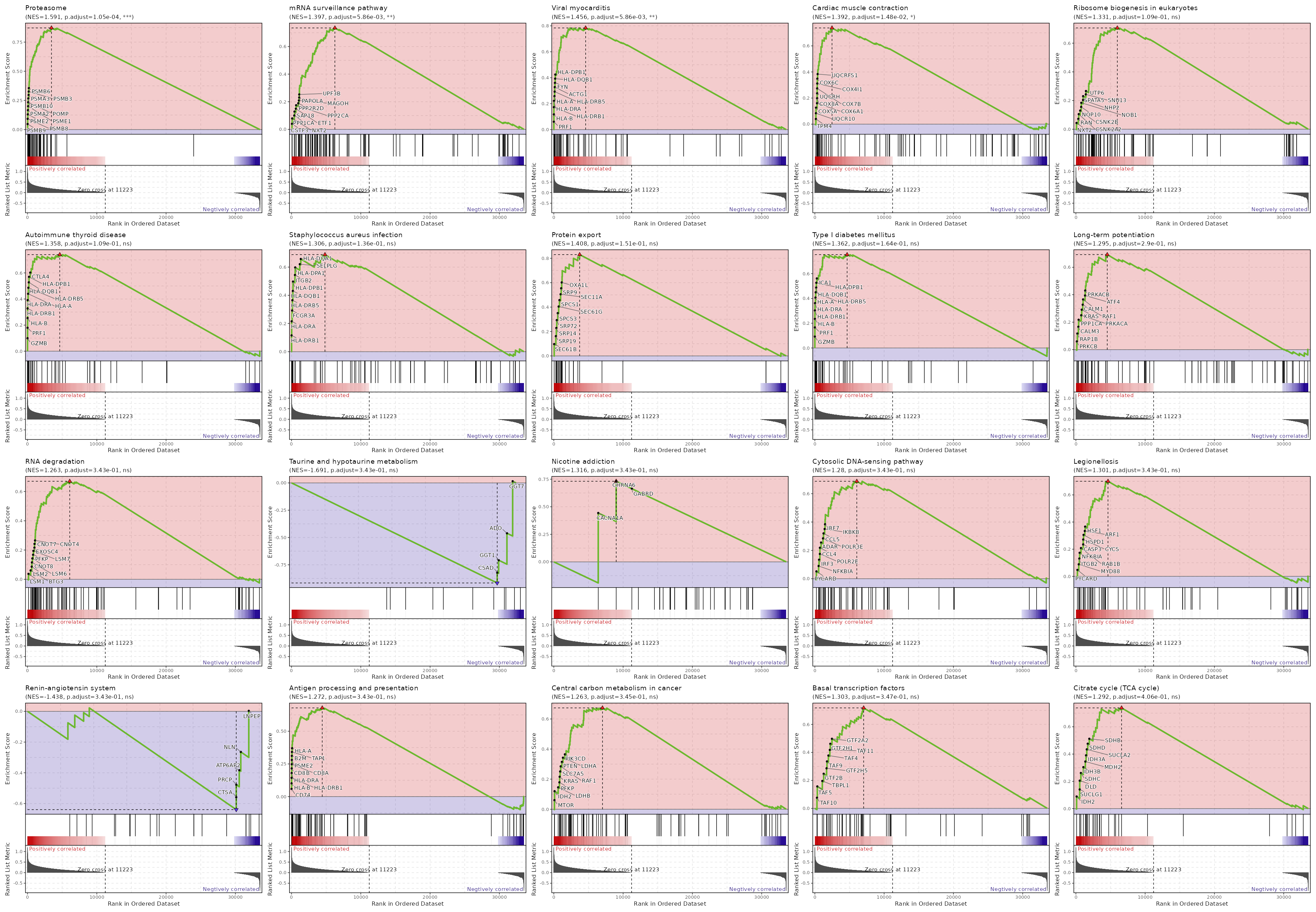
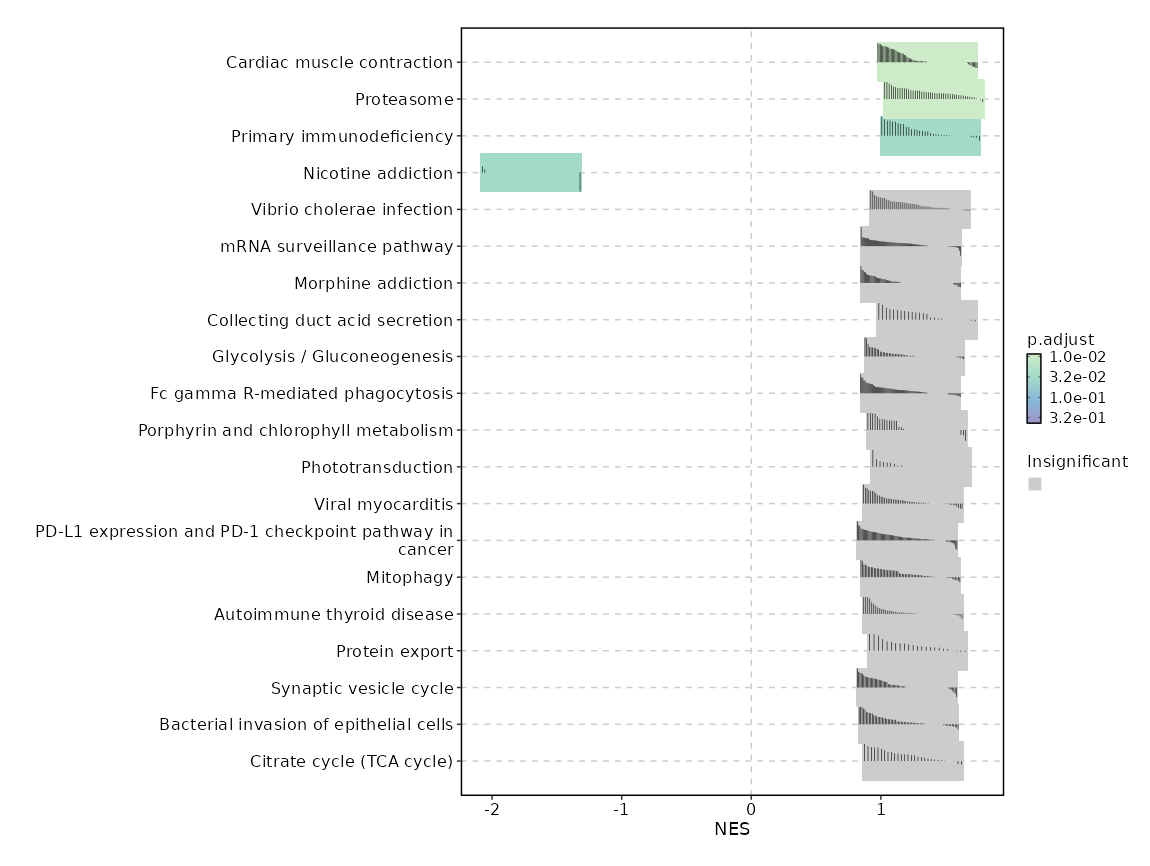
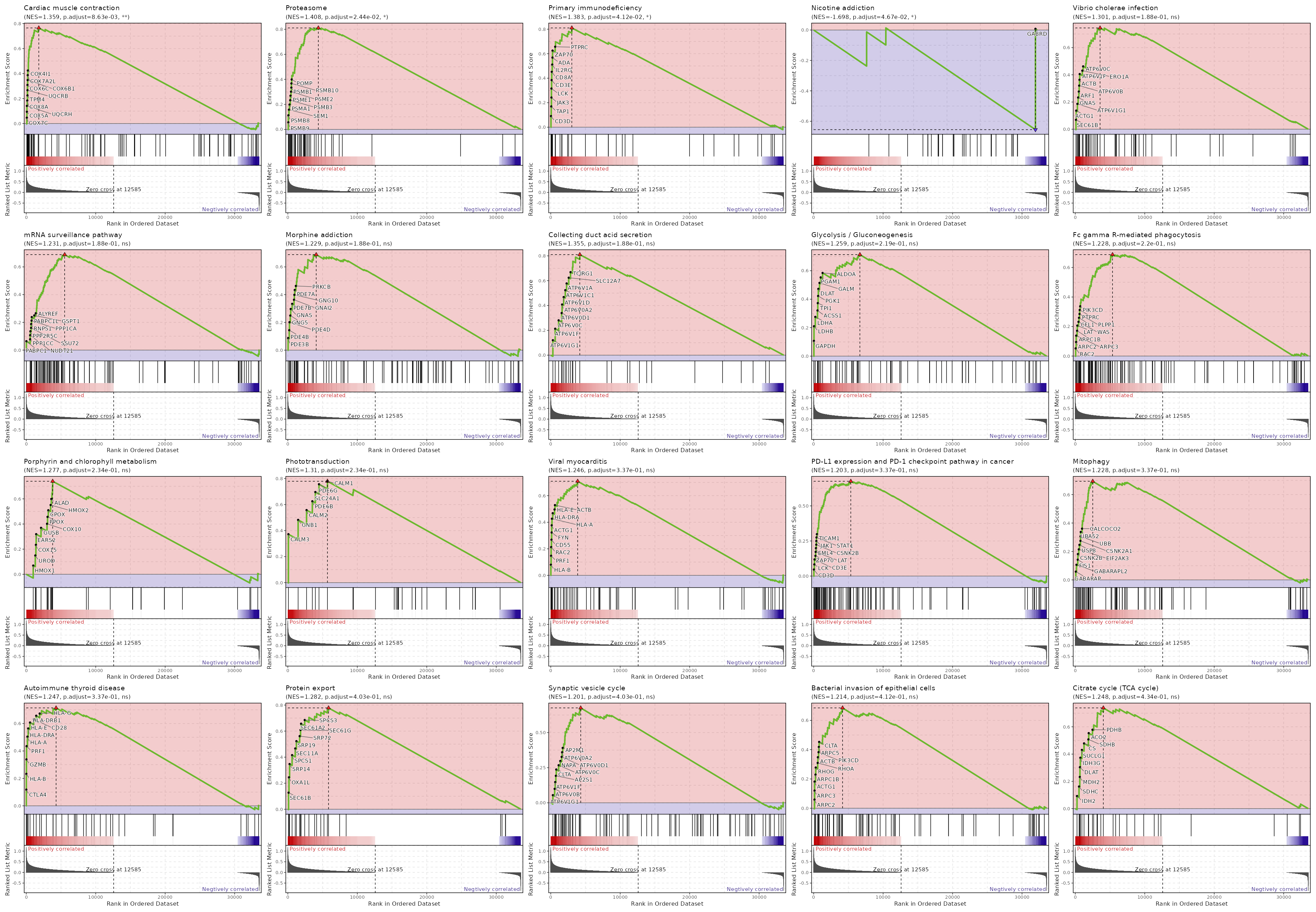
c10
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
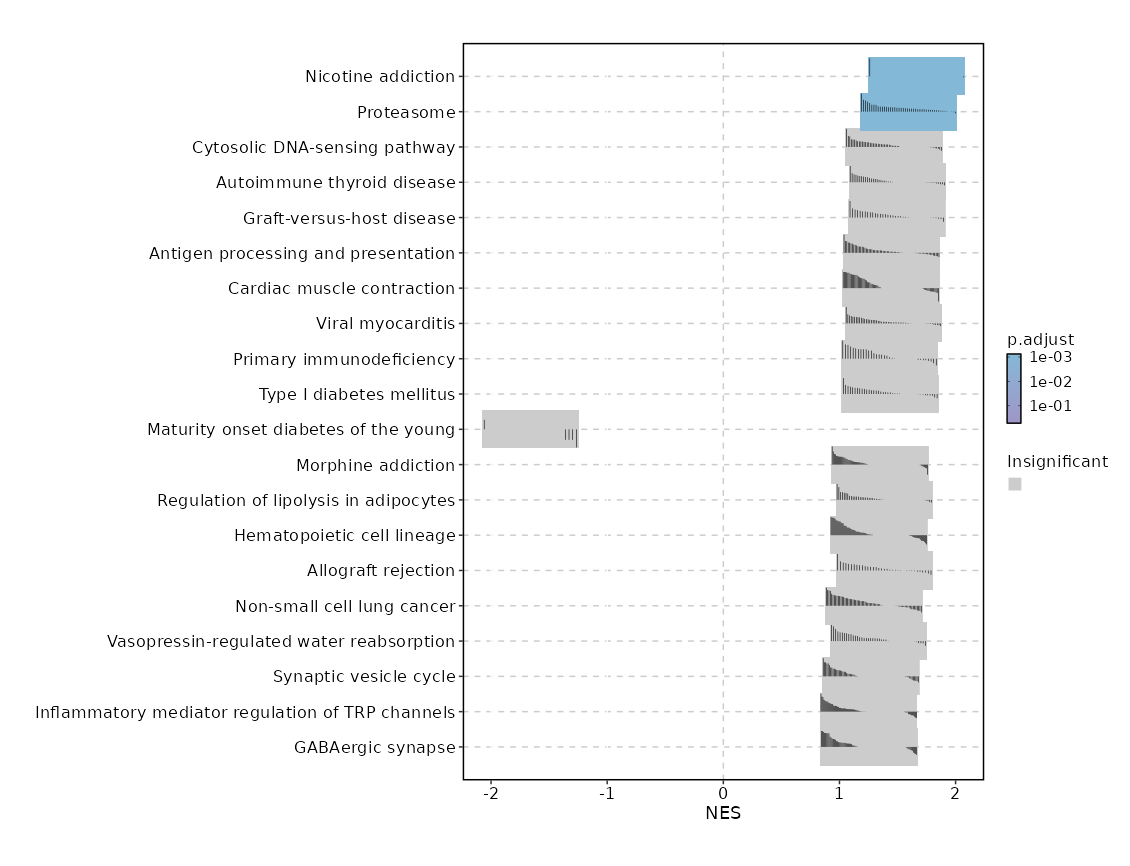
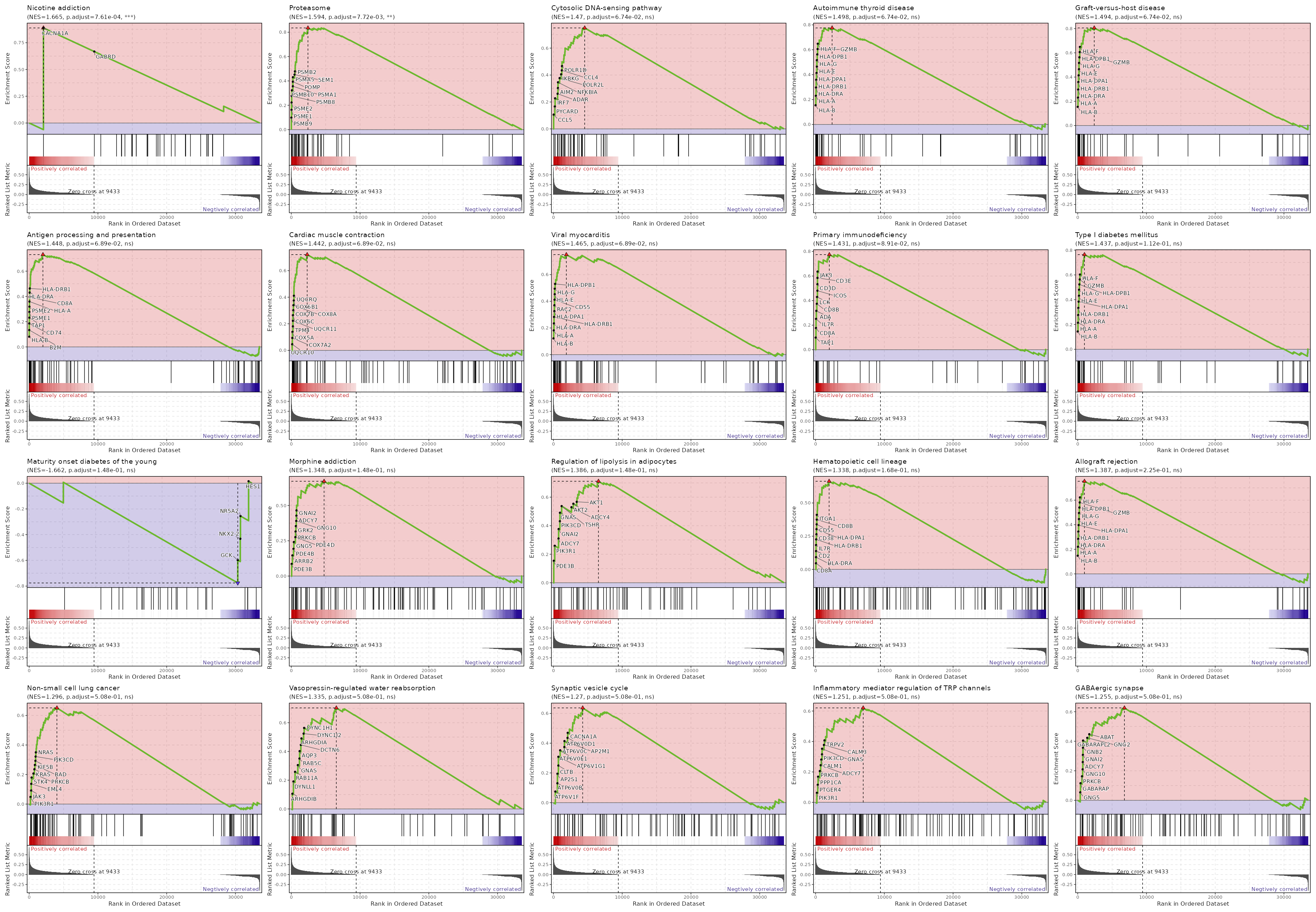
| Proteasome | 0 | 0.0001 | 0.6594 | 0.8716 | 1.5907 | 46 | PSMB9,PSMB8,PSME2,PSMA2,PSME1,POMP,PSMB10,PSMA3,PSMB3,PSMB6,PSMB2,PSMA5,PSMA7,PSMA6,PSMA1,SEM1,PSMB1,PSMD9,PSMA4,PSMD7,PSMC5,ADRM1,PSMD8,IFNG,PSMC1,PSMC6,PSMD13,PSMB7,PSME4,PSMA8,PSMD3,PSMC4,PSMB4,PSMF1,PSMB5,PSMC2,PSMD4,PSMD1,PSMC3,PSMD6,PSME3,PSMD11 |
| mRNA surveillance pathway | 0.0001 | 0.0059 | 0.5384 | 0.7347 | 1.3971 | 98 | CSTF3,NXT2,PPP1CA,SAP18,ETF1,PPP2CA,PPP2R2D,MAGOH,PAPOLA,UPF3B,PELO,RBM8A,PPP2R1A,RNPS1,GSPT1,PPP1CC,MAGOHB,PPP2R5C,DAZAP1,PYM1,SSU72,SMG7,PABPN1,PPP2R2A,PPP2R3C,NXT1,CPSF7,NCBP1,DDX19B,WDR33,NUDT21,ACIN1,CPSF3,CSTF2,UPF3A,CSTF2T,SRRM1,HBS1L,SMG1,CSTF1,DDX39B,PPP2R5B,SMG5,FUS,PABPC4,CLP1,EIF4A3,RNGTT,MSI2,FIP1L1,PCF11,CASC3,SMG6,ALYREF,CPSF2,DDX19A,CPSF4 |
| Viral myocarditis | 0.0001 | 0.0059 | 0.5384 | 0.7838 | 1.4561 | 57 | PRF1,HLA-B,HLA-DRB1,HLA-DRA,HLA-A,HLA-DRB5,ACTG1,FYN,HLA-DQB1,HLA-DPB1,ACTB,ITGB2,SGCB,HLA-DPA1,HLA-G,RAC2,CD55,HLA-F,RAC1,EIF4G1,HLA-C,CASP3,CYCS,HLA-DQA1,EIF4G2,CASP8,ABL2,CD86,BID,HLA-E,HLA-DMA,RAC3,DAG1,HLA-DMB,CXADR |
| Cardiac muscle contraction | 0.0003 | 0.0148 | 0.4985 | 0.7376 | 1.3917 | 83 | TPM4,UQCR10,COX5A,COX6A1,COX8A,UQCRH,COX7B,COX4I1,COX6C,UQCRFS1,COX7A2,ATP2A2,UQCRQ,COX6B1,COX5B,COX7A2L,ATP1B3,UQCR11,TPM3,ASPH,UQCRC1,UQCRB,UQCRHL,CYC1,UQCRC2 |
| Ribosome biogenesis in eukaryotes | 0.003 | 0.1087 | 0.4317 | 0.7063 | 1.3313 | 79 | NXT2,RAN,CSNK2A2,CSNK2B,NOP10,NOB1,NHP2,SPATA5,SNU13,UTP6,RPP25L,XRN2,SBDS,RMRP,REXO5,RIOK1,RBM28,NOP56,GNL2,RIOK2,UTP18,POP7,EIF6,RPP30,REXO2,TCOF1,GTPBP4,DKC1,NXT1,NMD3,GAR1,XRN1,AK6,UTP4,IMP3,XPO1,IMP4,EMG1,LSG1,POP5,POP4,NVL,RRP7A,GNL3,MDN1 |
| Autoimmune thyroid disease | 0.0034 | 0.1087 | 0.4317 | 0.7404 | 1.3583 | 50 | GZMB,PRF1,HLA-B,HLA-DRB1,HLA-DRA,HLA-A,HLA-DRB5,HLA-DQB1,HLA-DPB1,CTLA4,HLA-DPA1,HLA-G,HLA-F,HLA-C,HLA-DQA1,CD86,HLA-E,HLA-DMA,TSHR,FAS,HLA-DMB,FASLG,CD28 |
| Staphylococcus aureus infection | 0.005 | 0.1357 | 0.407 | 0.6888 | 1.306 | 91 | HLA-DRB1,HLA-DRA,FCGR3A,HLA-DRB5,HLA-DQB1,HLA-DPB1,ITGB2,HLA-DPA1,SELPLG,HLA-DQA1,KRT10,HLA-DMA,FCGR3B,ITGAM,HLA-DMB,FCGR2B |
| Protein export | 0.0063 | 0.1512 | 0.407 | 0.8303 | 1.4084 | 23 | SEC61B,SRP19,SRP14,SRP72,SPCS3,SEC61G,SPCS1,SEC11A,SRP9,OXA1L,SEC11C,SPCS2,SEC61A1,SRPRB,SRP68,SEC63,SEC62,HSPA5 |
| Type I diabetes mellitus | 0.0077 | 0.1636 | 0.407 | 0.7543 | 1.3615 | 41 | GZMB,PRF1,HLA-B,HLA-DRB1,HLA-DRA,HLA-A,HLA-DRB5,HLA-DQB1,HLA-DPB1,ICA1,HLA-DPA1,HLA-G,HLA-F,HLA-C,IFNG,HSPD1,PTPRN2,HLA-DQA1,CD86,HLA-E,HLA-DMA,FAS,IL12A,HLA-DMB,FASLG |
| Long-term potentiation | 0.0152 | 0.2895 | 0.3807 | 0.6923 | 1.2954 | 67 | PRKCB,RAP1B,CALM3,PPP1CA,PRKACA,KRAS,RAF1,CALM1,ATF4,PRKACB,PPP1CC,PPP3CC,CAMK4,MAP2K2,CAMK2D,HRAS,RPS6KA1,RPS6KA3,PPP3CB,CALM2,MAP2K1,MAPK1,RAP1A |
| RNA degradation | 0.0224 | 0.3425 | 0.3525 | 0.6702 | 1.2632 | 79 | LSM1,BTG3,LSM2,LSM6,CNOT8,PFKP,LSM7,EXOSC4,CNOT7,CNOT4,HSPD1,LSM3,LSM8,EXOSC3,EXOSC2,LSM5,CNOT6L,EXOSC7,ENO1,PARN,XRN2,C1D,DIS3,DCPS,LSM4,TTC37,DHX36,CNOT3,ZCCHC7,XRN1,PNPT1,DCP2,WDR61,PAN2,PABPC4,CNOT1,EXOSC8,DCP1B,MPHOSPH6,TOB2,EDC4,HSPA9,EXOSC1 |
| Taurine and hypotaurine metabolism | 0.0265 | 0.3425 | 0.3525 | -0.9161 | -1.6909 | 11 | GGT7,ADO,GGT1,CSAD |
| Nicotine addiction | 0.0265 | 0.3425 | 0.2879 | 0.7314 | 1.3157 | 40 | CACNA1A,CHRNA6,GABRD |
| Cytosolic DNA-sensing pathway | 0.0266 | 0.3425 | 0.282 | 0.6878 | 1.2803 | 61 | PYCARD,NFKBIA,IRF3,POLR2E,CCL4,ADAR,POLR3E,CCL5,IKBKB,IRF7,POLR2H,CGAS,IKBKG,POLR2L,POLR3B,CASP1,POLR1D,POLR2F,DDX58,POLR3GL,NFKBIB,POLR3H,POLR3F,POLR3K,POLR3A,POLR3C,TBK1,CXCL10,MAVS,ZBP1,POLR3D |
| Legionellosis | 0.0277 | 0.3425 | 0.2765 | 0.7005 | 1.3012 | 57 | PYCARD,MYD88,ITGB2,NFKBIA,RAB1B,CASP3,CYCS,HSPD1,ARF1,HSF1,BCL2L13,CASP8,RAB1A,SAR1B,SEC22B,NAIP,CASP1,CASP7,HSPA2,VCP,SAR1A,HBS1L,EEF1G,HSPA6,ITGAM,HSPA1L,IL12A,NFKB2,TLR5 |
| Renin-angiotensin system | 0.0289 | 0.3425 | 0.3525 | -0.6397 | -1.438 | 23 | LNPEP,NLN,ATP6AP2,PRCP,CTSA |
| Antigen processing and presentation | 0.0305 | 0.3425 | 0.2617 | 0.6782 | 1.272 | 69 | CD74,HLA-B,HLA-DRB1,HLA-DRA,CD8B,CD8A,PSME2,B2M,TAP1,HLA-A,PSME1,HLA-DRB5,HLA-DQB1,HLA-DPB1,HLA-DPA1,KLRC1,HLA-G,HLA-F,CIITA,HLA-C,IFNG,CALR,KLRD1,HLA-DQA1,HSPA4,RFX5,HSP90AA1,RFXANK,TAPBP,HSPA2,HLA-E,CANX,PSME3,HLA-DMA,CTSS,HSPA5,HSPA6,HSPA1L,PDIA3,HLA-DMB |
| Central carbon metabolism in cancer | 0.0325 | 0.3447 | 0.253 | 0.6734 | 1.2626 | 70 | MTOR,IDH2,LDHB,PFKP,KRAS,SLC7A5,RAF1,PTEN,LDHA,PIK3CD,MAP2K2,NTRK1,PGAM1,HRAS,G6PD,HIF1A,PIK3CB,TP53,HK1,PDHB,SIRT6,SCO2,HKDC1,MAP2K1,MAPK1,PKM,MYC,PDHA1,TIGAR,IDH1,GLS,NRAS,GLS2,AKT2,PFKL |
| Basal transcription factors | 0.0345 | 0.3466 | 0.2489 | 0.7176 | 1.3029 | 44 | TAF10,TAF5,TBPL1,GTF2B,GTF2H5,TAF9,TAF4,TAF11,GTF2H1,GTF2A2,GTF2F2,TAF3,TAF6,TAF12,GTF2I,TAF15,GTF2H3,GTF2H2C,CDK7,GTF2E2,GTF2F1,ERCC2,GTF2A1,TAF2,ERCC3 |
| Citrate cycle (TCA cycle) | 0.0436 | 0.4059 | 0.225 | 0.7384 | 1.2916 | 30 | IDH2,SUCLG1,DLD,SDHC,IDH3B,MDH2,IDH3A,SUCLA2,SDHD,SDHB,MDH1,CS,OGDH,SDHA,FH,IDH3G,PDHB,PCK2,PDHA1,IDH1,ACO2,ACLY,OGDHL |
Downloading entire data
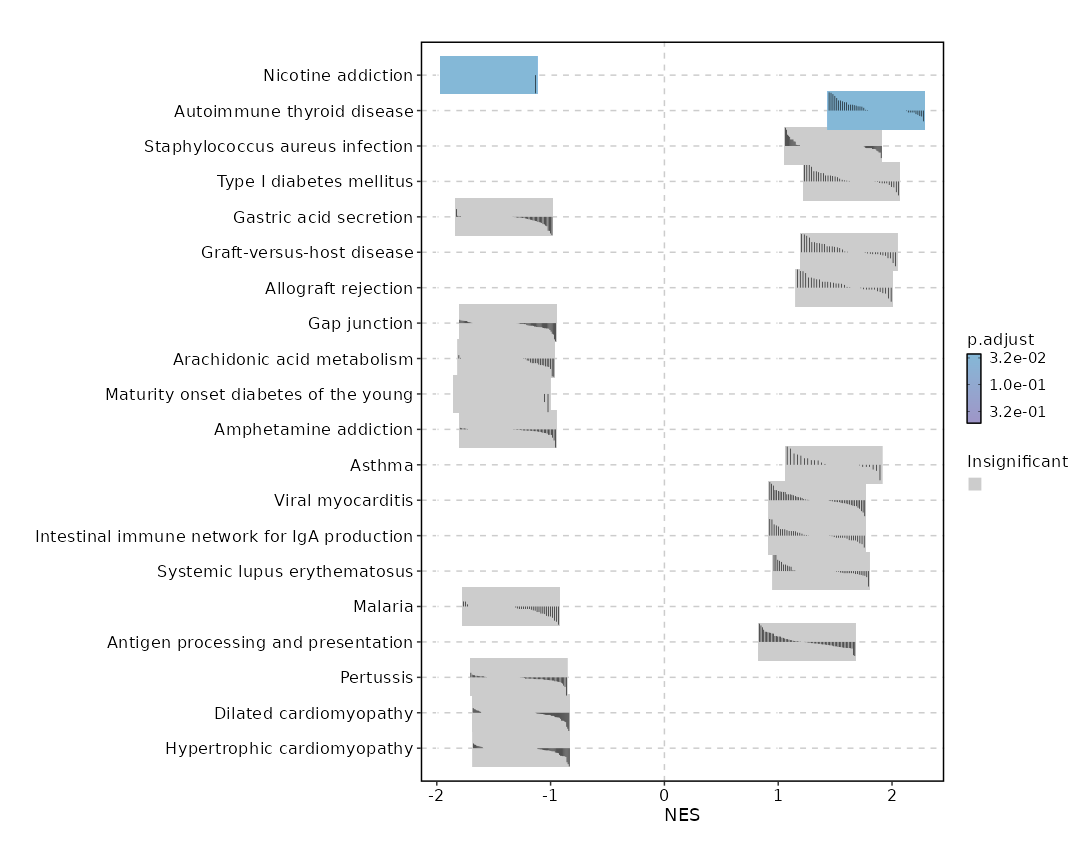
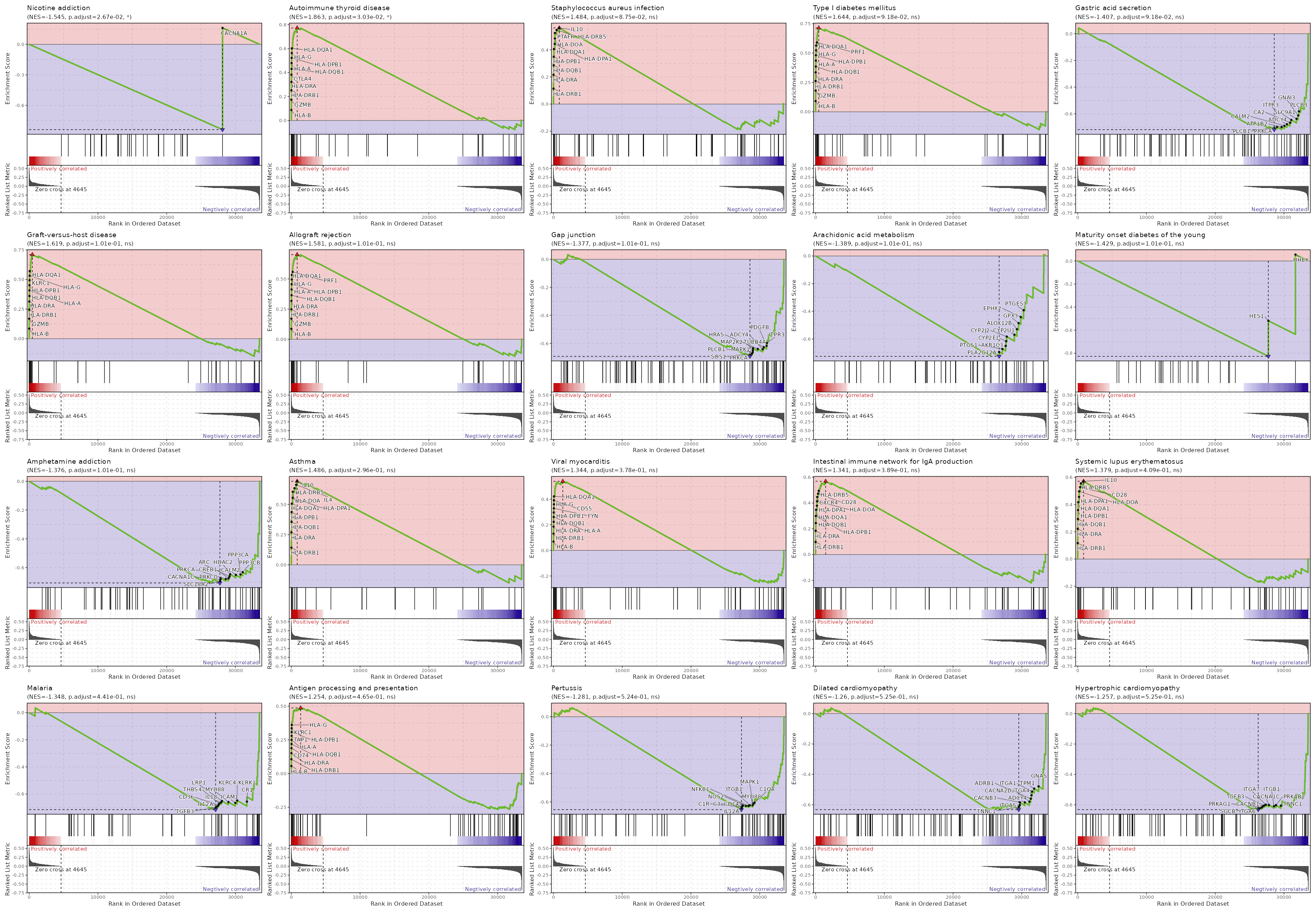
c7
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Nicotine addiction | 0.0001 | 0.0267 | 0.5188 | -0.8385 | -1.5446 | 40 | CACNA1A |
| Autoimmune thyroid disease | 0.0003 | 0.0303 | 0.4985 | 0.773 | 1.8626 | 50 | HLA-B,GZMB,HLA-DRB1,HLA-DRA,CTLA4,HLA-DQB1,HLA-A,HLA-DPB1,HLA-G,HLA-DQA1,PRF1,HLA-DPA1,HLA-DOA,HLA-E,CD28,HLA-DRB5,IL4,IL10,HLA-F |
| Staphylococcus aureus infection | 0.0014 | 0.0875 | 0.4551 | 0.5618 | 1.4844 | 91 | HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-DPB1,HLA-DQA1,HLA-DPA1,HLA-DOA,PTAFR,HLA-DRB5,IL10 |
| Type I diabetes mellitus | 0.0024 | 0.0918 | 0.4317 | 0.7142 | 1.6442 | 41 | HLA-B,GZMB,HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-A,HLA-DPB1,HLA-G,HLA-DQA1,PRF1,HLA-DPA1,HLA-DOA,HLA-E,CD28,HLA-DRB5 |
| Gastric acid secretion | 0.0024 | 0.0918 | 0.4317 | -0.7184 | -1.4075 | 76 | ACTB,ATP1A1,ACTG1,CALM1,PRKACB,ATP1B1,GNAQ,KCNQ1,GNAS,PLCB3,GNAI3,SLC9A1,ITPR3,CA2,ADCY4,CALM2,ATP1B2,PLCB1,PRKCA |
| Graft-versus-host disease | 0.004 | 0.1009 | 0.407 | 0.7107 | 1.6189 | 38 | HLA-B,GZMB,HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-A,HLA-DPB1,KLRC1,HLA-G,HLA-DQA1,PRF1,HLA-DPA1,HLA-DOA,HLA-E,CD28,HLA-DRB5 |
| Allograft rejection | 0.005 | 0.1009 | 0.407 | 0.7022 | 1.5807 | 35 | HLA-B,GZMB,HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-A,HLA-DPB1,HLA-G,HLA-DQA1,PRF1,HLA-DPA1,HLA-DOA,HLA-E,CD28,HLA-DRB5,IL4,IL10,HLA-F |
| Gap junction | 0.0051 | 0.1009 | 0.407 | -0.692 | -1.3773 | 88 | TUBB,TUBA1B,TUBA1A,TUBA1C,PRKACB,GNAQ,GNAS,TUBB2A,TUBB4B,RAF1,PLCB3,GNAI3,MAPK7,MAP2K5,ADRB1,CDK1,ITPR3,PDGFB,TUBB4A,ADCY4,HRAS,MAP2K2,MAPK1,PLCB1,SOS1,PRKCA |
| Arachidonic acid metabolism | 0.0054 | 0.1009 | 0.407 | -0.7253 | -1.3888 | 58 | LTC4S,GGT1,TBXAS1,CBR1,CBR3,PTGES3,EPHX2,GPX3,ALOX12B,CYP2U1,CYP2J2,CYP2E1,AKR1C3,PTGS1,PLA2G12A |
| Maturity onset diabetes of the young | 0.0058 | 0.1009 | 0.407 | -0.8268 | -1.4292 | 26 | HHEX,HES1 |
| Amphetamine addiction | 0.0058 | 0.1009 | 0.407 | -0.7065 | -1.3756 | 69 | FOS,FOSB,CALM1,PRKACB,JUN,PPP3R1,PPP1CC,HDAC1,GNAS,ATF6B,STX1A,PPP3CB,PPP3CA,CALM2,HDAC2,ARC,CREB1,PRKCA,PRKCG,CACNA1C,SLC18A2 |
| Asthma | 0.0186 | 0.2963 | 0.3525 | 0.6918 | 1.4863 | 28 | HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-DPB1,HLA-DQA1,HLA-DPA1,HLA-DOA,HLA-DRB5,IL4,IL10 |
| Viral myocarditis | 0.0257 | 0.3782 | 0.3525 | 0.5402 | 1.3437 | 57 | HLA-B,HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-A,HLA-DPB1,FYN,HLA-G,CD55,HLA-DQA1,PRF1,HLA-DPA1,HLA-DOA,HLA-E,CD28,HLA-DRB5,BID,CASP8,HLA-F |
| Intestinal immune network for IgA production | 0.0285 | 0.3895 | 0.3525 | 0.5686 | 1.3407 | 45 | HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-DPB1,HLA-DQA1,HLA-DPA1,HLA-DOA,CXCR4,CD28,HLA-DRB5,ICOSLG,IL4,IL10,ICOS,TNFRSF13C |
| Systemic lupus erythematosus | 0.0321 | 0.4085 | 0.3218 | 0.5711 | 1.379 | 51 | HLA-DRB1,HLA-DRA,HLA-DQB1,HLA-DPB1,HLA-DQA1,HLA-DPA1,HLA-DOA,CD28,HLA-DRB5,IL10 |
| Malaria | 0.037 | 0.4411 | 0.2489 | -0.7154 | -1.348 | 50 | TNF,KLRB1,ITGAL,PECAM1,TGFB1,KLRK1,THBS3,ITGB2,CD40LG,CD81,CD40,CR1,KLRC4-KLRK1,ICAM1,MYD88,LRP1,IL18,THBS4,CD36,IL12A,TGFB3 |
| Antigen processing and presentation | 0.0413 | 0.4646 | 0.3218 | 0.4873 | 1.2545 | 69 | HLA-B,HLA-DRB1,HLA-DRA,CD74,HLA-DQB1,HLA-A,TAP1,HLA-DPB1,KLRC1,HLA-G,HLA-DQA1,HLA-DPA1,HLA-DOA,B2M,HLA-E,CD8A,HLA-DRB5,PSME2,TAP2,KIR2DL4,HLA-F |
| Pertussis | 0.0494 | 0.5237 | 0.209 | -0.654 | -1.2812 | 76 | FOS,TNF,CALM1,CFL1,JUN,TICAM1,RELA,RHOA,ITGB2,CASP1,GNAI3,MAPK14,MAPK13,NLRP3,IL23A,CASP3,LY96,TRAF6,CALM2,ITGA5,C5,TICAM2,ITGAM,C1QA,MAPK1,MYD88,ITGB1,NFKB1,NOS2,CD14,C3,C1R,IL12A |
| Dilated cardiomyopathy | 0.0536 | 0.525 | 0.1978 | -0.632 | -1.26 | 95 | ACTB,TNF,ACTG1,PRKACB,TGFB1,TPM3,ITGB7,ITGA3,GNAS,TPM1,ITGA1,ITGA4,ADRB1,CACNA2D2,ADCY4,CACNB3,ITGA5,TNNC1 |
| Hypertrophic cardiomyopathy | 0.0581 | 0.525 | 0.19 | -0.6316 | -1.257 | 90 | ACTB,TNF,ACTG1,TGFB1,TPM3,ITGB7,ITGA3,PRKAB1,PRKAA1,PRKAG2,TPM1,ITGA1,ITGA4,CACNA2D2,CACNB3,ITGA5,TNNC1,PRKAB2,ITGB1,CACNA1C,ITGA7,TGFB3,CACNB1,PRKAG1,SGCB,ITGA6 |
Downloading entire data
c3
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Cardiac muscle contraction | 0 | 0.0086 | 0.5573 | 0.7653 | 1.3594 | 83 | COX7C,COX5A,UQCRH,COX8A,TPM4,UQCRB,COX6C,COX6B1,COX7A2L,COX4I1,UQCRFS1,UQCR10,COX5B,COX7A2,COX6A1,COX7B,UQCRC1,UQCR11,CYC1,UQCRC2,UQCRQ,CACNA2D4,ATP1B3,SLC9A6 |
| Proteasome | 0.0003 | 0.0244 | 0.4985 | 0.811 | 1.4082 | 46 | PSMB9,PSMB8,SEM1,PSMA1,PSMB3,PSME1,PSME2,PSMB1,PSMB10,POMP,PSMB4,PSMC5,PSMA6,PSMB6,PSMC3,PSMF1,PSMA4,PSMB2,PSMD3,PSMD7,PSMD4,PSME3,PSMA7,PSMA2,PSMD9,PSMC1,PSMC2,PSMA5,PSMD6,PSMD13,ADRM1,PSMD8,PSMB5,PSMD1,PSMB7,PSMC6 |
| Primary immunodeficiency | 0.0007 | 0.0412 | 0.4773 | 0.8104 | 1.3831 | 37 | CD3D,TAP1,JAK3,LCK,CD3E,CD8A,IL2RG,ADA,ZAP70,PTPRC,RFX5,CD8B,TNFRSF13C,ICOS,IKBKG,CD79A,RFXAP,TAP2 |
| Nicotine addiction | 0.001 | 0.0467 | 0.4551 | -0.655 | -1.6985 | 40 | GABRD |
| Vibrio cholerae infection | 0.0065 | 0.1885 | 0.407 | 0.7448 | 1.3012 | 50 | SEC61B,ACTG1,ATP6V1G1,GNAS,ARF1,ATP6V0B,ACTB,ATP6V1F,ERO1A,ATP6V0C,ATP6AP1,ATP6V0D1,ATP6V0A2,SEC61G,ATP6V1D,SEC61A2,ATP6V1C1,ATP6V1A,KDELR2,PRKACB,TCIRG1,ATP6V1B2,ATP6V0E1,ATP6V1E1,ATP6V0A1,PLCG2,KDELR1,SLC12A2,PRKACA,ADCY3 |
| mRNA surveillance pathway | 0.0067 | 0.1885 | 0.407 | 0.6886 | 1.2313 | 98 | PABPC1,PPP1CC,NUDT21,PPP2R5C,SSU72,RNPS1,PPP1CA,PABPC1L,GSPT1,ALYREF,PPP2R3C,PPP1CB,MSI2,CSTF2T,PPP2CA,SMG1,ACIN1,DDX19A,NXT1,PPP2R1A,HBS1L,GSPT2,SAP18,SYMPK,NCBP2,PYM1,RBM8A,UPF2,PPP2R5E,PPP2R2A,PELO,WDR33,PPP2R2D,UPF3A,SMG6,CASC3,EIF4A3,CSTF2,GLE1,ETF1,FIP1L1,PNN,CPSF1,PPP2R5A,SRRM1,CSTF1,CPSF6,PPP2R1B,WDR82,CPSF2 |
| Morphine addiction | 0.0076 | 0.1885 | 0.407 | 0.6886 | 1.2292 | 91 | PDE3B,PDE4B,PDE4D,GNG5,GNAS,PDE7B,GNAI2,GNG10,PDE7A,PRKCB,ARRB2,GNB2,GNGT2,GNB1,ARRB1,PDE1B,PRKACB,GRK2,GRK4,GNG4,PDE4A,GRK3,GNG2 |
| Collecting duct acid secretion | 0.0079 | 0.1885 | 0.3807 | 0.8102 | 1.3554 | 27 | ATP6V1G1,ATP6V1F,ATP6V0C,ATP6V0D1,ATP6V0A2,ATP6V1D,ATP6V1C1,ATP6V1A,SLC12A7,TCIRG1,ATP6V1B2,ATP6V0E1,ATP6V1E1,ATP6V0A1 |
| Glycolysis / Gluconeogenesis | 0.0104 | 0.2185 | 0.3807 | 0.7133 | 1.2586 | 66 | GAPDH,LDHB,LDHA,ACSS1,TPI1,PGK1,DLAT,GALM,PGAM1,ALDOA,PDHB,DLD,PCK2,MINPP1,AKR1A1,PGM2,ENO1,PFKP,PFKM,ALDOC,ADH5,ALDH3A2,HK2,PDHA1,ADPGK,HKDC1 |
| Fc gamma R-mediated phagocytosis | 0.0116 | 0.2203 | 0.3807 | 0.6872 | 1.2282 | 96 | RAC2,ARPC2,ARPC3,ARPC1B,LAT,WAS,CFL1,PLPP1,PTPRC,PIK3CD,ARPC5,INPP5D,PRKCB,WASF2,ASAP1,ACTR2,VASP,MARCKSL1,VAV1,PIK3R1,PIP5K1A,GSN,FCGR3A,INPPL1,PIK3R3,ARPC4,AKT2,RPS6KB1,BIN1,RAC1,MAP2K1,GAB2,LIMK1,ARPC1A,PLA2G6,RAF1,FCGR2B,RPS6KB2,PIP5K1C,PIP5K1B,JMJD7-PLA2G4B,SPHK2,CFL2,MAPK3,PRKCD,PLCG2,ARF6,AKT3,DNM2,ACTR3,ARPC5L,LIMK2,PIK3CA |
| Porphyrin and chlorophyll metabolism | 0.0141 | 0.2337 | 0.3807 | 0.7402 | 1.2771 | 42 | HMOX1,UROD,COX15,EARS2,GUSB,COX10,PPOX,CPOX,HMOX2,ALAD,UROS,FXN,ALAS1 |
| Phototransduction | 0.0148 | 0.2337 | 0.3807 | 0.7792 | 1.3099 | 28 | CALM3,GNB1,CALM2,PDE6B,SLC24A1,PDE6G,CALM1 |
| Viral myocarditis | 0.0263 | 0.3372 | 0.282 | 0.7086 | 1.2461 | 57 | HLA-B,PRF1,RAC2,CD55,FYN,ACTG1,HLA-A,HLA-DRA,HLA-E,ACTB,EIF4G3,CD28,HLA-DRB1,EIF4G2,HLA-G,HLA-DQB1,EIF4G1,CYCS,HLA-DOA,ABL2,RAC1,CASP3,HLA-DPA1,HLA-DPB1 |
| PD-L1 expression and PD-1 checkpoint pathway in cancer | 0.0271 | 0.3372 | 0.2765 | 0.6756 | 1.2034 | 89 | CD3D,LCK,CD3E,ZAP70,LAT,EML4,CSNK2B,JAK1,STAT1,TICAM1,BATF,MYD88,PIK3CD,CD247,CSNK2A1,NFKBIA,STAT3,CD3G,CD28,MAP3K3,MAP2K3,NRAS,RASGRP1,PDCD1,HIF1A,NFATC3,PRKCQ,IKBKB,KRAS,PTEN,PIK3R1,IKBKG,CD274,PIK3R3,AKT2,RPS6KB1,MAP2K1,NFATC1,PPP3R1,JAK2,RAF1,RPS6KB2,MAPK11,NFKBIB,NFATC2,CSNK2A2,MAPK3 |
| Mitophagy | 0.0282 | 0.3372 | 0.2713 | 0.6938 | 1.2278 | 68 | GABARAP,GABARAPL2,FIS1,CSNK2B,EIF2AK3,USP8,CSNK2A1,UBB,UBA52,CALCOCO2,ATF4,TBC1D15,TAX1BP1,TOMM7,NBR1,NRAS,BECN1,HIF1A,CITED2,RAB7A,GABARAPL1,RPS27A,KRAS,RRAS2,FOXO3,BNIP3L,ATG9B,BNIP3,CSNK2A2,RHOT2,TFE3,OPTN,TBC1D17,E2F1,FUNDC1,JUN,RHOT1,MAPK9,UBC,TP53,SQSTM1,USP15 |
| Autoimmune thyroid disease | 0.0284 | 0.3372 | 0.2713 | 0.7137 | 1.2469 | 50 | CTLA4,HLA-B,GZMB,PRF1,HLA-A,HLA-DRA,HLA-E,CD28,HLA-DRB1,HLA-G,HLA-DQB1,HLA-DOA,HLA-DPA1,HLA-DPB1,IL10 |
| Protein export | 0.0372 | 0.4027 | 0.2413 | 0.7781 | 1.2819 | 23 | SEC61B,OXA1L,SRP14,SPCS1,SEC11A,SRP19,SRP72,SEC61G,SEC61A2,SPCS3,SRP54,SEC11C,SRPRB,SEC62,SRP9,SPCS2,SRP68 |
| Synaptic vesicle cycle | 0.0382 | 0.4027 | 0.2311 | 0.6755 | 1.2006 | 78 | ATP6V1G1,ATP6V0B,ATP6V1F,AP2S1,CLTA,ATP6V0C,NAPA,ATP6V0D1,ATP6V0A2,AP2M1,ATP6V1D,CLTB,STX2,ATP6V1C1,ATP6V1A,NSF,AP2A1,TCIRG1,ATP6V1B2,ATP6V0E1,ATP6V1E1,CLTC,ATP6V0A1,STX1A |
| Bacterial invasion of epithelial cells | 0.0412 | 0.4121 | 0.2221 | 0.685 | 1.2139 | 69 | ARPC2,ARPC3,ACTG1,ARPC1B,RHOG,RHOA,ACTB,PIK3CD,ARPC5,CLTA,HCLS1,WASF2,CBL,ACTR2,CLTB,ELMO1,PIK3R1,PIK3R3,ARPC4,RAC1,CD2AP,CTNNB1,MAD2L2,ELMO2,CLTC,ARPC1A |
| Citrate cycle (TCA cycle) | 0.0542 | 0.4343 | 0.1958 | 0.7386 | 1.2475 | 30 | IDH2,SDHC,MDH2,DLAT,IDH3G,SUCLG1,CS,SDHB,ACO2,PDHB,SDHD,MDH1,DLD,PCK2,DLST,IDH1,OGDH,ACLY,PDHA1 |
Downloading entire data
c1
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
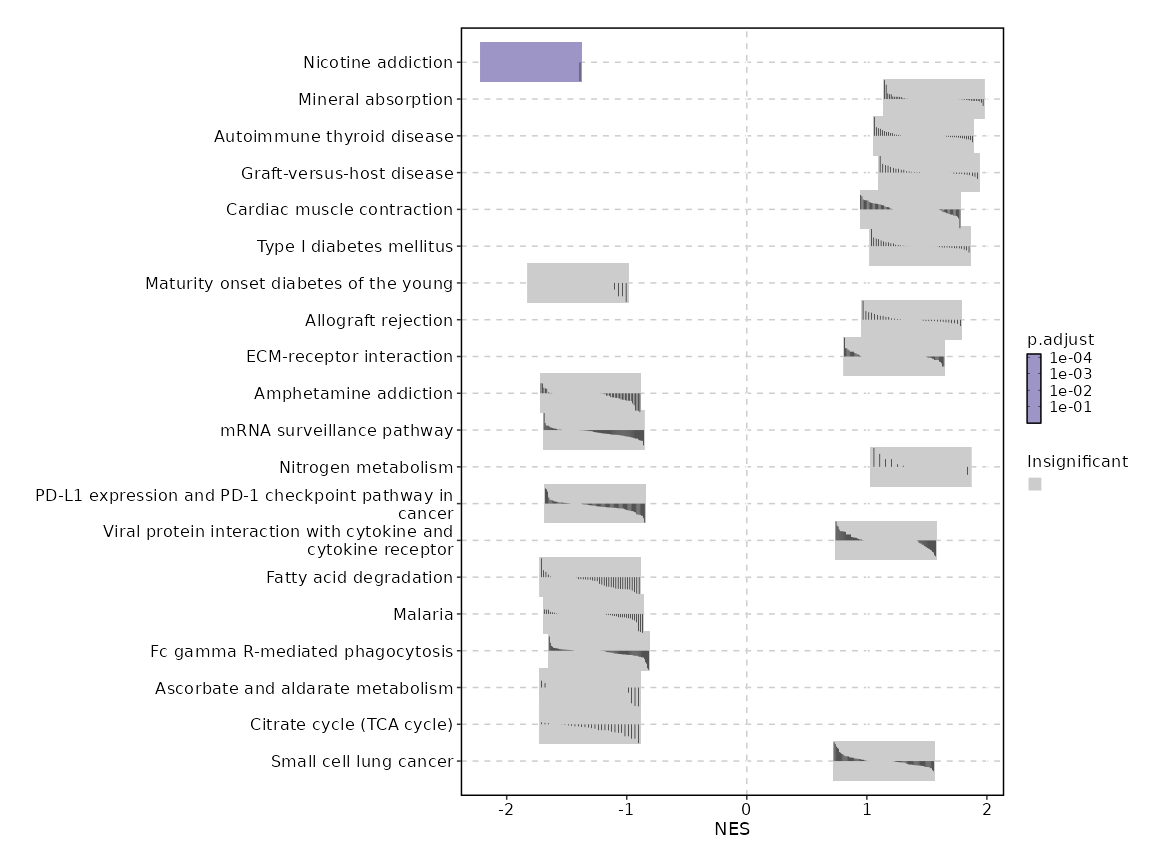
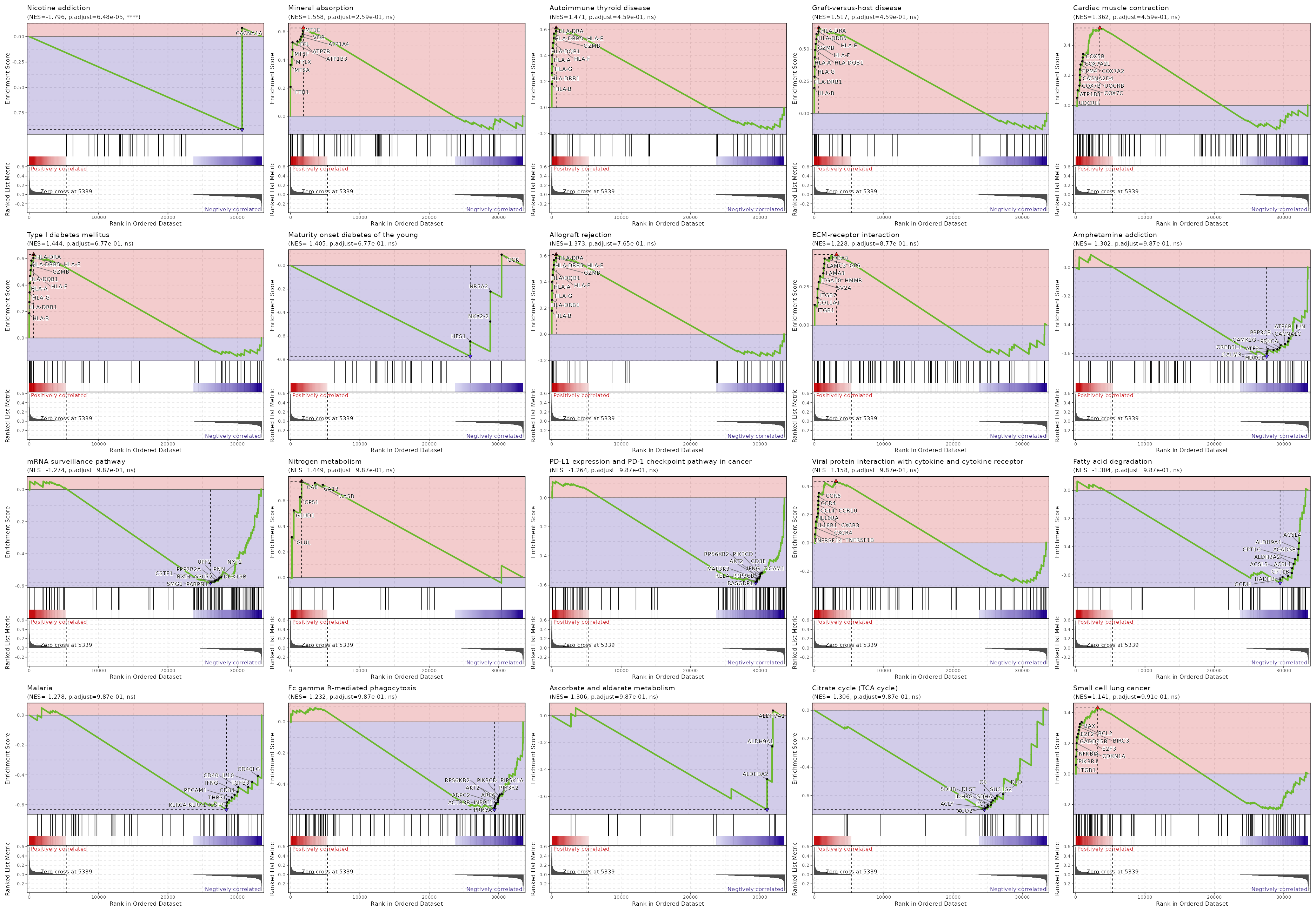
| Nicotine addiction | 0 | 0.0001 | 0.675 | -0.915 | -1.7961 | 40 | CACNA1A |
| Mineral absorption | 0.0027 | 0.259 | 0.4317 | 0.628 | 1.5577 | 60 | FTH1,MT2A,MT1X,MT1F,ATP1B3,FTL,ATP7B,ATP1A4,VDR,MT1E,ATP1B1 |
| Autoimmune thyroid disease | 0.01 | 0.459 | 0.3807 | 0.6145 | 1.471 | 50 | HLA-B,HLA-DRB1,HLA-G,HLA-A,HLA-DQB1,HLA-F,GZMB,HLA-DRB5,HLA-E,HLA-DRA |
| Graft-versus-host disease | 0.0118 | 0.459 | 0.3807 | 0.6645 | 1.5167 | 38 | HLA-B,HLA-DRB1,HLA-G,HLA-A,HLA-DQB1,HLA-F,GZMB,HLA-DRB5,HLA-E,HLA-DRA |
| Cardiac muscle contraction | 0.012 | 0.459 | 0.3807 | 0.5151 | 1.3623 | 83 | UQCRH,ATP1B3,COX7C,COX7B,CACNA2D4,UQCRB,TPM4,COX7A2,COX7A2L,COX5B,ATP1A4,COX6A1,COX8A,ATP1B1,COX6B1,UQCRQ,UQCR10,UQCRHL,UQCRFS1,COX4I1,TPM3,CACNA1F,SLC9A6,UQCRC2 |
| Type I diabetes mellitus | 0.0224 | 0.6773 | 0.3525 | 0.6307 | 1.4442 | 41 | HLA-B,HLA-DRB1,HLA-G,HLA-A,HLA-DQB1,HLA-F,GZMB,HLA-DRB5,HLA-E,HLA-DRA |
| Maturity onset diabetes of the young | 0.0248 | 0.6773 | 0.3525 | -0.7726 | -1.4052 | 26 | GCK,NR5A2,NKX2-2,HES1 |
| Allograft rejection | 0.0321 | 0.7654 | 0.3218 | 0.6088 | 1.3735 | 35 | HLA-B,HLA-DRB1,HLA-G,HLA-A,HLA-DQB1,HLA-F,GZMB,HLA-DRB5,HLA-E,HLA-DRA |
| ECM-receptor interaction | 0.0413 | 0.8771 | 0.3218 | 0.4598 | 1.2279 | 88 | ITGB1,COL1A1,ITGB7,SV2A,ITGA10,HMMR,LAMA3,LAMC3,GP6,ITGA3,ITGA2,THBS3,CD44 |
| Amphetamine addiction | 0.0568 | 0.9868 | 0.2066 | -0.62 | -1.3024 | 69 | FOS,PRKACB,PPP3R1,FOSB,PPP1CC,CREB1,HDAC2,CAMK2D,CREB3L4,CREB3L2,CAMK4,JUN,CACNA1C,ATF6B,PPP3CB,PRKCA,CAMK2G,CREB3L1,ATF2,CALM3,HDAC1 |
| mRNA surveillance pathway | 0.0631 | 0.9868 | 0.19 | -0.5821 | -1.2745 | 98 | FUS,FIP1L1,PPP2CA,PPP1CC,GLE1,PCF11,SMG6,PPP2R5D,PPP2R5A,SYMPK,PELO,PPP2R5E,EIF4A3,CPSF3,NCBP2,TARDBP,CPSF7,CPSF6,PPP2R2B,CPSF2,DDX19A,WDR82,DDX39B,UPF3A,CSTF2T,ACIN1,CPSF1,SMG7,RNGTT,NUDT21,PAPOLG,CASC3,PPP2R1B,PAPOLA,NCBP1,PPP2R1A,GSPT2,MAGOHB,DDX19B,NXT2,PNN,UPF2,PPP2R2A,SSU72,CSTF1,NXT1,SMG1,PABPN1 |
| Nitrogen metabolism | 0.0657 | 0.9868 | 0.2878 | 0.7538 | 1.4491 | 17 | GLUL,GLUD1,CPS1,CA8,CA5B,CA13 |
| PD-L1 expression and PD-1 checkpoint pathway in cancer | 0.0711 | 0.9868 | 0.1814 | -0.5855 | -1.2637 | 89 | LAT,LCK,FOS,CD4,PPP3R1,PRKCQ,NFATC1,ZAP70,CD28,PTPN6,HIF1A,CD3G,MAPK14,MAP2K6,STAT3,PLCG1,NFKBIE,CSNK2A1,IFNGR1,CD274,PIK3CB,MAPK13,KRAS,JAK1,TRAF6,NFATC2,JUN,CD3D,PIK3R2,TICAM1,CD3E,PIK3CD,IFNG,AKT2,RPS6KB2,MAP3K3,PPP3CB,RELA,RASGRP1 |
| Viral protein interaction with cytokine and cytokine receptor | 0.0754 | 0.9868 | 0.2878 | 0.4365 | 1.1576 | 98 | TNFRSF14,TNFRSF1B,CXCR4,IL18R1,IL10RA,CXCR3,CCL4,CCR10,CCR4,CCR6,CCR2,ACKR4,IL24,CXCL13,CCL3,IL10RB,CCL4L2,CCL5,CCR8,XCL1,CCL20 |
| Fatty acid degradation | 0.0804 | 0.9868 | 0.1767 | -0.6565 | -1.3043 | 43 | CPT1A,ACADVL,CYP2U1,ACSL5,ADH5,HADHA,ALDH7A1,EHHADH,ACSL4,ALDH9A1,ACADSB,CPT1C,ALDH3A2,ACSL1,ACSL3,CPT1B,HADHB,GCDH |
| Malaria | 0.0884 | 0.9868 | 0.1657 | -0.6326 | -1.2778 | 50 | ITGAL,ITGB2,KLRB1,CD40LG,IL10,TGFB3,CD40,IFNG,CD81,PECAM1,THBS1,KLRC4-KLRK1,CSF3 |
| Fc gamma R-mediated phagocytosis | 0.0971 | 0.9868 | 0.1511 | -0.5642 | -1.232 | 96 | LAT,PTPRC,ACTR3,CDC42,ACTR2,ARPC5,PLCG1,MARCKSL1,SPHK2,NCF1,VAV1,ARPC5L,PLPP1,CFL2,PIK3CB,PRKCD,DNM2,RAC2,CFL1,FCGR2B,PIK3R2,PIP5K1A,PIK3CD,AKT2,ARF6,RPS6KB2,ARPC2,INPPL1,ACTR3B,PRKCA |
| Ascorbate and aldarate metabolism | 0.0982 | 0.9868 | 0.1608 | -0.6987 | -1.306 | 30 | ALDH7A1,ALDH9A1,ALDH3A2 |
| Citrate cycle (TCA cycle) | 0.0982 | 0.9868 | 0.1608 | -0.6989 | -1.3064 | 30 | MDH1,ACO1,DLAT,PCK2,IDH3B,FH,OGDH,PDHA1,PDHB,SDHD,SUCLA2,SUCLG1,DLD,SUCLG2,CS,SDHA,DLST,SDHB,IDH3G,PC,ACLY,ACO2 |
| Small cell lung cancer | 0.1231 | 0.9913 | 0.2878 | 0.4309 | 1.141 | 92 | ITGB1,PIK3R1,NFKBIA,CDKN1A,GADD45B,E2F3,E2F2,BIRC3,BCL2,BAX,NOS2,LAMA3,LAMC3,PTEN,CDK2,ITGA3,CDKN1B,AKT1,ITGA2,RXRA,BCL2L1,FHIT,MAX,DDB2,GADD45A |
Downloading entire data
c4
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Nicotine addiction | 0 | 0.0008 | 0.6105 | 0.8881 | 1.6646 | 40 | CACNA1A,GABRD |
| Proteasome | 0.0001 | 0.0077 | 0.5384 | 0.8351 | 1.594 | 46 | PSMB9,PSME1,PSME2,PSMB8,PSMB10,PSMA1,POMP,PSMA5,SEM1,PSMB2,PSMB6,PSMA4,PSMB3,IFNG,PSMA2,PSMD9,PSMB4,PSMD8,PSMB1,PSMB7,PSME4,PSMA7,PSMD13,ADRM1,PSMC4,PSMF1,PSMD7,PSMC5,PSMC1,PSMA6,PSMC3,PSMA3,PSMC6,PSMC2,PSMD4,PSMD14,PSMB5 |
| Cytosolic DNA-sensing pathway | 0.0011 | 0.0674 | 0.4551 | 0.7501 | 1.4698 | 61 | CCL5,PYCARD,IRF7,ADAR,AIM2,NFKBIA,POLR2L,IKBKG,CCL4,POLR1D,CGAS,IKBKB,ZBP1,POLR2K,POLR3GL,DDX58,IKBKE,IRF3,TBK1,POLR2F,NFKBIB,CXCL10,POLR3E,CHUK,RIPK3,POLR2E,POLR3K,POLR3D |
| Autoimmune thyroid disease | 0.0015 | 0.0674 | 0.4551 | 0.7755 | 1.4983 | 50 | HLA-B,HLA-A,HLA-DRA,HLA-DRB1,HLA-DPA1,HLA-E,HLA-G,HLA-DPB1,GZMB,HLA-F,HLA-C,PRF1,HLA-DRB5,CD28,TSHR,HLA-DQB1,FASLG |
| Graft-versus-host disease | 0.0018 | 0.0674 | 0.4551 | 0.8055 | 1.4936 | 38 | HLA-B,HLA-A,HLA-DRA,HLA-DRB1,HLA-DPA1,HLA-E,HLA-G,HLA-DPB1,GZMB,HLA-F,HLA-C,IFNG,PRF1,HLA-DRB5,CD28,KLRD1,HLA-DQB1,FASLG |
| Antigen processing and presentation | 0.0023 | 0.0689 | 0.4317 | 0.7314 | 1.4479 | 69 | HLA-B,B2M,CD74,TAP1,PSME1,PSME2,HLA-A,CD8A,HLA-DRA,HLA-DRB1,HLA-DPA1,CD8B,HLA-E,HLA-G,HLA-DPB1,HLA-F,HLA-C,TAPBP,IFNG,CTSS,HLA-DRB5,NFYC,KLRD1,TAP2,IFI30,HLA-DQB1,HSP90AA1,RFX5 |
| Cardiac muscle contraction | 0.0027 | 0.0689 | 0.4317 | 0.7208 | 1.4419 | 83 | UQCR10,COX7A2,COX5A,TPM4,UQCR11,COX6C,COX7B,COX8A,COX6B1,UQCRQ,COX7C,UQCRFS1,UQCRH,COX5B,COX4I1,COX7A2L,COX6A1,ATP1B3,UQCRB,CYC1,CACNA2D4,ATP2A2,UQCRC2,TPM3 |
| Viral myocarditis | 0.0029 | 0.0689 | 0.4317 | 0.7501 | 1.4648 | 57 | HLA-B,HLA-A,HLA-DRA,HLA-DRB1,HLA-DPA1,RAC2,CD55,HLA-E,HLA-G,HLA-DPB1,HLA-F,FYN,HLA-C,ACTG1,PRF1,EIF4G3,HLA-DRB5,CD28,BID,RAC1,HLA-DQB1,ITGB2,DMD,HLA-DOB,CYCS,ITGAL,ACTB |
| Primary immunodeficiency | 0.0042 | 0.0891 | 0.407 | 0.7746 | 1.431 | 37 | TAP1,CD8A,IL7R,ADA,CD8B,LCK,ICOS,CD3D,CD3E,JAK3,IL2RG,ORAI1,IKBKG,TAP2,ZAP70,RFX5,CIITA,RFXANK |
| Type I diabetes mellitus | 0.0058 | 0.1115 | 0.407 | 0.7638 | 1.4371 | 41 | HLA-B,HLA-A,HLA-DRA,HLA-DRB1,HLA-DPA1,HLA-E,HLA-G,HLA-DPB1,GZMB,HLA-F,HSPD1,HLA-C,IFNG,PRF1,HLA-DRB5,CD28,HLA-DQB1,FASLG,PTPRN2,FAS,HLA-DOB |
| Maturity onset diabetes of the young | 0.0095 | 0.1476 | 0.3807 | -0.7757 | -1.6623 | 26 | HES1,NR5A2,NKX2-2,GCK |
| Morphine addiction | 0.0098 | 0.1476 | 0.3807 | 0.6714 | 1.3477 | 91 | PDE3B,ARRB2,PDE4B,GNG5,PDE4D,PRKCB,GRK2,GNG10,ADCY7,GNAI2,GNB2,PDE7A,GNAS,GNG2,GRK3,CACNA1A,GNAO1,GNB1,ADCY4,GNAI1,PDE7B,GNAI3,ADCY9,GNB4,PDE1B |
| Regulation of lipolysis in adipocytes | 0.01 | 0.1476 | 0.3807 | 0.7115 | 1.3862 | 54 | PDE3B,PIK3R1,ADCY7,GNAI2,PIK3CD,GNAS,TSHR,ADCY4,AKT2,AKT1,MGLL,GNAI1,ABHD5,GNAI3,ADCY9,IRS2,PLIN1,ADRB1,PIK3CA,PRKACB,PRKACA |
| Hematopoietic cell lineage | 0.0123 | 0.1683 | 0.3807 | 0.6634 | 1.338 | 96 | CD8A,HLA-DRA,CD2,IL7R,HLA-DRB1,CD38,HLA-DPA1,CD55,CD8B,ITGA1,HLA-DPB1,CD3D,CD3E,ITGA4,CD7,CD37,HLA-DRB5,FLT3LG,IL7,CD3G,CSF2,TFRC,HLA-DQB1,ITGA2 |
| Allograft rejection | 0.0177 | 0.2251 | 0.3525 | 0.7536 | 1.3865 | 35 | HLA-B,HLA-A,HLA-DRA,HLA-DRB1,HLA-DPA1,HLA-E,HLA-G,HLA-DPB1,GZMB,HLA-F,HLA-C,IFNG,PRF1,HLA-DRB5,CD28,HLA-DQB1,FASLG |
| Non-small cell lung cancer | 0.0443 | 0.5083 | 0.2193 | 0.6508 | 1.2955 | 72 | PIK3R1,JAK3,EML4,STK4,PRKCB,KRAS,BAD,KIF5B,PIK3CD,NRAS,BAK1,E2F1,GRB2,STAT5B,MAP2K1,CDKN2A,CDK6,E2F3,RXRB,E2F2,GADD45B,RB1,PLCG2,AKT2,ARAF,SOS1,AKT1,RET,HRAS,BAX |
| Vasopressin-regulated water reabsorption | 0.0514 | 0.5083 | 0.209 | 0.703 | 1.3345 | 44 | ARHGDIB,DYNLL1,RAB11A,GNAS,RAB5C,AQP3,DCTN6,ARHGDIA,DYNC1I2,DYNC1H1,STX4,DCTN5,CREB3L2,DCTN2,ADCY9,CREB3,CREB1,DYNC1LI2,RAB5A,CREB3L4,DCTN1,PRKACB,PRKACA |
| Synaptic vesicle cycle | 0.0561 | 0.5083 | 0.1938 | 0.637 | 1.2696 | 78 | ATP6V1F,ATP6V0B,AP2S1,CLTB,ATP6V1G1,ATP6V0E1,ATP6V0C,AP2M1,ATP6V0D1,CACNA1A,DNM2,ATP6V1C2,STX2,CLTA,ATP6V1D,STXBP1,NAPA,TCIRG1,STX1A |
| Inflammatory mediator regulation of TRP channels | 0.0582 | 0.5083 | 0.1882 | 0.6205 | 1.2515 | 98 | PIK3R1,PTGER4,PPP1CA,PRKCB,ADCY7,CALM1,PIK3CD,GNAS,CALM3,TRPV2,ITPR1,MAP2K3,PRKCQ,MAPK11,ADCY4,NTRK1,PLCG2,MAP2K6,CALML4,ADCY9,PRKCD,TRPV1,MAPK12,PPP1CC,ITPR2,JMJD7-PLA2G4B,PIK3CA,PRKACB,PLCG1,PRKCH,PRKACA,PRKCE,CALM2,PRKCA,MAPK9 |
| GABAergic synapse | 0.0586 | 0.5083 | 0.1882 | 0.6259 | 1.2545 | 89 | GNG5,GABARAP,PRKCB,GNG10,ADCY7,GNAI2,GNB2,GABARAPL2,GNG2,ABAT,CACNA1A,GNAO1,GNB1,ADCY4,GNAI1,SLC38A1,GNAI3,ADCY9,GNB4,CACNA1D,GNG12,SLC12A5,GNGT2,SLC38A5,GLS,GNB5,PRKACB,PRKACA,PRKCA |
Downloading entire data
c5
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
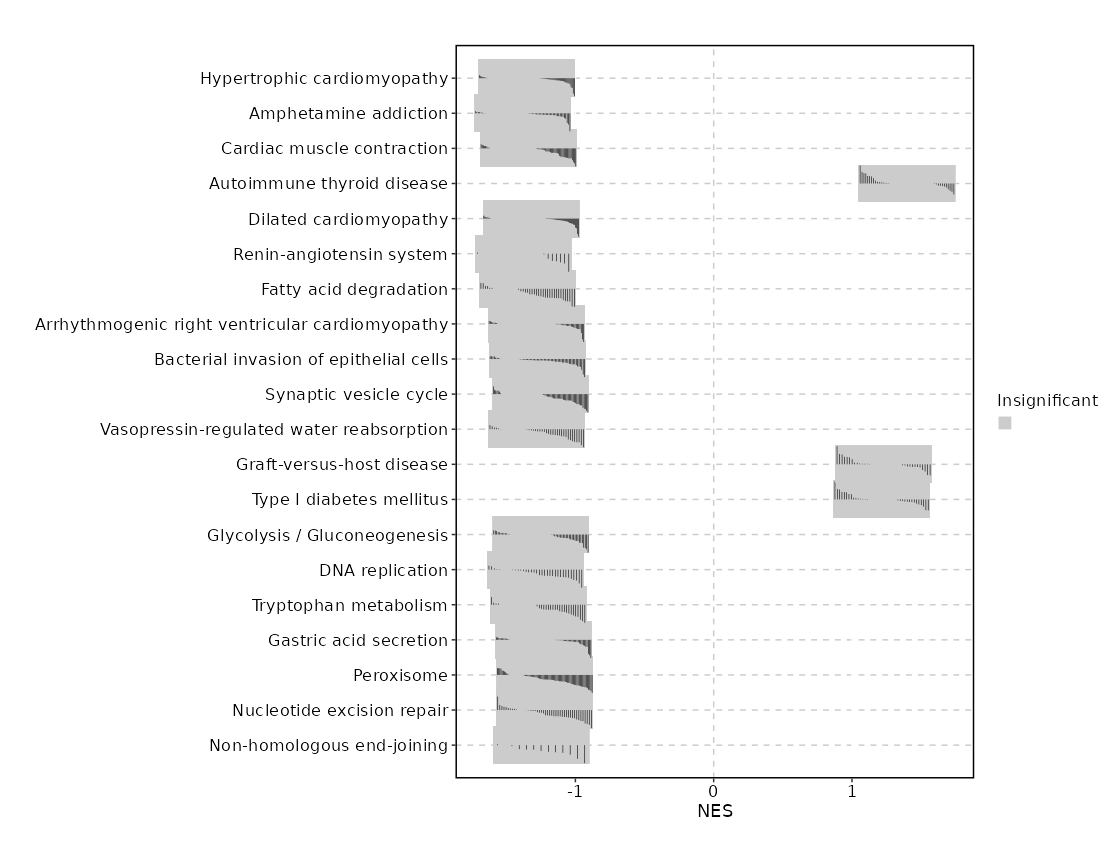
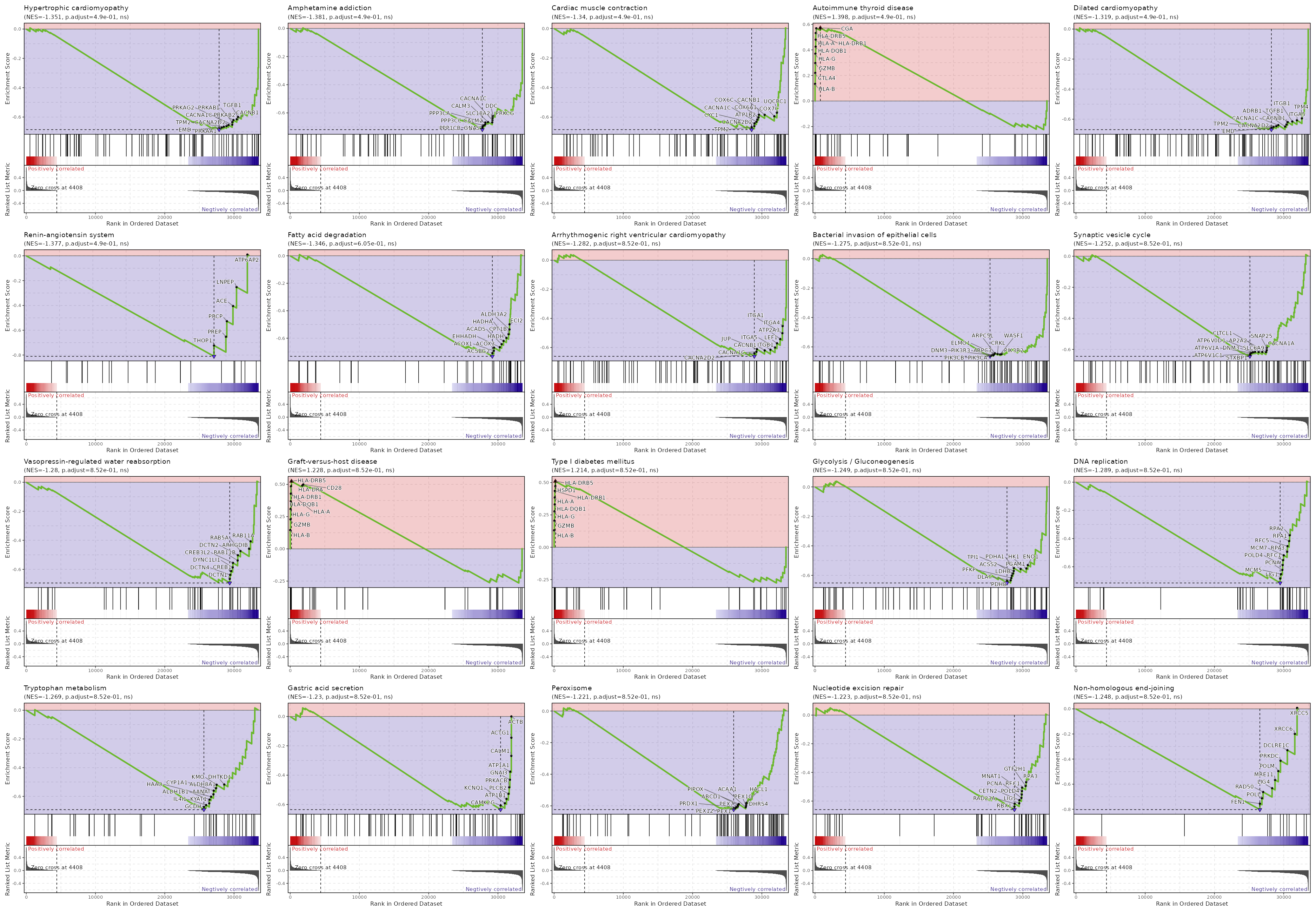
| Hypertrophic cardiomyopathy | 0.0048 | 0.4903 | 0.407 | -0.6859 | -1.3507 | 90 | ACTB,ACTG1,TNF,ITGB7,TPM3,ITGA4,ITGA1,ATP2A3,TPM4,ITGA5,ITGB1,ACE,TGFB1,CACNB1,PRKAB2,PRKAB1,PRKAG2,CACNA1C,CACNA2D2,TPM2,EMD,PRKAA1 |
| Amphetamine addiction | 0.0055 | 0.4903 | 0.407 | -0.719 | -1.3809 | 69 | FOS,CALM1,FOSB,PPP1CA,PRKACB,HDAC1,ATF6B,PPP3R1,JUN,CAMK2G,STX1A,SIRT1,CREB3L2,PPP1CC,CREB1,DDC,PRKCG,CACNA1C,SLC18A2,CALM3,PPP3CA,CALM2,PPP3CC,PPP1CB,GNAS |
| Cardiac muscle contraction | 0.0101 | 0.4903 | 0.3807 | -0.6845 | -1.3396 | 83 | COX4I1,ATP1A1,TPM3,COX6B1,ATP1B1,UQCRB,UQCR10,ATP2A3,TPM4,FXYD2,COX8A,COX5B,UQCRC2,UQCRC1,COX7C,CACNB1,COX6A1,COX6C,ATP1B2,CACNA1C,CYC1,CACNA2D2,TPM2 |
| Autoimmune thyroid disease | 0.0128 | 0.4903 | 0.3807 | 0.5721 | 1.3979 | 50 | HLA-B,CTLA4,GZMB,HLA-G,HLA-DQB1,HLA-A,HLA-DRB1,HLA-DRB5,CGA |
| Dilated cardiomyopathy | 0.0133 | 0.4903 | 0.3807 | -0.6696 | -1.3191 | 95 | ACTB,ACTG1,TNF,ITGB7,TPM3,PRKACB,ITGA4,ITGA1,ATP2A3,TPM4,ITGA5,ITGB1,TGFB1,CACNB1,ADRB1,CACNA1C,CACNA2D2,TPM2,EMD |
| Renin-angiotensin system | 0.0154 | 0.4903 | 0.3807 | -0.81 | -1.377 | 23 | ATP6AP2,LNPEP,ACE,PRCP,PREP,THOP1 |
| Fatty acid degradation | 0.0222 | 0.6045 | 0.3525 | -0.7315 | -1.3455 | 43 | CPT1A,ACADVL,ACSL6,ECHS1,ADH5,ACSL5,ACSL3,ACAA2,ALDH9A1,HADHB,ACADSB,ECI2,ALDH3A2,CPT1B,HADHA,HADH,ACADS,EHHADH,ACOX3,ACOX1,ACSBG2 |
| Arrhythmogenic right ventricular cardiomyopathy | 0.0389 | 0.852 | 0.2344 | -0.6606 | -1.2817 | 77 | ACTB,ACTG1,ITGB7,CTNNA1,ITGA4,ITGA1,ATP2A3,LEF1,ITGA5,ITGB1,CACNB1,JUP,CACNA1C,CACNA2D2 |
| Bacterial invasion of epithelial cells | 0.0491 | 0.852 | 0.209 | -0.6639 | -1.275 | 69 | ACTB,ACTG1,ACTR3,ARPC5,ARPC2,CDC42,RHOA,CLTA,HCLS1,CTNNA1,ILK,ACTR2,RAC1,CLTC,SRC,ITGA5,ARPC4,ITGB1,CRK,PIK3CD,ACTR3B,DNM2,ARPC1B,PIK3R1,DNM1,SHC1,CBL,ELMO3,WASF2,VCL,PTK2,CLTCL1,DOCK1,GAB1,PIK3R2,WASF1,CRKL,ARPC5L,ELMO1,ARPC3,PIK3R3,DNM3,PIK3CB,PIK3CA |
| Synaptic vesicle cycle | 0.0641 | 0.852 | 0.1798 | -0.6443 | -1.2517 | 78 | ATP6V0A1,CLTA,VAMP2,STX3,AP2M1,CLTC,AP2S1,ATP6V0E1,ATP6V1G1,CPLX1,STX1A,RAB3A,DNM2,AP2B1,ATP6V1D,DNM1,SLC18A2,UNC13B,ATP6V1G2,AP2A1,ATP6V1B2,ATP6V1F,ATP6V1E1,SLC32A1,SNAP25,CACNA1A,CLTCL1,SLC6A9,AP2A2,ATP6V0D1,DNM3,ATP6V1A,ATP6V1C1,STXBP1 |
| Vasopressin-regulated water reabsorption | 0.0671 | 0.852 | 0.1814 | -0.6938 | -1.2799 | 44 | RAB5B,PRKACB,DYNC2H1,VAMP2,ARHGDIA,STX4,RAB11A,ARHGDIB,RAB5A,DCTN2,RAB11B,CREB3L2,DYNC1LI1,CREB1,DCTN4,DCTN1 |
| Graft-versus-host disease | 0.0682 | 0.852 | 0.2878 | 0.5241 | 1.2285 | 38 | HLA-B,GZMB,HLA-G,HLA-DQB1,HLA-A,HLA-DRB1,HLA-DRB5,HLA-DRA,CD28 |
| Type I diabetes mellitus | 0.0685 | 0.852 | 0.2878 | 0.5102 | 1.2141 | 41 | HLA-B,GZMB,HLA-G,HLA-DQB1,HLA-A,HLA-DRB1,HSPD1,HLA-DRB5 |
| Glycolysis / Gluconeogenesis | 0.0719 | 0.852 | 0.1709 | -0.6518 | -1.2492 | 66 | PKM,FBP1,ALDOA,PGM2,ADH5,GAPDH,ENO2,ALDH9A1,BPGM,ALDH3A2,GPI,ENO1,HK1,PGAM1,PDHA1,TPI1,ACSS2,LDHB,PFKP,DLAT,PDHB |
| DNA replication | 0.0766 | 0.852 | 0.1709 | -0.7151 | -1.2894 | 36 | POLA1,RNASEH2C,MCM2,MCM4,MCM3,RNASEH1,DNA2,RPA2,RPA1,RFC5,RPA3,MCM7,RFC1,POLD4,PCNA,MCM5,LIG1 |
| Tryptophan metabolism | 0.078 | 0.852 | 0.1682 | -0.6924 | -1.2693 | 42 | CAT,ECHS1,KYAT3,ALDH9A1,ALDH3A2,HADHA,HADH,EHHADH,DLST,DDC,TPH1,DHTKD1,KMO,ALDH8A1,CYP1A1,AANAT,HAAO,ALDH1B1,KYAT1,IL4I1,GCDH |
| Gastric acid secretion | 0.0847 | 0.852 | 0.1552 | -0.6349 | -1.2301 | 76 | ACTB,ACTG1,CALM1,ATP1A1,GNAI3,PRKACB,PLCB2,ATP1B1,KCNQ1,CAMK2G |
| Peroxisome | 0.0932 | 0.852 | 0.1464 | -0.6239 | -1.2212 | 82 | SCP2,CAT,ACSL6,PRDX5,ACSL5,PEX26,MPV17L2,PECR,EPHX2,CROT,ACSL3,HSD17B4,ECI2,ABCD2,PAOX,AGPS,SLC25A17,PEX13,PEX11G,EHHADH,ACOT8,PHYH,DECR2,PXMP2,ACOX3,PEX19,ACOX1,NUDT7,PXMP4,PEX3,MLYCD,PEX16,PEX11B,ABCD3,MPV17,PEX5,HACL1,DHRS4,PEX10,ACAA1,PIPOX,ABCD1,PEX7,PRDX1,PEX12,PEX1 |
| Nucleotide excision repair | 0.1065 | 0.852 | 0.1404 | -0.6598 | -1.2231 | 46 | ERCC5,ERCC1,CDK7,GTF2H2,CUL4A,XPA,GTF2H4,RAD23B,DDB1,CUL4B,RPA2,RPA1,RFC5,RPA3,GTF2H1,RFC1,MNAT1,POLD4,PCNA,CETN2,LIG1,RAD23A,RBX1 |
| Non-homologous end-joining | 0.1129 | 0.852 | 0.1464 | -0.8028 | -1.2483 | 13 | XRCC5,XRCC6,DCLRE1C,PRKDC,POLM,MRE11,LIG4,RAD50,POLL,FEN1 |
Downloading entire data
c11
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
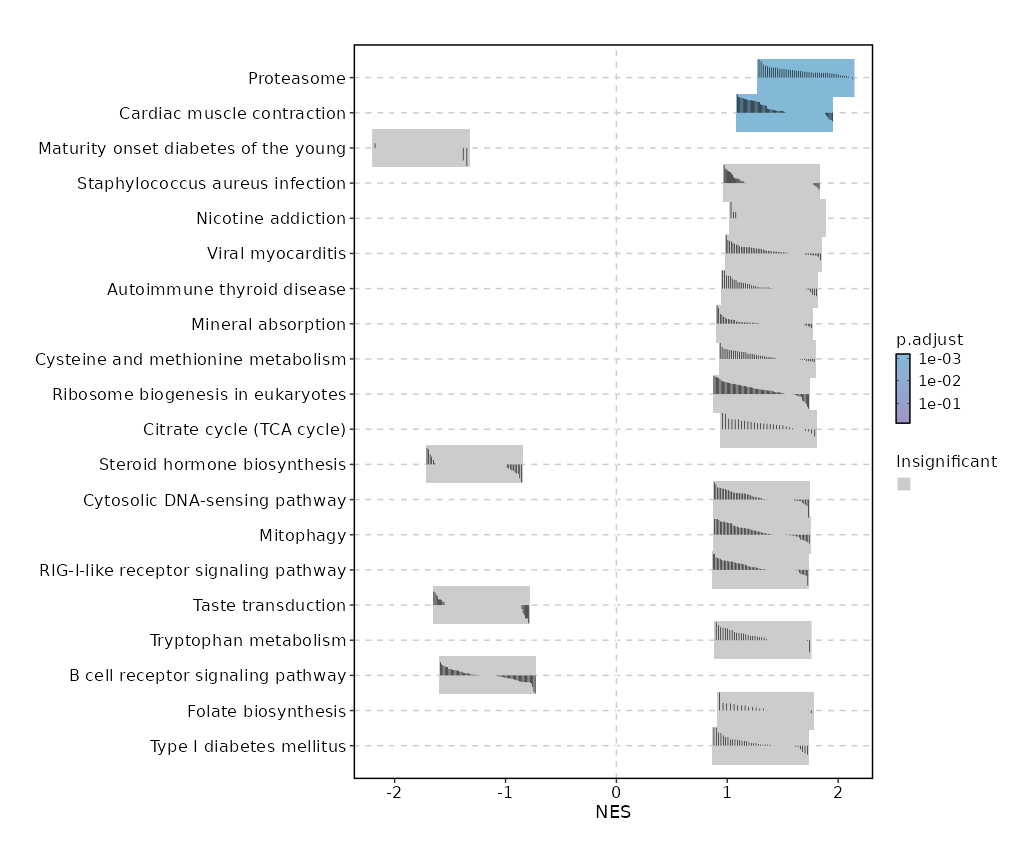
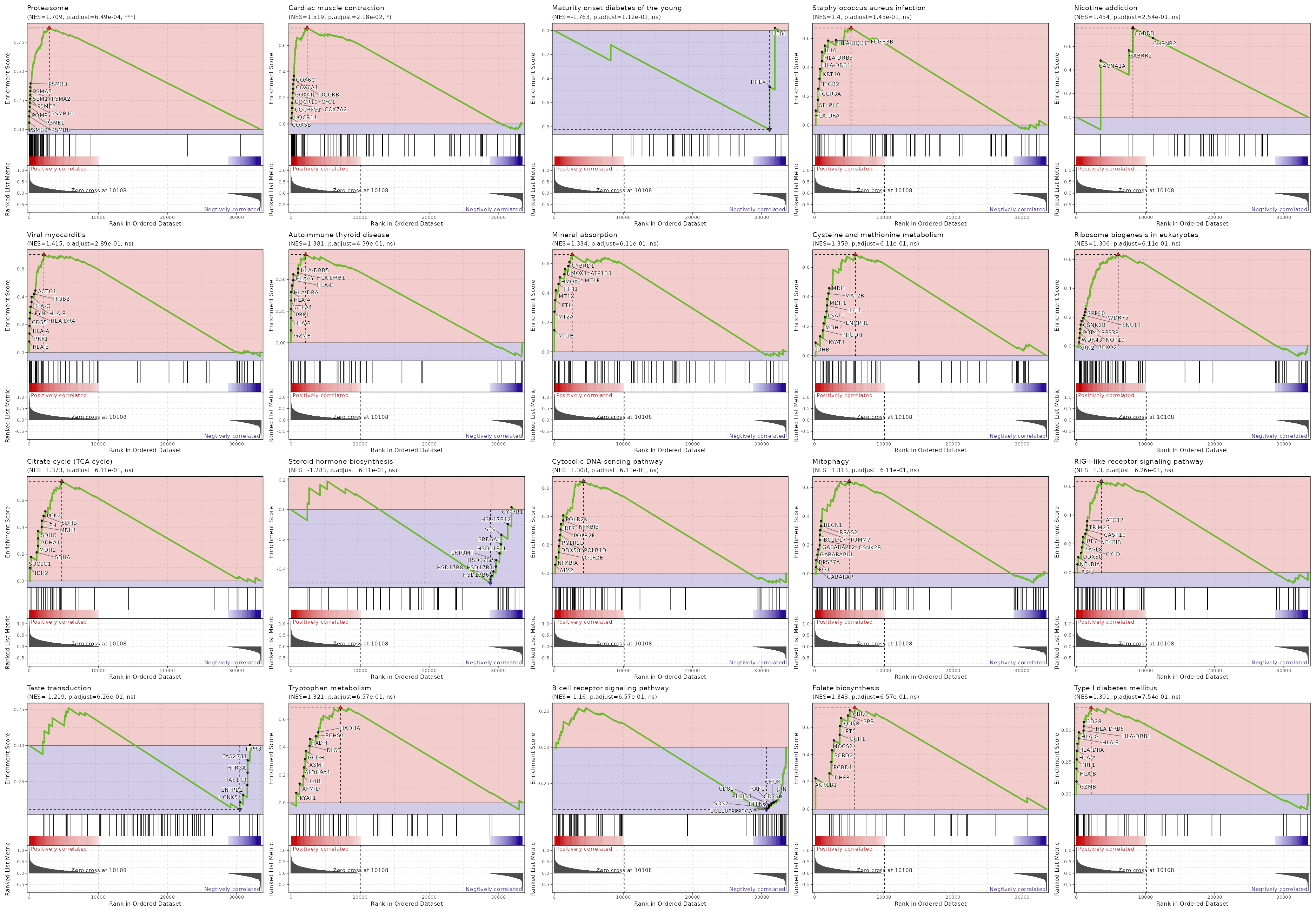
| Proteasome | 0 | 0.0006 | 0.6273 | 0.8722 | 1.7088 | 46 | PSMB9,PSMB8,PSME1,POMP,PSMB10,PSME2,SEM1,PSMA2,PSMA5,PSMB3,PSMB4,PSMD7,PSMA1,PSMA7,PSMB6,PSMB2,PSMC6,PSMA3,PSMA4,PSMB7,PSMB1,PSMD8,IFNG,PSMD4,PSMC1,PSMD12,PSMA6,PSME3,PSMD13,PSME4,ADRM1,PSMC5,PSMD11,PSMD1,PSMB5,PSMC3,PSMC2,PSMD3 |
| Cardiac muscle contraction | 0.0002 | 0.0218 | 0.5188 | 0.7351 | 1.5193 | 83 | COX5B,UQCR11,UQCRFS1,COX7A2,UQCR10,CYC1,COX4I1,COX6A1,UQCRB,COX6C,COX7B,COX6B1,UQCRQ,SLC9A1,COX7C,ATP2A2,UQCRC2,UQCRH,COX8A,COX5A,COX7A2L,UQCRC1,TPM4,UQCRHL,ATP1B3,ATP1A3 |
| Maturity onset diabetes of the young | 0.0018 | 0.1118 | 0.4551 | -0.8238 | -1.7627 | 26 | HES1,HHEX |
| Staphylococcus aureus infection | 0.003 | 0.1445 | 0.4317 | 0.6724 | 1.3996 | 91 | HLA-DRA,SELPLG,FCGR3A,ITGB2,KRT10,HLA-DRB1,HLA-DRB5,IL10,HLA-DQB1,FCGR3B,KRT18,ITGAM,HLA-DPB1,FCGR2B,C5,ITGAL |
| Nicotine addiction | 0.0066 | 0.2538 | 0.407 | 0.7561 | 1.4545 | 40 | CACNA1A,GABRR2,GABRD,CHRNB2 |
| Viral myocarditis | 0.0091 | 0.2894 | 0.3807 | 0.7032 | 1.4151 | 57 | HLA-B,PRF1,HLA-A,CD55,HLA-DRA,FYN,HLA-E,HLA-G,ITGB2,ACTG1,CASP8,CYCS,HLA-DRB1,HLA-DRB5,CD28,ACTB,HLA-F,EIF4G1,CASP3,HLA-DQB1,CASP9,HLA-C,RAC2,EIF4G3,CD80,HLA-DPB1,RAC1 |
| Autoimmune thyroid disease | 0.0161 | 0.4394 | 0.3525 | 0.6969 | 1.3813 | 50 | GZMB,HLA-B,PRF1,CTLA4,HLA-A,HLA-DRA,HLA-E,HLA-G,HLA-DRB1,HLA-DRB5,CD28,HLA-F,IL10,HLA-DQB1,HLA-C |
| Mineral absorption | 0.0323 | 0.6113 | 0.2664 | 0.6612 | 1.3336 | 60 | MT1E,MT2A,FTL,MT1X,FTH1,HMOX2,MT1F,HMOX1,ATP1B3,CYBRD1,ATP1A3,MT1G,ATOX1,SLC11A2,FXYD2,SLC39A4,SLC46A1,ATP2B1,SLC26A6,VDR,SLC31A1,STEAP1 |
| Cysteine and methionine metabolism | 0.0342 | 0.6113 | 0.2617 | 0.6859 | 1.3594 | 50 | LDHB,KYAT1,PHGDH,MDH2,ENOPH1,PSAT1,IL4I1,MDH1,MAT2B,MRI1,BCAT1,APIP,LDHA,MPST,AHCY,TST,CTH,MTR,BCAT2,SRM,GCLM,GCLC,AHCYL1 |
| Ribosome biogenesis in eukaryotes | 0.037 | 0.6113 | 0.245 | 0.6341 | 1.3065 | 79 | XRN2,WDR43,REXO2,POP4,NOP10,CSNK2B,RPP38,SNU13,WDR75,RPP40,IMP3,NOL6,MDN1,GNL3,NOP56,FBL,RRP7A,RAN,AK6,GNL2,REXO1,UTP15,RPP30,RIOK1,CSNK2A2,EIF6,HEATR1,NHP2,UTP18,DKC1,WDR36,NXT2,IMP4,SBDS,EMG1,WDR3,NOB1,TBL3,LSG1,DROSHA,RCL1,XPO1,GTPBP4,CSNK2A1,XRN1,GNL3L |
| Citrate cycle (TCA cycle) | 0.0403 | 0.6113 | 0.2489 | 0.7425 | 1.3725 | 30 | IDH2,SUCLG1,SDHA,MDH2,PDHA1,SDHC,MDH1,FH,SDHB,PCK2,DLST,ACO2,ACLY,DLAT,IDH3A,OGDH,IDH3B,OGDHL,PDHB,SUCLG2 |
| Steroid hormone biosynthesis | 0.0408 | 0.6113 | 0.3218 | -0.4945 | -1.2827 | 61 | CYP7B1,HSD17B12,STS,SRD5A1,HSD11B1L,LRTOMT,HSD17B7,HSD17B1,HSD17B8,HSD17B6 |
| Cytosolic DNA-sensing pathway | 0.0435 | 0.6113 | 0.228 | 0.6479 | 1.3082 | 61 | AIM2,NFKBIA,POLR2E,DDX58,POLR1D,POLR2L,POLR2F,IRF7,NFKBIB,POLR2K,POLR3G,IRF3,RELA,PYCARD,ADAR,CCL4,IKBKE,POLR3GL,RIPK3,CCL5,POLR3D,TBK1,POLR3A,POLR2H |
| Mitophagy | 0.0448 | 0.6113 | 0.2221 | 0.6425 | 1.3126 | 68 | GABARAP,FIS1,RPS27A,GABARAPL1,GABARAPL2,CSNK2B,TBC1D17,TOMM7,RRAS2,BECN1,TFEB,UBA52,ATG9A,BNIP3L,USP15,UBB,RELA,ULK1,CSNK2A2,PGAM5,NRAS,RRAS,ATF4,KRAS,TBK1,RAB7A,AMBRA1,USP8,ATG5,OPTN |
| RIG-I-like receptor signaling pathway | 0.0514 | 0.6258 | 0.2066 | 0.6357 | 1.2998 | 69 | AZI2,NFKBIA,DDX58,CYLD,CASP8,IRF7,NFKBIB,CASP10,TRIM25,ATG12,TANK,TRADD,IRF3,PIN1,RNF125,RELA,TRAF3,SIKE1,IKBKE,IL12A,MAPK12,ISG15,TBK1,DHX58 |
| Taste transduction | 0.0524 | 0.6258 | 0.3218 | -0.4443 | -1.2185 | 85 | ITPR3,TAS2R31,HTR3A,TAS1R3,ENTPD2,KCNK5 |
| Tryptophan metabolism | 0.0634 | 0.657 | 0.19 | 0.6809 | 1.3214 | 42 | KYAT1,AFMID,IL4I1,ALDH9A1,ASMT,GCDH,DLST,HADH,ECHS1,HADHA,EHHADH,DHTKD1,KMO,ACAT2,HAAO,CAT,DDC,ALDH8A1,ACAT1,DLD |
| B cell receptor signaling pathway | 0.0641 | 0.657 | 0.2878 | -0.4303 | -1.1601 | 79 | FOS,LYN,VAV3,LILRB1,VAV2,CD19,BLNK,BTK,CD22,PLCG2,SOS1,CD79A,CR2,GSK3B,AKT3,CD72,JUN,CD79B,CHUK,RAF1,CD81,PIK3R3,PTPN6,SOS2,BCL10,PPP3CA |
| Folate biosynthesis | 0.0654 | 0.657 | 0.1938 | 0.7414 | 1.3426 | 26 | AKR1B1,DHFR,PCBD1,PCBD2,MOCS2,GCH1,PTS,QDPR,SPR,CBR1,DHFR2 |
| Type I diabetes mellitus | 0.0798 | 0.7536 | 0.1682 | 0.6734 | 1.3009 | 41 | GZMB,HLA-B,PRF1,HLA-A,HLA-DRA,HLA-E,HLA-G,HLA-DRB1,HLA-DRB5,CD28,IFNG,HLA-F,HSPD1,HLA-DQB1,HLA-C,IL12A,CD80,ICA1,HLA-DPB1 |
Downloading entire data
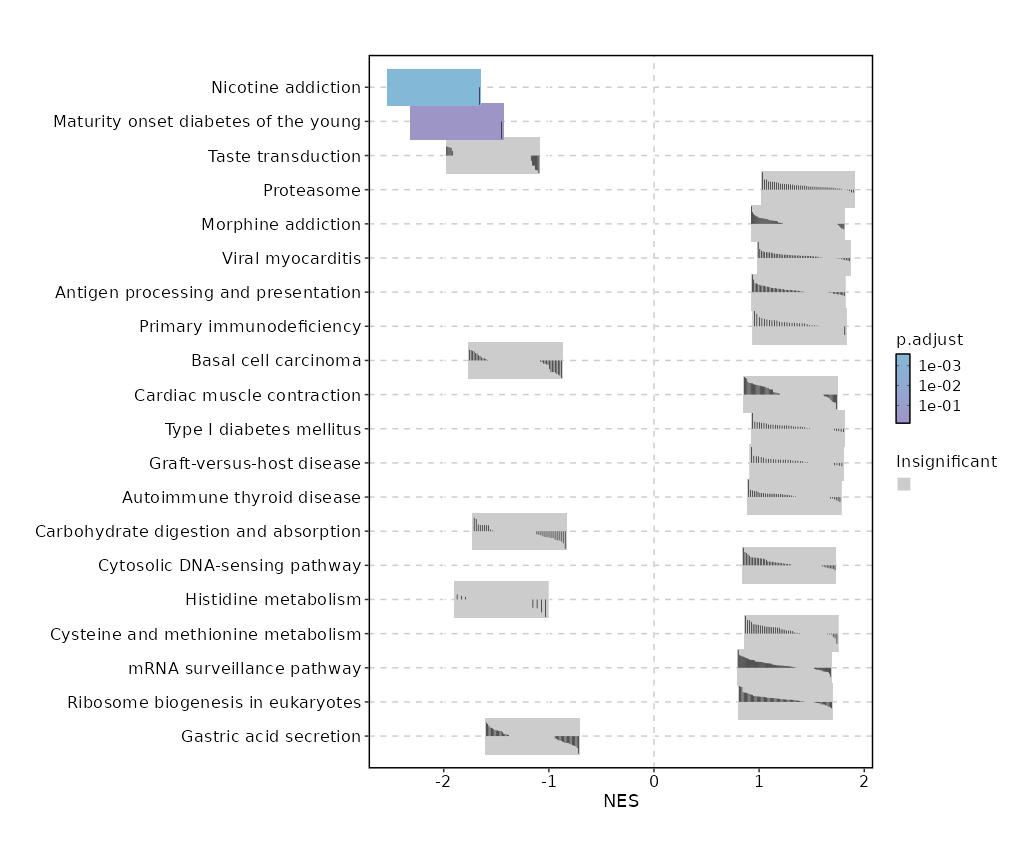
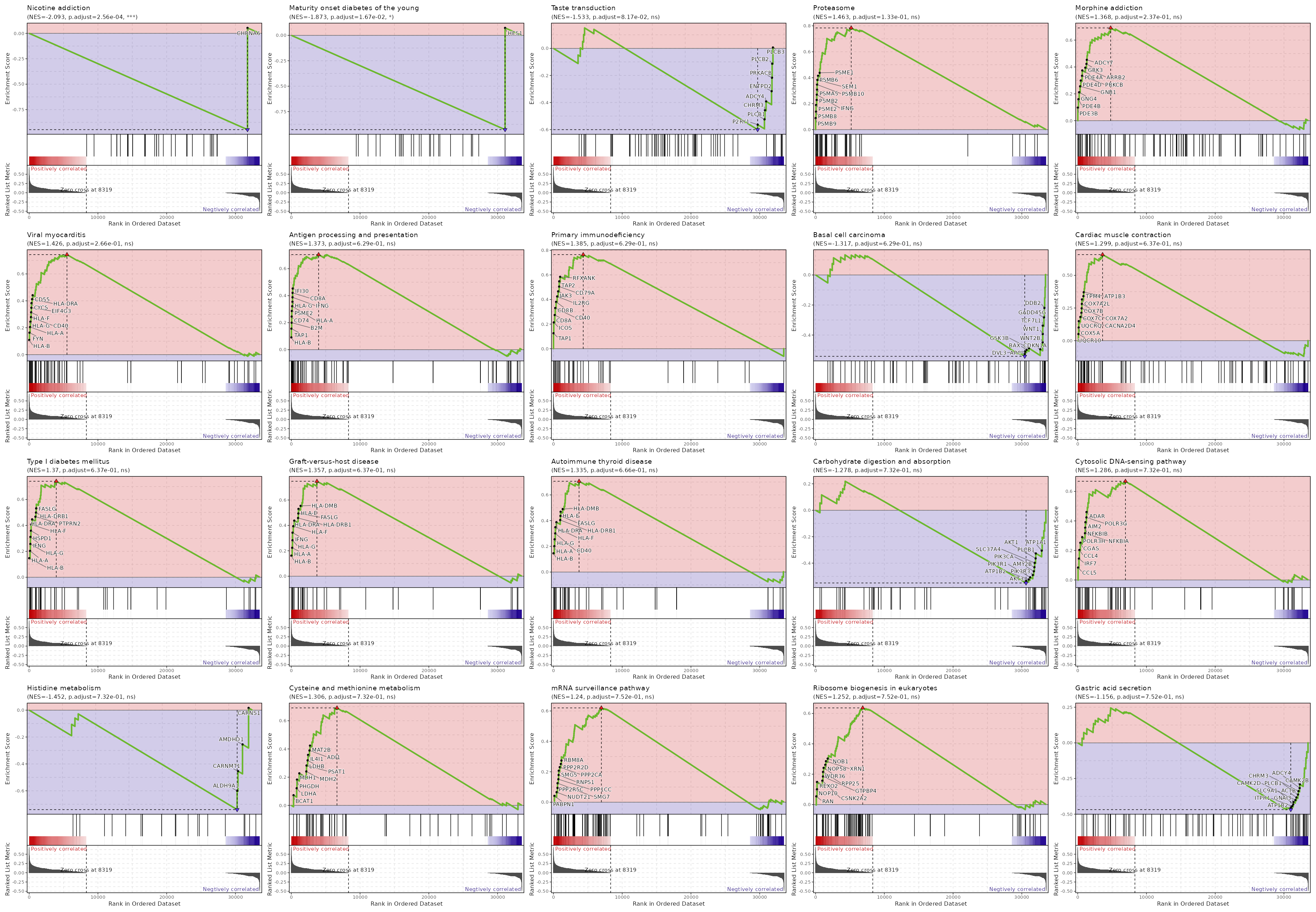
c9
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Nicotine addiction | 0 | 0.0003 | 0.6436 | -0.9469 | -2.0926 | 40 | CHRNA6 |
| Maturity onset diabetes of the young | 0.0002 | 0.0167 | 0.5188 | -0.9272 | -1.8729 | 26 | HES1 |
| Taste transduction | 0.0013 | 0.0817 | 0.4551 | -0.6003 | -1.533 | 85 | PLCB3,PLCB2,PRKACB,ENTPD2,ADCY4,CHRM3,PLCB1,P2RY1 |
| Proteasome | 0.0028 | 0.1333 | 0.4317 | 0.7837 | 1.4633 | 46 | PSMB9,PSMB8,PSME2,PSMB2,IFNG,PSMA5,PSMB10,SEM1,PSMB6,PSME1,PSMA2,POMP,PSMA1,PSMB3,PSMD6,PSMA7,PSMB4,PSMD3,PSMA3,PSMA4,PSMD13,PSMC3,PSMF1,PSMC2,ADRM1,PSMD7,PSMB1,PSMA6,PSMD8,PSMD4,PSMC6,PSMC4,PSMD9,PSME3,PSMC5,PSMB7 |
| Morphine addiction | 0.0062 | 0.2367 | 0.407 | 0.6907 | 1.3677 | 91 | PDE3B,PDE4B,GNG4,GNB1,PDE4D,PRKCB,PDE4A,ARRB2,GRK3,ADCY7,GNG2,GNB2,GNG5,GNG10,GRK2,GNB4,PDE7B,PDE7A,GNAO1,PDE2A,GNG12,PRKACA,PDE1B,GNAS,GNB5 |
| Viral myocarditis | 0.0084 | 0.2661 | 0.3807 | 0.7435 | 1.4261 | 57 | HLA-B,FYN,HLA-A,HLA-G,CD40,HLA-F,EIF4G3,CYCS,HLA-DRA,CD55,RAC2,CCND1,HLA-DRB1,HLA-E,HLA-DMB,HLA-DPA1,HLA-C,PRF1,CD80,CD40LG,CD28,CASP9,EIF4G1,RAC1,SGCB,ABL2,HLA-DPB1,CXADR,DMD,HLA-DOA,CAV1,ABL1,ICAM1,HLA-DMA,HLA-DRB5,EIF4G2,CASP3 |
| Antigen processing and presentation | 0.0263 | 0.6294 | 0.3525 | 0.7043 | 1.3728 | 69 | HLA-B,TAP1,B2M,CD74,PSME2,HLA-A,HLA-G,IFNG,CD8A,IFI30,HLA-F,CD8B,HLA-DRA,PSME1,TAP2,RFXANK,HLA-DRB1,HLA-E,HLA-DMB,HSPA2,HLA-DPA1,HLA-C,KLRD1,TAPBP,RFXAP,HLA-DPB1,CD8B2,HLA-DOA,HSPA6,CIITA,HLA-DMA,CTSB,PSME3,HLA-DRB5 |
| Primary immunodeficiency | 0.0296 | 0.6294 | 0.3525 | 0.7651 | 1.3847 | 37 | TAP1,ICOS,CD8A,CD40,CD8B,IL2RG,JAK3,CD79A,TAP2,RFXANK,LCK,CD40LG,ORAI1,ADA,CD3D,IL7R,DCLRE1C,RFXAP,CD8B2,CIITA,PTPRC |
| Basal cell carcinoma | 0.0297 | 0.6294 | 0.3525 | -0.5413 | -1.3175 | 63 | WNT10A,POLK,PTCH2,DDB2,GADD45G,TCF7L1,WNT1,WNT2B,CDKN1A,GSK3B,BAX,DVL3,APC |
| Cardiac muscle contraction | 0.0381 | 0.6367 | 0.2413 | 0.6585 | 1.2994 | 83 | UQCR10,COX5A,UQCRQ,CACNA2D4,COX7C,COX7A2,COX7B,COX7A2L,TPM4,ATP1B3,CYC1,ATP2A2,UQCR11,COX6C,UQCRFS1,COX8A,COX4I1,COX6B1,UQCRH,ASPH,COX6A1,UQCRC1,COX5B,MYL3,TNNC1,FXYD2 |
| Type I diabetes mellitus | 0.0398 | 0.6367 | 0.245 | 0.7407 | 1.3695 | 41 | HLA-B,HLA-A,HLA-G,IFNG,HSPD1,HLA-F,HLA-DRA,PTPRN2,HLA-DRB1,FASLG,HLA-E,HLA-DMB,GZMB,HLA-DPA1,HLA-C,PRF1,CD80,CD28,HLA-DPB1,IL2,HLA-DOA,CPE |
| Graft-versus-host disease | 0.04 | 0.6367 | 0.2489 | 0.7477 | 1.3569 | 38 | HLA-B,HLA-A,HLA-G,IFNG,HLA-F,HLA-DRA,HLA-DRB1,FASLG,HLA-E,HLA-DMB,GZMB,HLA-DPA1,HLA-C,PRF1,CD80,KLRD1,CD28,HLA-DPB1,IL2,HLA-DOA |
| Autoimmune thyroid disease | 0.0453 | 0.6663 | 0.228 | 0.7062 | 1.3349 | 50 | HLA-B,HLA-A,HLA-G,CD40,HLA-F,HLA-DRA,HLA-DRB1,FASLG,HLA-E,HLA-DMB,GZMB,HLA-DPA1,HLA-C,PRF1,CD80,TG,TSHR,CD40LG,CD28,HLA-DPB1,IL2,HLA-DOA |
| Carbohydrate digestion and absorption | 0.0569 | 0.7316 | 0.3218 | -0.5499 | -1.2776 | 46 | ATP1B1,PLCB3,PLCB2,CACNA1D,ATP1A4,ATP1A1,PLCB1,AKT1,SLC37A4,PIK3CA,AMY2B,PIK3R3,PIK3R1,ATP1B2,AKT3 |
| Cytosolic DNA-sensing pathway | 0.0615 | 0.7316 | 0.1919 | 0.6686 | 1.2865 | 61 | CCL5,IRF7,CCL4,CGAS,POLR3H,NFKBIA,NFKBIB,AIM2,POLR3G,ADAR,POLR2L,POLR3A,PYCARD,CCL4L2,CXCL10,POLR2K,POLR2E,IKBKB,IKBKE,CHUK,POLR3E,POLR3D,IRF3,TBK1,MAVS,POLR2H,POLR1C,DDX58,ZBP1,IKBKG,TREX1 |
| Histidine metabolism | 0.0631 | 0.7316 | 0.2878 | -0.7401 | -1.4521 | 22 | CARNS1,AMDHD1,CARNMT1,ALDH9A1 |
| Cysteine and methionine metabolism | 0.0651 | 0.7316 | 0.1882 | 0.691 | 1.3062 | 50 | BCAT1,LDHA,PHGDH,MDH1,MDH2,PSAT1,LDHB,IL4I1,ADI1,MAT2B,SMS,AMD1,MPST,CDO1,GOT2,SRM,GCLM,APIP,TST,GSS,MRI1,MAT2A,GOT1,DNMT3A,MTR,KYAT3 |
| mRNA surveillance pathway | 0.0724 | 0.7516 | 0.1696 | 0.62 | 1.2401 | 98 | PABPN1,NUDT21,PPP2R5C,SMG7,PPP1CC,RNPS1,SMG5,PPP2CA,PPP2R2D,RBM8A,SSU72,HBS1L,PYM1,GSPT1,CSTF2T,PPP1CA,ACIN1,PPP2R3C,WDR33,SRRM1,ALYREF,MAGOH,SMG1,RNMT,NCBP2,SAP18,NXF1,WDR82,PPP2CB,PNN,UPF2,CSTF2,UPF3A,NXT1,PAPOLA,CSTF1,PPP1CB,UPF3B,PPP2R5D,SYMPK,CASC3,PPP2R5E,PABPC1L,PPP2R5A,CPSF7,ETF1,FIP1L1,MAGOHB,CPSF2,DDX19A,PABPC1,CLP1,UPF1,CPSF4 |
| Ribosome biogenesis in eukaryotes | 0.0783 | 0.7516 | 0.1657 | 0.6365 | 1.2516 | 79 | RAN,NOP10,REXO2,CSNK2A2,GTPBP4,RPP25,WDR36,NOP58,XRN1,NOB1,SBDS,RIOK1,NMD3,POP1,BMS1,CSNK2B,GNL3,SNU13,HEATR1,GAR1,NXF1,WDR3,NHP2,EMG1,WDR43,IMP3,NXT1,RPP30,DKC1,POP7,RRP7A,IMP4,EIF6,UTP14C,UTP14A,GNL2,XRN2,MPHOSPH10,TCOF1,NVL,SPATA5,RCL1,UTP15,RMRP,UTP4,XPO1,NOP56,RPP25L,UTP6 |
| Gastric acid secretion | 0.0787 | 0.7516 | 0.2878 | -0.4682 | -1.1564 | 76 | ATP1B1,PLCB3,PLCB2,PRKACB,KCNQ1,ATP1A4,ATP1A1,CALM1,CAMK2B,ADCY4,CHRM3,PLCB1,ACTB,CAMK2D,GNAI3,SLC9A1,ITPR1,ATP1B2 |
Downloading entire data
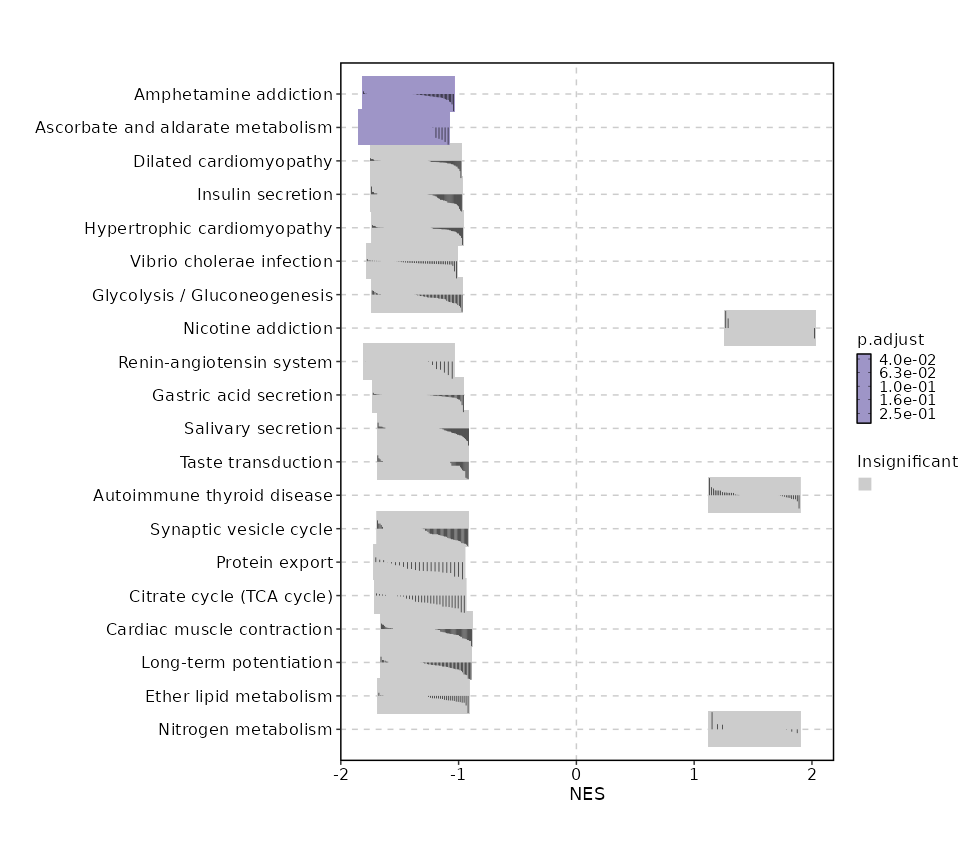
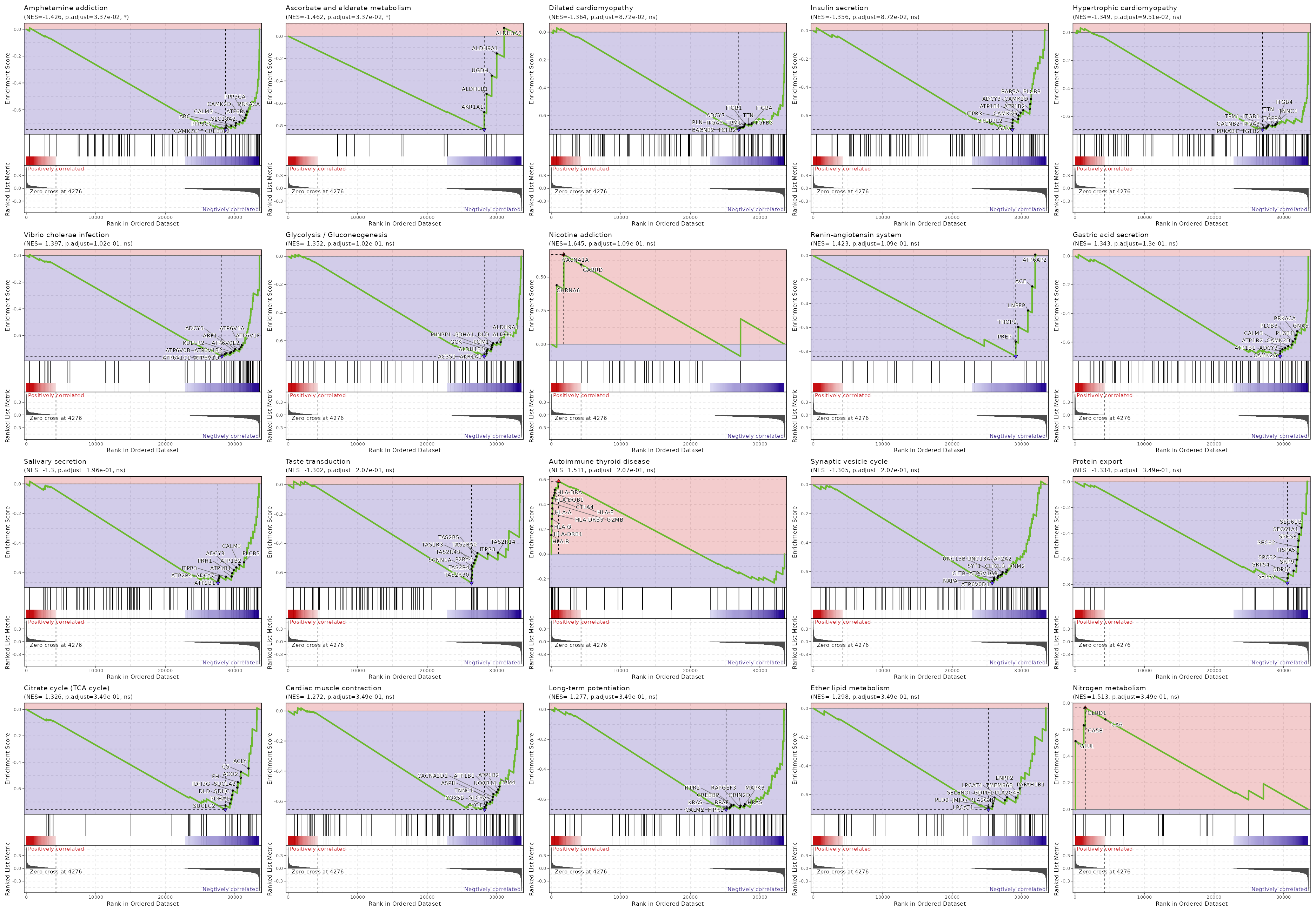
c2
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Amphetamine addiction | 0.0002 | 0.0337 | 0.5188 | -0.7476 | -1.4263 | 69 | FOS,FOSB,CALM1,PPP1CA,PRKACB,PPP1CC,HDAC1,HDAC2,PPP1CB,PPP3R1,GNAS,PRKACA,PPP3CA,ATF6B,CAMK2D,CALM3,SLC18A2,ARC,CAMK2G,PPP3CC,CREB3L2 |
| Ascorbate and aldarate metabolism | 0.0004 | 0.0337 | 0.4985 | -0.8358 | -1.4623 | 30 | ALDH3A2,ALDH9A1,UGDH,ALDH1B1,AKR1A1 |
| Dilated cardiomyopathy | 0.0014 | 0.0872 | 0.4551 | -0.7009 | -1.3637 | 95 | ACTB,ACTG1,TNF,ITGA1,TPM3,ITGB7,PRKACB,ITGA4,CACNB1,ATP2A3,SGCB,GNAS,PRKACA,TGFB1,ITGA2,TPM4,ADCY3,DAG1,CACNA2D2,TNNC1,ITGB4,TGFB3,TTN,ITGB1,ADCY7,TPM1,ITGA5,PLN,CACNB2,TGFB2 |
| Insulin secretion | 0.0018 | 0.0872 | 0.4551 | -0.6996 | -1.3564 | 86 | PRKACB,PLCB2,ATP1A1,FXYD2,KCNN4,GNAS,PRKACA,PLCB1,KCNMB3,ATF6B,VAMP2,PLCB3,RAB3A,CAMK2D,ATP1B2,ADCY3,ATP1B1,CAMK2G,ITPR3,CREB3L2,GCK |
| Hypertrophic cardiomyopathy | 0.0025 | 0.0951 | 0.4317 | -0.6943 | -1.3491 | 90 | ACTB,ACTG1,TNF,ITGA1,TPM3,ITGB7,ITGA4,PRKAB2,CACNB1,ACE,ATP2A3,SGCB,TGFB1,ITGA2,TPM4,PRKAG1,DAG1,CACNA2D2,TNNC1,ITGB4,TGFB3,TTN,ITGB1,TPM1,ITGA5,PRKAB1,CACNB2,TGFB2 |
| Vibrio cholerae infection | 0.0033 | 0.1018 | 0.4317 | -0.7561 | -1.3968 | 50 | ACTB,ACTG1,PRKACB,PDIA4,SEC61G,KCNQ1,SEC61B,ATP6V1G1,TCIRG1,SEC61A1,ATP6V0A1,ATP6AP1,GNAS,PLCG1,PRKACA,ATP6V1E1,ATP6V1G2,KDELR1,ATP6V1E2,ATP6V1F,ATP6V1A,ATP6V0E2,ARF1,ADCY3,KDELR2,ATP6V1B2,ATP6V0B,ATP6V1C1,ATP6V1D |
| Glycolysis / Gluconeogenesis | 0.0037 | 0.1018 | 0.4317 | -0.7106 | -1.3523 | 66 | PKM,GAPDH,ENO1,ALDOA,GPI,PGAM1,ENO2,PFKL,ADH5,TPI1,FBP1,PGK1,HK1,ALDH3A2,PGM2,LDHA,ALDOC,ALDH9A1,DLD,PGM1,PDHA1,MINPP1,GCK,ALDH1B1,ACSS1,AKR1A1 |
| Nicotine addiction | 0.005 | 0.1087 | 0.407 | 0.6674 | 1.6454 | 40 | CHRNA6,CACNA1A,GABRD |
| Renin-angiotensin system | 0.0051 | 0.1087 | 0.407 | -0.8406 | -1.4232 | 23 | ATP6AP2,ACE,LNPEP,THOP1,PREP |
| Gastric acid secretion | 0.0068 | 0.1298 | 0.407 | -0.6989 | -1.3428 | 76 | ACTB,ACTG1,CALM1,PRKACB,PLCB2,ATP1A1,KCNQ1,GNAS,PRKACA,PLCB1,PLCB3,CAMK2D,CALM3,ATP1B2,ADCY3,ATP1B1,CAMK2G |
| Salivary secretion | 0.0113 | 0.1956 | 0.3807 | -0.6687 | -1.3004 | 93 | CALM1,PRKACB,PLCB2,ATP1A1,ADRB2,FXYD2,KCNN4,GNAS,PRKACA,PLCB1,VAMP2,PLCB3,CALM3,ATP1B2,ADCY3,PRH1,ATP1B1,ITPR3,ADCY7,ATP2B4,ATP2B1 |
| Taste transduction | 0.0131 | 0.2066 | 0.3807 | -0.6705 | -1.3024 | 85 | KCNK5,PRKACB,PLCB2,PRKACA,PLCB1,PLCB3,TAS2R14,ITPR3,TAS2R50,TAS2R5,TAS1R3,TAS2R43,P2RY4,SCNN1A,TAS2R4,TAS2R30 |
| Autoimmune thyroid disease | 0.015 | 0.2066 | 0.3807 | 0.5878 | 1.5111 | 50 | HLA-B,HLA-DRB1,HLA-G,HLA-DRB5,HLA-A,GZMB,CTLA4,HLA-DQB1,HLA-E,HLA-DRA,CGA,IL4,TPO,FAS |
| Synaptic vesicle cycle | 0.0151 | 0.2066 | 0.3807 | -0.678 | -1.3049 | 78 | CLTA,AP2B1,ATP6V1G1,TCIRG1,ATP6V0A1,AP2S1,ATP6V1E1,ATP6V1G2,VAMP2,ATP6V1E2,RAB3A,ATP6V1F,ATP6V1A,CPLX1,ATP6V0E2,CLTC,SLC18A2,STX3,ATP6V1B2,AP2M1,ATP6V0B,ATP6V1C1,NSF,ATP6V1D,DNM2,AP2A2,CLTCL1,UNC13A,UNC13B,SYT1,ATP6V1G3,CLTB,NAPA,ATP6V0D1 |
| Protein export | 0.0301 | 0.3492 | 0.282 | -0.788 | -1.334 | 23 | SRPRA,SPCS1,SRP68,SEC63,SEC61G,SEC61B,SEC61A1,SPCS3,SEC62,HSPA5,SPCS2,SRP9,SRP14,SRP54,SRP72 |
| Citrate cycle (TCA cycle) | 0.0305 | 0.3492 | 0.2765 | -0.7577 | -1.3256 | 30 | MDH1,DLST,ACO1,PC,MDH2,SUCLG1,IDH2,ACLY,CS,ACO2,FH,SUCLA2,IDH3G,SDHC,DLD,PDHA1,SUCLG2 |
| Cardiac muscle contraction | 0.0369 | 0.3492 | 0.2378 | -0.6563 | -1.272 | 83 | TPM3,ATP1A1,COX8A,CACNB1,COX6B1,ATP2A3,FXYD2,COX4I1,UQCRC2,UQCRC1,COX6A1,SLC9A6,COX5A,COX7A2,TPM4,ATP1B2,UQCR11,ATP1B1,CACNA2D2,ASPH,TNNC1,SLC9A7,COX5B,CYC1 |
| Long-term potentiation | 0.0417 | 0.3492 | 0.225 | -0.6707 | -1.2772 | 67 | CALM1,RAP1A,PPP1CA,PRKACB,PPP1CC,PLCB2,RPS6KA1,PPP1CB,PPP3R1,PRKACA,PLCB1,PPP3CA,PLCB3,CAMK2D,CALM3,NRAS,RPS6KA2,CAMK2G,MAPK1,PPP3CC,ITPR3,RAF1,MAPK3,HRAS,GRIN2D,RAPGEF3,ITPR2,CREBBP,BRAF,KRAS,CALM2,ITPR1 |
| Ether lipid metabolism | 0.0432 | 0.3492 | 0.225 | -0.7041 | -1.2981 | 46 | CHPT1,PLD1,PAFAH2,PAFAH1B3,PLPP2,PLA2G6,PLD3,CEPT1,PLPP1,AGPS,PAFAH1B1,ENPP2,PLA2G4B,TMEM86B,LPCAT4,GDPD1,SELENOI,JMJD7-PLA2G4B,PLD2,LPCAT1 |
| Nitrogen metabolism | 0.0449 | 0.3492 | 0.3218 | 0.7628 | 1.5131 | 17 | GLUL,CA5B,GLUD1,CA6 |
Downloading entire data
c8
Showing top 50 pathways by padj in descending order. Use 'Download the entire data' button to download all pathways.
| Nicotine addiction | 0.0001 | 0.0109 | 0.5573 | -0.8792 | -1.6304 | 40 | CACNA1A |
| Autoimmune thyroid disease | 0.0001 | 0.0133 | 0.5188 | 0.8061 | 1.857 | 50 | GZMB,HLA-B,PRF1,HLA-A,HLA-DRB1,HLA-DQB1,CTLA4,HLA-E,HLA-DPB1,HLA-G,HLA-F,HLA-DRB5,HLA-DRA,HLA-DQA1,HLA-DOA |
| Viral myocarditis | 0.0004 | 0.02 | 0.4985 | 0.7168 | 1.699 | 57 | HLA-B,PRF1,CD55,HLA-A,HLA-DRB1,HLA-DQB1,FYN,HLA-E,HLA-DPB1,HLA-G,HLA-F,HLA-DRB5,HLA-DRA,HLA-DQA1,EIF4G3,BID,CASP3,HLA-DOA,CASP8,CD28,ABL1,DAG1,RAC1 |
| Maturity onset diabetes of the young | 0.0004 | 0.02 | 0.4985 | -0.9128 | -1.5763 | 26 | HHEX,HES1,NEUROG3 |
| Type I diabetes mellitus | 0.0011 | 0.0406 | 0.4551 | 0.7792 | 1.7175 | 41 | GZMB,HLA-B,PRF1,HLA-A,HLA-DRB1,HLA-DQB1,HLA-E,HLA-DPB1,HLA-G,HLA-F,HLA-DRB5,HLA-DRA,HLA-DQA1,HSPD1,HLA-DOA,PTPRN,IL12A,CD28,PTPRN2 |
| Phototransduction | 0.0061 | 0.1947 | 0.407 | -0.8253 | -1.4404 | 28 | CALM1,PDE6G,GUCY2D,CALM3,PDE6B,CALM2 |
| Allograft rejection | 0.0093 | 0.255 | 0.3807 | 0.7644 | 1.6466 | 35 | GZMB,HLA-B,PRF1,HLA-A,HLA-DRB1,HLA-DQB1,HLA-E,HLA-DPB1,HLA-G,HLA-F,HLA-DRB5,HLA-DRA,HLA-DQA1,HLA-DOA,IL12A,CD28 |
| Asthma | 0.0252 | 0.6014 | 0.3525 | 0.713 | 1.4467 | 28 | HLA-DRB1,HLA-DQB1,HLA-DPB1,HLA-DRB5,HLA-DRA,HLA-DQA1,FCER1G,HLA-DOA |
| Graft-versus-host disease | 0.0319 | 0.6776 | 0.3218 | 0.6558 | 1.429 | 38 | GZMB,HLA-B,PRF1,HLA-A,HLA-DRB1,HLA-DQB1,HLA-E,KLRC1,HLA-DPB1,HLA-G,HLA-F,HLA-DRB5,HLA-DRA,HLA-DQA1,KIR2DL1,HLA-DOA |
| Pyruvate metabolism | 0.0398 | 0.7594 | 0.2489 | -0.7171 | -1.3578 | 47 | GLO1,DLAT,ACACB,LDHB,ADH5,ACYP1,MDH1,FH,ACAT2,PDHA1,ALDH9A1,PKM,ALDH1B1,ACSS2 |
| Taste transduction | 0.0458 | 0.795 | 0.3218 | 0.4989 | 1.2703 | 85 | P2RY1,ITPR3,PDE1B,TAS2R20,CACNA1C,ADCY4,PRKACA,KCNK5 |
| Glycolysis / Gluconeogenesis | 0.0537 | 0.8553 | 0.209 | -0.6631 | -1.3024 | 66 | DLAT,ENO2,PGM1,LDHB,ADH5,ALDOC,GPI,BPGM,PGM2,MINPP1,PDHA1,FBP1,ALDH9A1,ENO1,ALDOA,PGAM1,HK1,PKM,ALDH1B1,GAPDH |
| Tyrosine metabolism | 0.0996 | 0.9947 | 0.1563 | -0.7124 | -1.3006 | 36 | FAH,GOT2,ADH5,MIF,IL4I1,LRTOMT |
| Amphetamine addiction | 0.108 | 0.9947 | 0.1429 | -0.6244 | -1.2295 | 69 | FOS,FOSB,CALM1,PRKACB,HDAC1,PPP1CA,PPP3R1,PPP1CB,CREB3L3,PRKCA,HDAC2,CREB1,CALM3,PPP3CC,CREB3,CREB3L4,CREB3L2,CALM2,SLC18A2,ARC,GNAS,CAMK2G,JUN,PPP1CC,ATF6B,PRKCB,PPP3CB |
| Endocrine and other factor-regulated calcium reabsorption | 0.1308 | 0.9947 | 0.1301 | -0.6342 | -1.2165 | 53 | PRKACB,ATP1A1,ATP1B1,PRKCA,DNM1,PLCB1,AP2M1,DNM2,AP2B1,ADCY6,DNM3,GNAQ,ATP1A3,CLTCL1,GNAS,RAB11A,AP2A1,PLCB2,PLCB3,PRKCB,CLTB |
| Thyroid hormone synthesis | 0.1313 | 0.9947 | 0.1275 | -0.6105 | -1.2119 | 74 | PRKACB,ATP1A1,ATP1B1,CANX,HSPA5,HSP90B1,CREB3L3,PRKCA,PLCB1,CREB1,ITPR2,PDIA4,ADCY3,ADCY6,GNAQ,ATP1A3,TTF2,CREB3,PAX8,CREB3L4,CREB3L2,GPX3 |
| Metabolism of xenobiotics by cytochrome P450 | 0.1354 | 0.9947 | 0.1256 | -0.6061 | -1.1988 | 72 | GSTP1,MGST3,GSTA4,GSTM3,ADH5,GSTM1,AKR7A2,GSTK1,DHDH,CBR1,CYP2F1,CYP2D6 |
| Viral protein interaction with cytokine and cytokine receptor | 0.1392 | 0.9947 | 0.1204 | -0.5786 | -1.1808 | 98 | XCL1,CXCR3,XCL2,CCL4L2,CCL3L1,TNFSF14,IL2,TNFRSF10A,TNF,TNFRSF1A,IL2RG,IL10RB,IL6R |
| Morphine addiction | 0.1424 | 0.9947 | 0.2492 | 0.4646 | 1.1815 | 91 | PDE4B,PDE3B,PDE4D,GNG2,GRK5,PDE1B,GNG4,ADCY4,GNB4,GRK3,ARRB2,PRKACA,PDE7B,ADCY9,PDE4A,PDE7A,GNG10 |
| Fanconi anemia pathway | 0.1425 | 0.9947 | 0.1238 | -0.6294 | -1.2096 | 54 | RPA1,CENPS-CORT,FANCE,FANCD2,MUS81,CENPS,EME2,ERCC4,RAD51C,RPA2,PMS2,TOP3A,HES1,TELO2,FAAP100,FAN1,ATRIP,FANCI,POLN,RPA4,SLX1A,UBE2T,ERCC1,TOP3B,FANCB,SLX1B,WDR48,REV3L,RMI1,POLK,FANCM,FANCF |
Downloading entire data
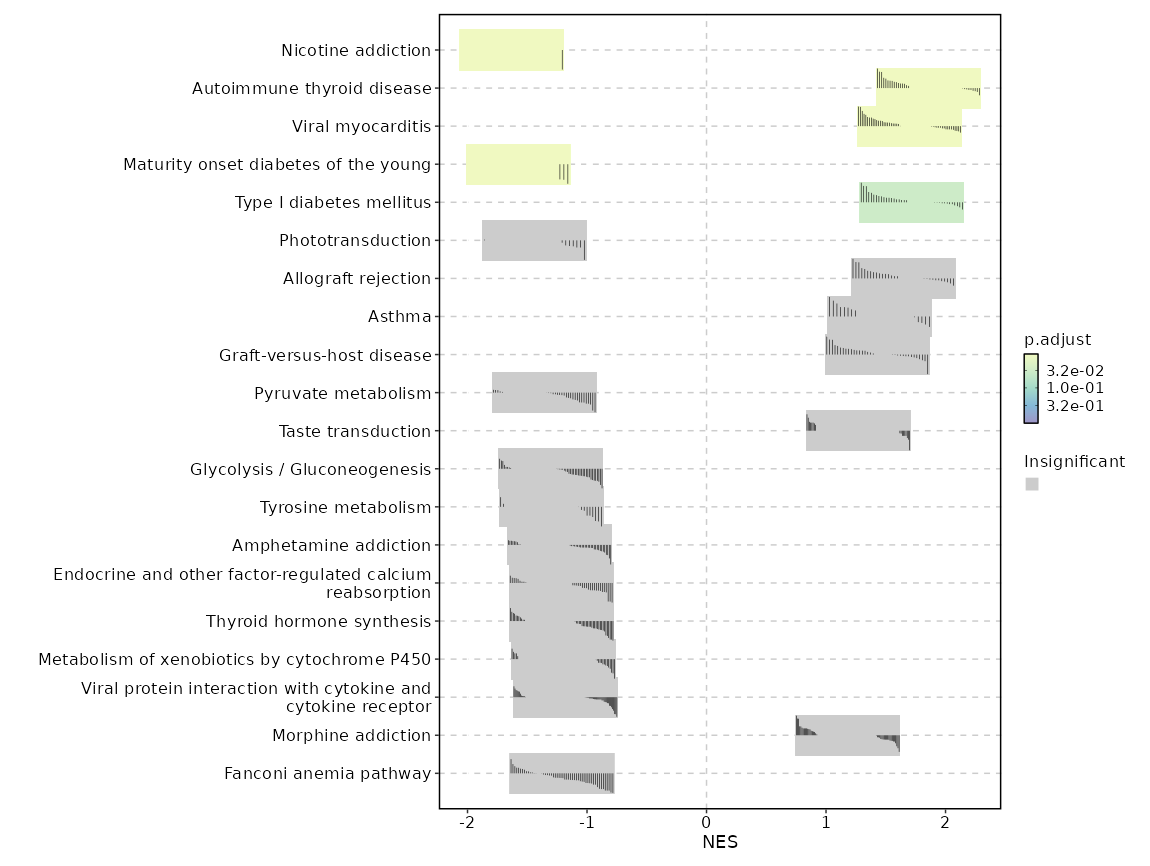
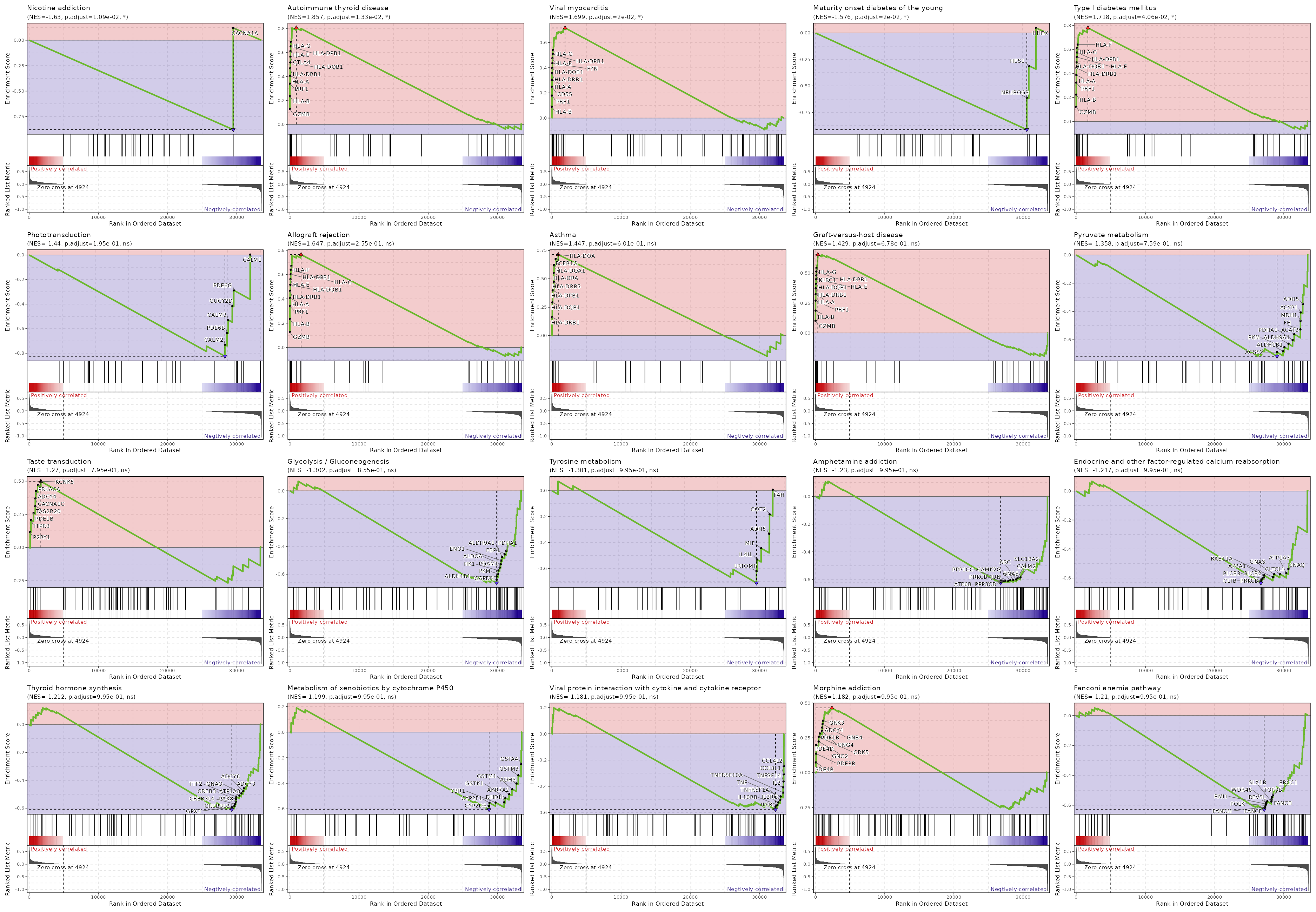
GSEA (all seurat_clusters)
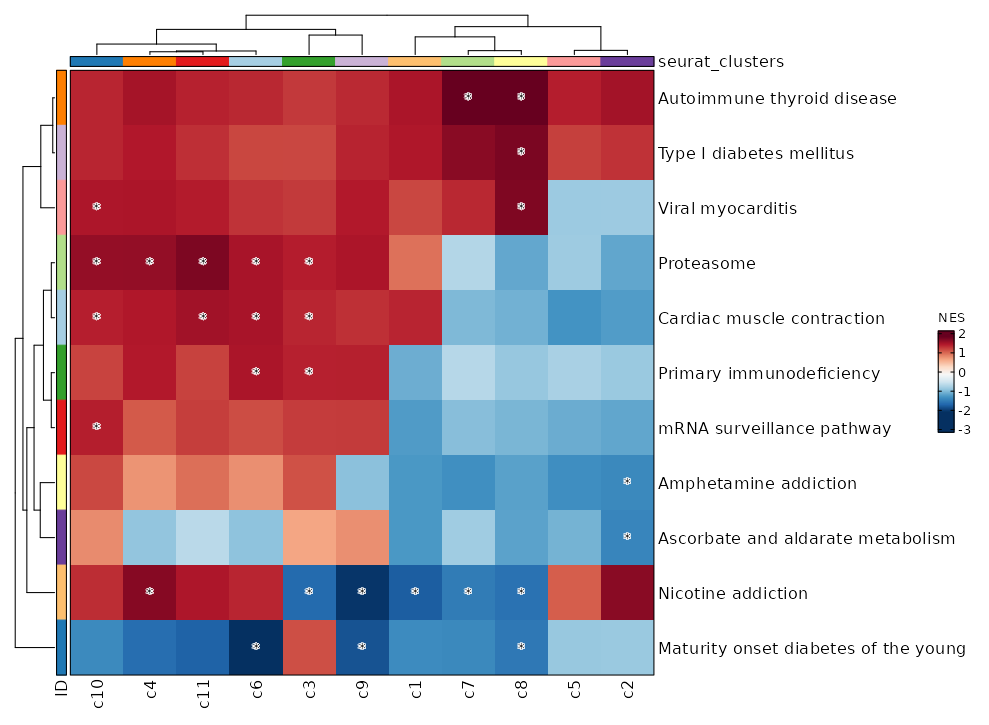
Heatmap
Pathways for all seurat_clusters.
Image loading error!
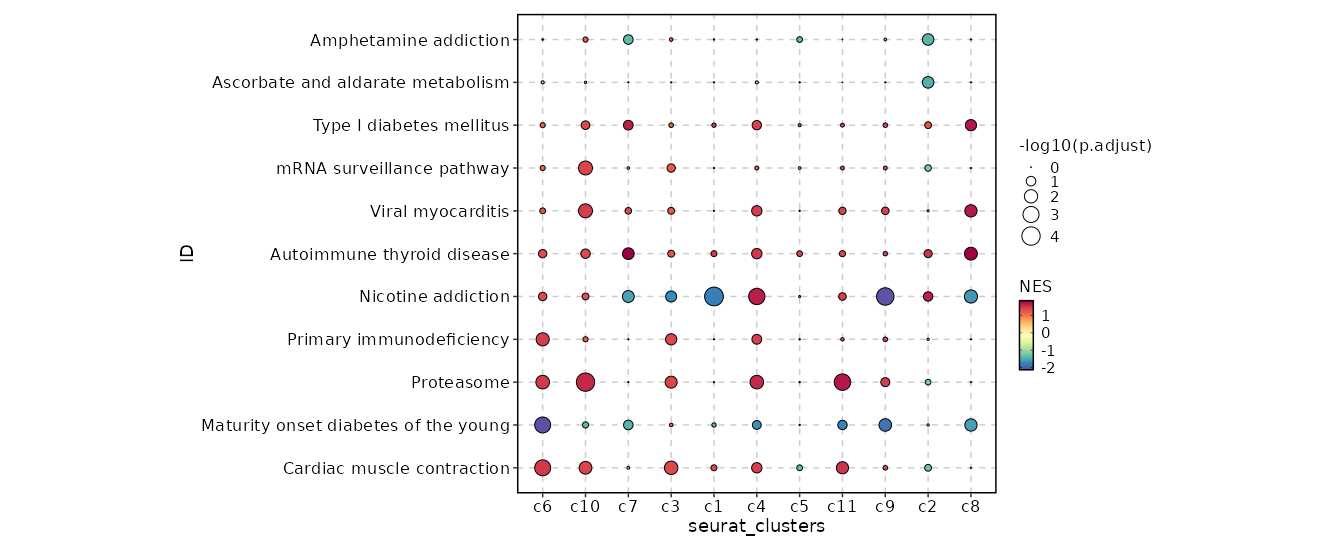
DotPlot
Pathways for all seurat_clusters.
Image loading error!